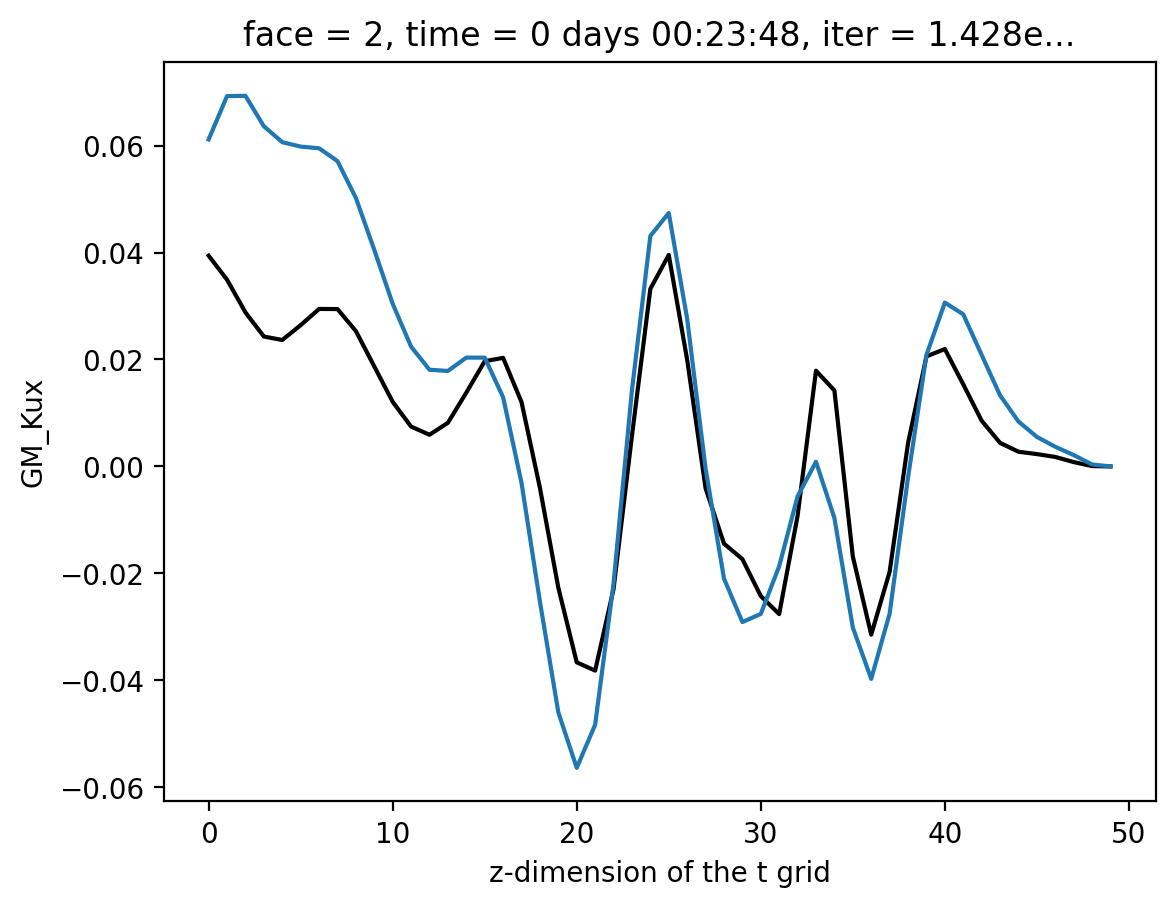
Reading ECCO \(χ_{redi}\)#
%load_ext watermark
from pathlib import Path
import cf_xarray as cfxr
import dask
import ecco_v4_py as ecco
import matplotlib as mpl
import ncar_jobqueue
import numpy as np
import xgcm
from distributed import Client
import xarray as xr
%watermark -iv
xr.set_options(display_expand_data=False)
dirname = "/glade/work/dcherian/mitgcm/ECCOV4/release4/run/diags/"
def open_directory(dirname):
import xmitgcm
ds = xr.merge(
[
dask.optimize(
xmitgcm.open_mdsdataset(
str(path),
dirname + "../",
geometry="llc",
chunks={"face": 1, "k_l": -1, "k": -1},
llc_method="smallchunks",
read_grid=idx == 0,
default_dtype=np.float32,
)
.isel(time=slice(-12))
.chunk({"time": 120})
)[0]
for idx, path in enumerate(Path(dirname).glob("*"))
],
compat="override",
) # .sel(face=2)
ds.coords["drC_kl"] = ds.drC.isel(k_p1=slice(-1)).rename({"k_p1": "k_l"}).variable
grid = xgcm.Grid(
ds,
periodic=False,
boundary="fill",
fill_value=np.nan,
metrics={
("X",): ["dxC", "dxG"], # X distances
("Y",): ["dyC", "dyG"], # Y distances
("Z",): ["drF", "drC", "drC_kl"], # Z distances
("X", "Y"): ["rA", "rAz", "rAs", "rAw"], # Areas
},
)
ds["Tx"] = grid.derivative(ds.THETA, "X")
ds["Ty"] = grid.derivative(ds.THETA, "Y")
ds["Tz"] = -1 * grid.derivative(ds.THETA, "Z")
ds["GM_CHI"] = (
grid.interp(ds.GM_CHIX + ds.GM_CHIXO, "X", to="center")
+ grid.interp(ds.GM_CHIY + ds.GM_CHIYO, "Y", to="center")
+ grid.interp(ds.GM_CHIZ, "Z", to="center")
)
return ds, grid
The watermark extension is already loaded. To reload it, use:
%reload_ext watermark
pandas : 1.5.3
cf_xarray : 0.8.0
dask : 2023.3.2
numpy : 1.23.5
ncar_jobqueue: 2021.4.14
xgcm : 0.6.1
json : 2.0.9
ecco_v4_py : 1.5.5
xmitgcm : 0.5.2
sys : 3.10.10 | packaged by conda-forge | (main, Mar 24 2023, 20:08:06) [GCC 11.3.0]
matplotlib : 3.7.1
xarray : 2023.3.0
cluster = ncar_jobqueue.NCARCluster(
local_directory="/local_scratch/pbs.$PBS_JOBID/dask/spill"
)
cluster
cluster.adapt(minimum_jobs=1, maximum_jobs=32)
/glade/u/home/dcherian/miniconda3/envs/eddydiff/lib/python3.10/site-packages/dask_jobqueue/core.py:255: FutureWarning: job_extra has been renamed to job_extra_directives. You are still using it (even if only set to []; please also check config files). If you did not set job_extra_directives yet, job_extra will be respected for now, but it will be removed in a future release. If you already set job_extra_directives, job_extra is ignored and you can remove it.
warnings.warn(warn, FutureWarning)
/glade/u/home/dcherian/miniconda3/envs/eddydiff/lib/python3.10/site-packages/dask_jobqueue/core.py:274: FutureWarning: env_extra has been renamed to job_script_prologue. You are still using it (even if only set to []; please also check config files). If you did not set job_script_prologue yet, env_extra will be respected for now, but it will be removed in a future release. If you already set job_script_prologue, env_extra is ignored and you can remove it.
warnings.warn(warn, FutureWarning)
/glade/u/home/dcherian/miniconda3/envs/eddydiff/lib/python3.10/site-packages/dask_jobqueue/core.py:255: FutureWarning: job_extra has been renamed to job_extra_directives. You are still using it (even if only set to []; please also check config files). If you did not set job_extra_directives yet, job_extra will be respected for now, but it will be removed in a future release. If you already set job_extra_directives, job_extra is ignored and you can remove it.
warnings.warn(warn, FutureWarning)
/glade/u/home/dcherian/miniconda3/envs/eddydiff/lib/python3.10/site-packages/dask_jobqueue/core.py:274: FutureWarning: env_extra has been renamed to job_script_prologue. You are still using it (even if only set to []; please also check config files). If you did not set job_script_prologue yet, env_extra will be respected for now, but it will be removed in a future release. If you already set job_script_prologue, env_extra is ignored and you can remove it.
warnings.warn(warn, FutureWarning)
<distributed.deploy.adaptive.Adaptive at 0x2abdfae84310>
client = Client(cluster)
client
Client
Client-56c116a5-e061-11ed-bfe1-3cecef1acc54
| Connection method: Cluster object | Cluster type: dask_jobqueue.PBSCluster |
| Dashboard: https://jupyterhub.hpc.ucar.edu/stable/user/dcherian/proxy/8787/status |
Cluster Info
PBSCluster
a2e14ca0
| Dashboard: https://jupyterhub.hpc.ucar.edu/stable/user/dcherian/proxy/8787/status | Workers: 0 |
| Total threads: 0 | Total memory: 0 B |
Scheduler Info
Scheduler
Scheduler-857d4c85-b42a-4e09-88ac-de3f3562c798
| Comm: tcp://10.12.206.19:44019 | Workers: 0 |
| Dashboard: https://jupyterhub.hpc.ucar.edu/stable/user/dcherian/proxy/8787/status | Total threads: 0 |
| Started: Just now | Total memory: 0 B |
Workers
%ls /glade/work/dcherian/mitgcm/ECCOV4/release4/run/diags/
GM_CHIX/ GM_CHIY/ GM_CHIZ/ GM_Kuz/ GM_Kvz/ GM_Kwy/ GM_KwzTz/ THETA/
GM_CHIXO/ GM_CHIYO/ GM_Kux/ GM_Kvy/ GM_Kwx/ GM_Kwz/ GM_ubT/
Compare#
adj, grid = open_directory("/glade/work/dcherian/mitgcm/ECCOV4/release4/run/diags/")
const, _ = open_directory("/glade/work/dcherian/mitgcm/ECCOV4/release4/run2/diags/")
display(adj)
display(const)
<xarray.Dataset>
Dimensions: (i: 90, i_g: 90, j: 90, j_g: 90, k: 50, k_u: 50, k_l: 50,
k_p1: 51, face: 13, time: 176)
Coordinates: (12/45)
* i (i) int64 0 1 2 3 4 5 6 7 8 9 ... 80 81 82 83 84 85 86 87 88 89
* i_g (i_g) int64 0 1 2 3 4 5 6 7 8 9 ... 80 81 82 83 84 85 86 87 88 89
* j (j) int64 0 1 2 3 4 5 6 7 8 9 ... 80 81 82 83 84 85 86 87 88 89
* j_g (j_g) int64 0 1 2 3 4 5 6 7 8 9 ... 80 81 82 83 84 85 86 87 88 89
* k (k) int64 0 1 2 3 4 5 6 7 8 9 ... 40 41 42 43 44 45 46 47 48 49
* k_u (k_u) int64 0 1 2 3 4 5 6 7 8 9 ... 40 41 42 43 44 45 46 47 48 49
... ...
maskCtrlC (k, face, j, i) bool dask.array<chunksize=(50, 1, 90, 90), meta=np.ndarray>
maskCtrlS (k, face, j_g, i) bool dask.array<chunksize=(50, 1, 90, 90), meta=np.ndarray>
rhoRef (k) >f4 dask.array<chunksize=(50,), meta=np.ndarray>
maskCtrlW (k, face, j, i_g) bool dask.array<chunksize=(50, 1, 90, 90), meta=np.ndarray>
iter (time) float64 dask.array<chunksize=(120,), meta=np.ndarray>
drC_kl (k_l) >f4 dask.array<chunksize=(50,), meta=np.ndarray>
Data variables: (12/19)
GM_CHIXO (time, k, face, j, i_g) float32 dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
GM_CHIZ (time, k_l, face, j, i) float32 dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
GM_ubT (time, k, face, j, i_g) float32 dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
GM_CHIX (time, k, face, j, i_g) float32 dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
GM_CHIYO (time, k, face, j_g, i) float32 dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
GM_Kvz (time, k, face, j_g, i) float32 dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
... ...
GM_CHIY (time, k, face, j_g, i) float32 dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
GM_Kuz (time, k, face, j, i_g) float32 dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
Tx (time, k, face, j, i_g) float32 dask.array<chunksize=(120, 50, 1, 90, 1), meta=np.ndarray>
Ty (time, k, face, j_g, i) float32 dask.array<chunksize=(120, 50, 1, 1, 90), meta=np.ndarray>
Tz (time, k_l, face, j, i) float32 dask.array<chunksize=(120, 1, 1, 90, 90), meta=np.ndarray>
GM_CHI (time, k, face, j, i) float32 dask.array<chunksize=(120, 49, 1, 89, 89), meta=np.ndarray>
Attributes:
Conventions: CF-1.6
title: netCDF wrapper of MITgcm MDS binary data
source: MITgcm
history: Created by calling `open_mdsdataset(grid_dir='/glade/work/d...xarray.Dataset
- i: 90
- i_g: 90
- j: 90
- j_g: 90
- k: 50
- k_u: 50
- k_l: 50
- k_p1: 51
- face: 13
- time: 176
- i(i)int640 1 2 3 4 5 6 ... 84 85 86 87 88 89
- standard_name :
- x_grid_index
- axis :
- X
- long_name :
- x-dimension of the t grid
- swap_dim :
- XC
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89]) - i_g(i_g)int640 1 2 3 4 5 6 ... 84 85 86 87 88 89
- standard_name :
- x_grid_index_at_u_location
- axis :
- X
- long_name :
- x-dimension of the u grid
- c_grid_axis_shift :
- -0.5
- swap_dim :
- XG
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89]) - j(j)int640 1 2 3 4 5 6 ... 84 85 86 87 88 89
- standard_name :
- y_grid_index
- axis :
- Y
- long_name :
- y-dimension of the t grid
- swap_dim :
- YC
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89]) - j_g(j_g)int640 1 2 3 4 5 6 ... 84 85 86 87 88 89
- standard_name :
- y_grid_index_at_v_location
- axis :
- Y
- long_name :
- y-dimension of the v grid
- c_grid_axis_shift :
- -0.5
- swap_dim :
- YG
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89]) - k(k)int640 1 2 3 4 5 6 ... 44 45 46 47 48 49
- standard_name :
- z_grid_index
- axis :
- Z
- long_name :
- z-dimension of the t grid
- swap_dim :
- Z
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49]) - k_u(k_u)int640 1 2 3 4 5 6 ... 44 45 46 47 48 49
- standard_name :
- z_grid_index_at_upper_w_location
- axis :
- Z
- long_name :
- z-dimension of the w grid
- c_grid_axis_shift :
- 0.5
- swap_dim :
- Zu
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49]) - k_l(k_l)int640 1 2 3 4 5 6 ... 44 45 46 47 48 49
- standard_name :
- z_grid_index_at_lower_w_location
- axis :
- Z
- long_name :
- z-dimension of the w grid
- c_grid_axis_shift :
- -0.5
- swap_dim :
- Zl
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49]) - k_p1(k_p1)int640 1 2 3 4 5 6 ... 45 46 47 48 49 50
- standard_name :
- z_grid_index_at_w_location
- axis :
- Z
- long_name :
- z-dimension of the w grid
- c_grid_axis_shift :
- (-0.5, 0.5)
- swap_dim :
- Zp1
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50]) - face(face)int640 1 2 3 4 5 6 7 8 9 10 11 12
- standard_name :
- face_index
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12])
- time(time)timedelta64[ns]00:12:12 ... 1 days 11:42:36
- standard_name :
- time
- long_name :
- Time
- axis :
- T
- calendar :
- gregorian
array([ 732000000000, 1428000000000, 2172000000000, 2892000000000, 3636000000000, 4356000000000, 5100000000000, 5844000000000, 6564000000000, 7308000000000, 8028000000000, 8772000000000, 9516000000000, 10188000000000, 10932000000000, 11652000000000, 12396000000000, 13116000000000, 13860000000000, 14604000000000, 15324000000000, 16068000000000, 16788000000000, 17532000000000, 18276000000000, 18948000000000, 19692000000000, 20412000000000, 21156000000000, 21876000000000, 22620000000000, 23364000000000, 24084000000000, 24828000000000, 25548000000000, 26292000000000, 27036000000000, 27708000000000, 28452000000000, 29172000000000, 29916000000000, 30636000000000, 31380000000000, 32124000000000, 32844000000000, 33588000000000, 34308000000000, 35052000000000, 35796000000000, 36492000000000, 37236000000000, 37956000000000, 38700000000000, 39420000000000, 40164000000000, 40908000000000, 41628000000000, 42372000000000, 43092000000000, 43836000000000, 44580000000000, 45252000000000, 45996000000000, 46716000000000, 47460000000000, 48180000000000, 48924000000000, 49668000000000, 50388000000000, 51132000000000, 51852000000000, 52596000000000, 53340000000000, 54012000000000, 54756000000000, 55476000000000, 56220000000000, 56940000000000, 57684000000000, 58428000000000, 59148000000000, 59892000000000, 60612000000000, 61356000000000, 62100000000000, 62772000000000, 63516000000000, 64236000000000, 64980000000000, 65700000000000, 66444000000000, 67188000000000, 67908000000000, 68652000000000, 69372000000000, 70116000000000, 70860000000000, 71556000000000, 72300000000000, 73020000000000, 73764000000000, 74484000000000, 75228000000000, 75972000000000, 76692000000000, 77436000000000, 78156000000000, 78900000000000, 79644000000000, 80316000000000, 81060000000000, 81780000000000, 82524000000000, 83244000000000, 83988000000000, 84732000000000, 85452000000000, 86196000000000, 86916000000000, 87660000000000, 88404000000000, 89076000000000, 89820000000000, 90540000000000, 91284000000000, 92004000000000, 92748000000000, 93492000000000, 94212000000000, 94956000000000, 95676000000000, 96420000000000, 97164000000000, 97836000000000, 98580000000000, 99300000000000, 100044000000000, 100764000000000, 101508000000000, 102252000000000, 102972000000000, 103716000000000, 104436000000000, 105180000000000, 105924000000000, 106620000000000, 107364000000000, 108084000000000, 108828000000000, 109548000000000, 110292000000000, 111036000000000, 111756000000000, 112500000000000, 113220000000000, 113964000000000, 114708000000000, 115380000000000, 116124000000000, 116844000000000, 117588000000000, 118308000000000, 119052000000000, 119796000000000, 120516000000000, 121260000000000, 121980000000000, 122724000000000, 123468000000000, 124140000000000, 124884000000000, 125604000000000, 126348000000000, 127068000000000, 127812000000000, 128556000000000], dtype='timedelta64[ns]') - XC(face, j, i)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- longitude
- long_name :
- longitude
- units :
- degrees_east
- coordinate :
- YC XC
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - YC(face, j, i)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- latitude
- long_name :
- latitude
- units :
- degrees_north
- coordinate :
- YC XC
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - XG(face, j_g, i_g)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- longitude_at_f_location
- long_name :
- longitude
- units :
- degrees_east
- coordinate :
- YG XG
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - YG(face, j_g, i_g)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- latitude_at_f_location
- long_name :
- latitude
- units :
- degrees_north
- coordinate :
- YG XG
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - CS(face, j, i)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- Cos of grid orientation angle
- long_name :
- AngleCS
- units :
- coordinate :
- YC XC
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - SN(face, j, i)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- Sin of grid orientation angle
- long_name :
- AngleSN
- units :
- coordinate :
- YC XC
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - Z(k)>f4dask.array<chunksize=(50,), meta=np.ndarray>
- standard_name :
- depth
- long_name :
- vertical coordinate of cell center
- units :
- m
- positive :
- down
Array Chunk Bytes 200 B 200 B Shape (50,) (50,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - Zp1(k_p1)>f4dask.array<chunksize=(51,), meta=np.ndarray>
- standard_name :
- depth_at_w_location
- long_name :
- vertical coordinate of cell interface
- units :
- m
- positive :
- down
Array Chunk Bytes 204 B 204 B Shape (51,) (51,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - Zu(k_u)>f4dask.array<chunksize=(50,), meta=np.ndarray>
- standard_name :
- depth_at_upper_w_location
- long_name :
- vertical coordinate of upper cell interface
- units :
- m
- positive :
- down
Array Chunk Bytes 200 B 200 B Shape (50,) (50,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - Zl(k_l)>f4dask.array<chunksize=(50,), meta=np.ndarray>
- standard_name :
- depth_at_lower_w_location
- long_name :
- vertical coordinate of lower cell interface
- units :
- m
- positive :
- down
Array Chunk Bytes 200 B 200 B Shape (50,) (50,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - rA(face, j, i)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- cell_area
- long_name :
- cell area
- units :
- m2
- coordinate :
- YC XC
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - dxG(face, j_g, i)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- cell_x_size_at_v_location
- long_name :
- cell x size
- units :
- m
- coordinate :
- YG XC
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - dyG(face, j, i_g)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- cell_y_size_at_u_location
- long_name :
- cell y size
- units :
- m
- coordinate :
- YC XG
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - Depth(face, j, i)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- ocean_depth
- long_name :
- ocean depth
- units :
- m
- coordinate :
- XC YC
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - rAz(face, j_g, i_g)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- cell_area_at_f_location
- long_name :
- cell area
- units :
- m
- coordinate :
- YG XG
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - dxC(face, j, i_g)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- cell_x_size_at_u_location
- long_name :
- cell x size
- units :
- m
- coordinate :
- YC XG
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - dyC(face, j_g, i)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- cell_y_size_at_v_location
- long_name :
- cell y size
- units :
- m
- coordinate :
- YG XC
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - rAw(face, j, i_g)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- cell_area_at_u_location
- long_name :
- cell area
- units :
- m2
- coordinate :
- YG XC
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - rAs(face, j_g, i)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- cell_area_at_v_location
- long_name :
- cell area
- units :
- m2
- coordinate :
- YG XC
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - drC(k_p1)>f4dask.array<chunksize=(51,), meta=np.ndarray>
- standard_name :
- cell_z_size_at_w_location
- long_name :
- cell z size
- units :
- m
Array Chunk Bytes 204 B 204 B Shape (51,) (51,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - drF(k)>f4dask.array<chunksize=(50,), meta=np.ndarray>
- standard_name :
- cell_z_size
- long_name :
- cell z size
- units :
- m
Array Chunk Bytes 200 B 200 B Shape (50,) (50,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - PHrefC(k)>f4dask.array<chunksize=(50,), meta=np.ndarray>
- standard_name :
- cell_reference_pressure
- long_name :
- Reference Hydrostatic Pressure
- units :
- m2 s-2
Array Chunk Bytes 200 B 200 B Shape (50,) (50,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - PHrefF(k_p1)>f4dask.array<chunksize=(51,), meta=np.ndarray>
- standard_name :
- cell_reference_pressure
- long_name :
- Reference Hydrostatic Pressure
- units :
- m2 s-2
Array Chunk Bytes 204 B 204 B Shape (51,) (51,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - hFacC(k, face, j, i)>f4dask.array<chunksize=(50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- cell_vertical_fraction
- long_name :
- vertical fraction of open cell
Array Chunk Bytes 20.08 MiB 1.54 MiB Shape (50, 13, 90, 90) (50, 1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - hFacW(k, face, j, i_g)>f4dask.array<chunksize=(50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- cell_vertical_fraction_at_u_location
- long_name :
- vertical fraction of open cell
Array Chunk Bytes 20.08 MiB 1.54 MiB Shape (50, 13, 90, 90) (50, 1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - hFacS(k, face, j_g, i)>f4dask.array<chunksize=(50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- cell_vertical_fraction_at_v_location
- long_name :
- vertical fraction of open cell
Array Chunk Bytes 20.08 MiB 1.54 MiB Shape (50, 13, 90, 90) (50, 1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - maskC(k, face, j, i)booldask.array<chunksize=(50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- sea_binary_mask_at_t_location
- long_name :
- mask denoting wet point at center
Array Chunk Bytes 5.02 MiB 395.51 kiB Shape (50, 13, 90, 90) (50, 1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type bool numpy.ndarray - maskW(k, face, j, i_g)booldask.array<chunksize=(50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- cell_vertical_fraction_at_u_location
- long_name :
- mask denoting wet point at interface
Array Chunk Bytes 5.02 MiB 395.51 kiB Shape (50, 13, 90, 90) (50, 1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type bool numpy.ndarray - maskS(k, face, j_g, i)booldask.array<chunksize=(50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- cell_vertical_fraction_at_v_location
- long_name :
- mask denoting wet point at interface
Array Chunk Bytes 5.02 MiB 395.51 kiB Shape (50, 13, 90, 90) (50, 1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type bool numpy.ndarray - maskCtrlC(k, face, j, i)booldask.array<chunksize=(50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- ctrl_vector_3d_mask
- long_name :
- CTRL 3D mask where ctrl vector is active at tracer location
- units :
Array Chunk Bytes 5.02 MiB 395.51 kiB Shape (50, 13, 90, 90) (50, 1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type bool numpy.ndarray - maskCtrlS(k, face, j_g, i)booldask.array<chunksize=(50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- ctrl_vector_3d_mask_at_v_location
- long_name :
- CTRL 3D mask where ctrl vector is active at v location
- units :
Array Chunk Bytes 5.02 MiB 395.51 kiB Shape (50, 13, 90, 90) (50, 1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type bool numpy.ndarray - rhoRef(k)>f4dask.array<chunksize=(50,), meta=np.ndarray>
- standard_name :
- reference_density_profile
- long_name :
- 1D, vertical reference density profile
- coordinate :
- Z
- units :
- kg m-3
Array Chunk Bytes 200 B 200 B Shape (50,) (50,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - maskCtrlW(k, face, j, i_g)booldask.array<chunksize=(50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- ctrl_vector_3d_mask_at_u_location
- long_name :
- CTRL 3D mask where ctrl vector is active at u location
- units :
Array Chunk Bytes 5.02 MiB 395.51 kiB Shape (50, 13, 90, 90) (50, 1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type bool numpy.ndarray - iter(time)float64dask.array<chunksize=(120,), meta=np.ndarray>
- standard_name :
- timestep
- long_name :
- model timestep number
Array Chunk Bytes 1.38 kiB 0.94 kiB Shape (176,) (120,) Dask graph 2 chunks in 7 graph layers Data type float64 numpy.ndarray - drC_kl(k_l)>f4dask.array<chunksize=(50,), meta=np.ndarray>
- standard_name :
- cell_z_size_at_w_location
- long_name :
- cell z size
- units :
- m
Array Chunk Bytes 200 B 200 B Shape (50,) (50,) Dask graph 1 chunks in 2 graph layers Data type >f4 numpy.ndarray
- GM_CHIXO(time, k, face, j, i_g)float32dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_CHIXO
- long_name :
- Redi Temp variance dissipation rate: X component
- units :
- degC^2/s
- mate :
- GM_CHIX
Array Chunk Bytes 3.45 GiB 185.39 MiB Shape (176, 50, 13, 90, 90) (120, 50, 1, 90, 90) Dask graph 26 chunks in 18 graph layers Data type float32 numpy.ndarray - GM_CHIZ(time, k_l, face, j, i)float32dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_CHIZ
- long_name :
- Redi Temp variance dissipation rate: Z component
- units :
- degC^2/s
Array Chunk Bytes 3.45 GiB 185.39 MiB Shape (176, 50, 13, 90, 90) (120, 50, 1, 90, 90) Dask graph 26 chunks in 1 graph layer Data type float32 numpy.ndarray - GM_ubT(time, k, face, j, i_g)float32dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_ubT
- long_name :
- Zonal Mass-Weight Bolus Transp of Pot Temp
- units :
- degC.m^3/s
- mate :
- GM_vbT
Array Chunk Bytes 3.45 GiB 185.39 MiB Shape (176, 50, 13, 90, 90) (120, 50, 1, 90, 90) Dask graph 26 chunks in 1 graph layer Data type float32 numpy.ndarray - GM_CHIX(time, k, face, j, i_g)float32dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_CHIX
- long_name :
- Redi Temp variance dissipation rate: X component
- units :
- degC^2/s
- mate :
- GM_KwzTz
Array Chunk Bytes 3.45 GiB 185.39 MiB Shape (176, 50, 13, 90, 90) (120, 50, 1, 90, 90) Dask graph 26 chunks in 1 graph layer Data type float32 numpy.ndarray - GM_CHIYO(time, k, face, j_g, i)float32dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_CHIYO
- long_name :
- Redi Temp variance dissipation rate: Y component
- units :
- degC^2/s
- mate :
- GM_CHIY
Array Chunk Bytes 3.45 GiB 185.39 MiB Shape (176, 50, 13, 90, 90) (120, 50, 1, 90, 90) Dask graph 26 chunks in 18 graph layers Data type float32 numpy.ndarray - GM_Kvz(time, k, face, j_g, i)float32dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_Kvz
- long_name :
- K_23 element (V.point, Z.dir) of GM-Redi tensor
- units :
- m^2/s
- mate :
- GM_Kuz
Array Chunk Bytes 3.45 GiB 185.39 MiB Shape (176, 50, 13, 90, 90) (120, 50, 1, 90, 90) Dask graph 26 chunks in 18 graph layers Data type float32 numpy.ndarray - GM_Kwz(time, k_l, face, j, i)float32dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_Kwz
- long_name :
- K_33 element (W.point, Z.dir) of GM-Redi tensor
- units :
- m^2/s
Array Chunk Bytes 3.45 GiB 185.39 MiB Shape (176, 50, 13, 90, 90) (120, 50, 1, 90, 90) Dask graph 26 chunks in 1 graph layer Data type float32 numpy.ndarray - THETA(time, k, face, j, i)float32dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- THETA
- long_name :
- Potential Temperature
- units :
- degC
Array Chunk Bytes 3.45 GiB 185.39 MiB Shape (176, 50, 13, 90, 90) (120, 50, 1, 90, 90) Dask graph 26 chunks in 18 graph layers Data type float32 numpy.ndarray - GM_Kvy(time, k, face, j_g, i)float32dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_Kvy
- long_name :
- K_22 element (V.point, Y.dir) of GM-Redi tensor
- units :
- m^2/s
- mate :
- GM_Kux
Array Chunk Bytes 3.45 GiB 185.39 MiB Shape (176, 50, 13, 90, 90) (120, 50, 1, 90, 90) Dask graph 26 chunks in 1 graph layer Data type float32 numpy.ndarray - GM_Kwy(time, k_l, face, j, i)float32dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_Kwy
- long_name :
- K_32 element (W.point, Y.dir) of GM-Redi tensor
- units :
- m^2/s
- mate :
- GM_Kwx
Array Chunk Bytes 3.45 GiB 185.39 MiB Shape (176, 50, 13, 90, 90) (120, 50, 1, 90, 90) Dask graph 26 chunks in 18 graph layers Data type float32 numpy.ndarray - GM_KwzTz(time, k_l, face, j, i)float32dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_KwzTz
- long_name :
- Redi main-diagonal vertical Temperature flux
- units :
- degC.m^3/s
Array Chunk Bytes 3.45 GiB 185.39 MiB Shape (176, 50, 13, 90, 90) (120, 50, 1, 90, 90) Dask graph 26 chunks in 1 graph layer Data type float32 numpy.ndarray - GM_Kux(time, k, face, j, i_g)float32dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_Kux
- long_name :
- K_11 element (U.point, X.dir) of GM-Redi tensor
- units :
- m^2/s
- mate :
- GM_Kvy
Array Chunk Bytes 3.45 GiB 185.39 MiB Shape (176, 50, 13, 90, 90) (120, 50, 1, 90, 90) Dask graph 26 chunks in 1 graph layer Data type float32 numpy.ndarray - GM_Kwx(time, k_l, face, j, i)float32dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_Kwx
- long_name :
- K_31 element (W.point, X.dir) of GM-Redi tensor
- units :
- m^2/s
- mate :
- GM_Kwy
Array Chunk Bytes 3.45 GiB 185.39 MiB Shape (176, 50, 13, 90, 90) (120, 50, 1, 90, 90) Dask graph 26 chunks in 18 graph layers Data type float32 numpy.ndarray - GM_CHIY(time, k, face, j_g, i)float32dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_CHIY
- long_name :
- Redi Temp variance dissipation rate: Y component
- units :
- degC^2/s
- mate :
- GM_CHIXO
Array Chunk Bytes 3.45 GiB 185.39 MiB Shape (176, 50, 13, 90, 90) (120, 50, 1, 90, 90) Dask graph 26 chunks in 1 graph layer Data type float32 numpy.ndarray - GM_Kuz(time, k, face, j, i_g)float32dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_Kuz
- long_name :
- K_13 element (U.point, Z.dir) of GM-Redi tensor
- units :
- m^2/s
- mate :
- GM_Kvz
Array Chunk Bytes 3.45 GiB 185.39 MiB Shape (176, 50, 13, 90, 90) (120, 50, 1, 90, 90) Dask graph 26 chunks in 18 graph layers Data type float32 numpy.ndarray - Tx(time, k, face, j, i_g)float32dask.array<chunksize=(120, 50, 1, 90, 1), meta=np.ndarray>
Array Chunk Bytes 3.45 GiB 183.33 MiB Shape (176, 50, 13, 90, 90) (120, 50, 1, 90, 89) Dask graph 52 chunks in 27 graph layers Data type float32 numpy.ndarray - Ty(time, k, face, j_g, i)float32dask.array<chunksize=(120, 50, 1, 1, 90), meta=np.ndarray>
Array Chunk Bytes 3.45 GiB 183.33 MiB Shape (176, 50, 13, 90, 90) (120, 50, 1, 89, 90) Dask graph 52 chunks in 27 graph layers Data type float32 numpy.ndarray - Tz(time, k_l, face, j, i)float32dask.array<chunksize=(120, 1, 1, 90, 90), meta=np.ndarray>
Array Chunk Bytes 3.45 GiB 181.69 MiB Shape (176, 50, 13, 90, 90) (120, 49, 1, 90, 90) Dask graph 52 chunks in 30 graph layers Data type float32 numpy.ndarray - GM_CHI(time, k, face, j, i)float32dask.array<chunksize=(120, 49, 1, 89, 89), meta=np.ndarray>
Array Chunk Bytes 3.45 GiB 177.67 MiB Shape (176, 50, 13, 90, 90) (120, 49, 1, 89, 89) Dask graph 208 chunks in 50 graph layers Data type float32 numpy.ndarray
- iPandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89], dtype='int64', name='i')) - i_gPandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89], dtype='int64', name='i_g')) - jPandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89], dtype='int64', name='j')) - j_gPandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89], dtype='int64', name='j_g')) - kPandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49], dtype='int64', name='k')) - k_uPandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49], dtype='int64', name='k_u')) - k_lPandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49], dtype='int64', name='k_l')) - k_p1PandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50], dtype='int64', name='k_p1')) - facePandasIndex
PandasIndex(Int64Index([0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12], dtype='int64', name='face'))
- timePandasIndex
PandasIndex(TimedeltaIndex(['0 days 00:12:12', '0 days 00:23:48', '0 days 00:36:12', '0 days 00:48:12', '0 days 01:00:36', '0 days 01:12:36', '0 days 01:25:00', '0 days 01:37:24', '0 days 01:49:24', '0 days 02:01:48', ... '1 days 09:53:00', '1 days 10:05:24', '1 days 10:17:48', '1 days 10:29:00', '1 days 10:41:24', '1 days 10:53:24', '1 days 11:05:48', '1 days 11:17:48', '1 days 11:30:12', '1 days 11:42:36'], dtype='timedelta64[ns]', name='time', length=176, freq=None))
- Conventions :
- CF-1.6
- title :
- netCDF wrapper of MITgcm MDS binary data
- source :
- MITgcm
- history :
- Created by calling `open_mdsdataset(grid_dir='/glade/work/dcherian/mitgcm/ECCOV4/release4/run/diags/../', iters=None, prefix=None, read_grid=True, delta_t=1, ref_date=None, calendar='gregorian', geometry='llc', grid_vars_to_coords=False, swap_dims=False, endian='>', chunks={'face': 1, 'k_l': -1, 'k': -1}, ignore_unknown_vars=False, default_dtype=<class 'numpy.float64'>, nx=None, ny=None, nz=None, llc_method='smallchunks', extra_metadata=None, extra_variables=None, data_dir='/glade/work/dcherian/mitgcm/ECCOV4/release4/run/diags/GM_CHIXO', levels=None)`
<xarray.Dataset>
Dimensions: (i: 90, i_g: 90, j: 90, j_g: 90, k: 50, k_u: 50, k_l: 50,
k_p1: 51, face: 13, time: 176)
Coordinates: (12/45)
* i (i) int64 0 1 2 3 4 5 6 7 8 9 ... 80 81 82 83 84 85 86 87 88 89
* i_g (i_g) int64 0 1 2 3 4 5 6 7 8 9 ... 80 81 82 83 84 85 86 87 88 89
* j (j) int64 0 1 2 3 4 5 6 7 8 9 ... 80 81 82 83 84 85 86 87 88 89
* j_g (j_g) int64 0 1 2 3 4 5 6 7 8 9 ... 80 81 82 83 84 85 86 87 88 89
* k (k) int64 0 1 2 3 4 5 6 7 8 9 ... 40 41 42 43 44 45 46 47 48 49
* k_u (k_u) int64 0 1 2 3 4 5 6 7 8 9 ... 40 41 42 43 44 45 46 47 48 49
... ...
maskCtrlC (k, face, j, i) bool dask.array<chunksize=(50, 1, 90, 90), meta=np.ndarray>
maskCtrlS (k, face, j_g, i) bool dask.array<chunksize=(50, 1, 90, 90), meta=np.ndarray>
rhoRef (k) >f4 dask.array<chunksize=(50,), meta=np.ndarray>
maskCtrlW (k, face, j, i_g) bool dask.array<chunksize=(50, 1, 90, 90), meta=np.ndarray>
iter (time) float64 dask.array<chunksize=(120,), meta=np.ndarray>
drC_kl (k_l) >f4 dask.array<chunksize=(50,), meta=np.ndarray>
Data variables: (12/19)
GM_CHIXO (time, k, face, j, i_g) float32 dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
GM_CHIZ (time, k_l, face, j, i) float32 dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
GM_ubT (time, k, face, j, i_g) float32 dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
GM_CHIX (time, k, face, j, i_g) float32 dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
GM_CHIYO (time, k, face, j_g, i) float32 dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
GM_Kvz (time, k, face, j_g, i) float32 dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
... ...
GM_CHIY (time, k, face, j_g, i) float32 dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
GM_Kuz (time, k, face, j, i_g) float32 dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
Tx (time, k, face, j, i_g) float32 dask.array<chunksize=(120, 50, 1, 90, 1), meta=np.ndarray>
Ty (time, k, face, j_g, i) float32 dask.array<chunksize=(120, 50, 1, 1, 90), meta=np.ndarray>
Tz (time, k_l, face, j, i) float32 dask.array<chunksize=(120, 1, 1, 90, 90), meta=np.ndarray>
GM_CHI (time, k, face, j, i) float32 dask.array<chunksize=(120, 49, 1, 89, 89), meta=np.ndarray>
Attributes:
Conventions: CF-1.6
title: netCDF wrapper of MITgcm MDS binary data
source: MITgcm
history: Created by calling `open_mdsdataset(grid_dir='/glade/work/d...xarray.Dataset
- i: 90
- i_g: 90
- j: 90
- j_g: 90
- k: 50
- k_u: 50
- k_l: 50
- k_p1: 51
- face: 13
- time: 176
- i(i)int640 1 2 3 4 5 6 ... 84 85 86 87 88 89
- standard_name :
- x_grid_index
- axis :
- X
- long_name :
- x-dimension of the t grid
- swap_dim :
- XC
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89]) - i_g(i_g)int640 1 2 3 4 5 6 ... 84 85 86 87 88 89
- standard_name :
- x_grid_index_at_u_location
- axis :
- X
- long_name :
- x-dimension of the u grid
- c_grid_axis_shift :
- -0.5
- swap_dim :
- XG
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89]) - j(j)int640 1 2 3 4 5 6 ... 84 85 86 87 88 89
- standard_name :
- y_grid_index
- axis :
- Y
- long_name :
- y-dimension of the t grid
- swap_dim :
- YC
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89]) - j_g(j_g)int640 1 2 3 4 5 6 ... 84 85 86 87 88 89
- standard_name :
- y_grid_index_at_v_location
- axis :
- Y
- long_name :
- y-dimension of the v grid
- c_grid_axis_shift :
- -0.5
- swap_dim :
- YG
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89]) - k(k)int640 1 2 3 4 5 6 ... 44 45 46 47 48 49
- standard_name :
- z_grid_index
- axis :
- Z
- long_name :
- z-dimension of the t grid
- swap_dim :
- Z
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49]) - k_u(k_u)int640 1 2 3 4 5 6 ... 44 45 46 47 48 49
- standard_name :
- z_grid_index_at_upper_w_location
- axis :
- Z
- long_name :
- z-dimension of the w grid
- c_grid_axis_shift :
- 0.5
- swap_dim :
- Zu
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49]) - k_l(k_l)int640 1 2 3 4 5 6 ... 44 45 46 47 48 49
- standard_name :
- z_grid_index_at_lower_w_location
- axis :
- Z
- long_name :
- z-dimension of the w grid
- c_grid_axis_shift :
- -0.5
- swap_dim :
- Zl
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49]) - k_p1(k_p1)int640 1 2 3 4 5 6 ... 45 46 47 48 49 50
- standard_name :
- z_grid_index_at_w_location
- axis :
- Z
- long_name :
- z-dimension of the w grid
- c_grid_axis_shift :
- (-0.5, 0.5)
- swap_dim :
- Zp1
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50]) - face(face)int640 1 2 3 4 5 6 7 8 9 10 11 12
- standard_name :
- face_index
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12])
- time(time)timedelta64[ns]00:12:12 ... 1 days 11:42:36
- standard_name :
- time
- long_name :
- Time
- axis :
- T
- calendar :
- gregorian
array([ 732000000000, 1428000000000, 2172000000000, 2892000000000, 3636000000000, 4356000000000, 5100000000000, 5844000000000, 6564000000000, 7308000000000, 8028000000000, 8772000000000, 9516000000000, 10188000000000, 10932000000000, 11652000000000, 12396000000000, 13116000000000, 13860000000000, 14604000000000, 15324000000000, 16068000000000, 16788000000000, 17532000000000, 18276000000000, 18948000000000, 19692000000000, 20412000000000, 21156000000000, 21876000000000, 22620000000000, 23364000000000, 24084000000000, 24828000000000, 25548000000000, 26292000000000, 27036000000000, 27708000000000, 28452000000000, 29172000000000, 29916000000000, 30636000000000, 31380000000000, 32124000000000, 32844000000000, 33588000000000, 34308000000000, 35052000000000, 35796000000000, 36492000000000, 37236000000000, 37956000000000, 38700000000000, 39420000000000, 40164000000000, 40908000000000, 41628000000000, 42372000000000, 43092000000000, 43836000000000, 44580000000000, 45252000000000, 45996000000000, 46716000000000, 47460000000000, 48180000000000, 48924000000000, 49668000000000, 50388000000000, 51132000000000, 51852000000000, 52596000000000, 53340000000000, 54012000000000, 54756000000000, 55476000000000, 56220000000000, 56940000000000, 57684000000000, 58428000000000, 59148000000000, 59892000000000, 60612000000000, 61356000000000, 62100000000000, 62772000000000, 63516000000000, 64236000000000, 64980000000000, 65700000000000, 66444000000000, 67188000000000, 67908000000000, 68652000000000, 69372000000000, 70116000000000, 70860000000000, 71556000000000, 72300000000000, 73020000000000, 73764000000000, 74484000000000, 75228000000000, 75972000000000, 76692000000000, 77436000000000, 78156000000000, 78900000000000, 79644000000000, 80316000000000, 81060000000000, 81780000000000, 82524000000000, 83244000000000, 83988000000000, 84732000000000, 85452000000000, 86196000000000, 86916000000000, 87660000000000, 88404000000000, 89076000000000, 89820000000000, 90540000000000, 91284000000000, 92004000000000, 92748000000000, 93492000000000, 94212000000000, 94956000000000, 95676000000000, 96420000000000, 97164000000000, 97836000000000, 98580000000000, 99300000000000, 100044000000000, 100764000000000, 101508000000000, 102252000000000, 102972000000000, 103716000000000, 104436000000000, 105180000000000, 105924000000000, 106620000000000, 107364000000000, 108084000000000, 108828000000000, 109548000000000, 110292000000000, 111036000000000, 111756000000000, 112500000000000, 113220000000000, 113964000000000, 114708000000000, 115380000000000, 116124000000000, 116844000000000, 117588000000000, 118308000000000, 119052000000000, 119796000000000, 120516000000000, 121260000000000, 121980000000000, 122724000000000, 123468000000000, 124140000000000, 124884000000000, 125604000000000, 126348000000000, 127068000000000, 127812000000000, 128556000000000], dtype='timedelta64[ns]') - XC(face, j, i)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- longitude
- long_name :
- longitude
- units :
- degrees_east
- coordinate :
- YC XC
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - YC(face, j, i)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- latitude
- long_name :
- latitude
- units :
- degrees_north
- coordinate :
- YC XC
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - XG(face, j_g, i_g)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- longitude_at_f_location
- long_name :
- longitude
- units :
- degrees_east
- coordinate :
- YG XG
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - YG(face, j_g, i_g)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- latitude_at_f_location
- long_name :
- latitude
- units :
- degrees_north
- coordinate :
- YG XG
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - CS(face, j, i)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- Cos of grid orientation angle
- long_name :
- AngleCS
- units :
- coordinate :
- YC XC
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - SN(face, j, i)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- Sin of grid orientation angle
- long_name :
- AngleSN
- units :
- coordinate :
- YC XC
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - Z(k)>f4dask.array<chunksize=(50,), meta=np.ndarray>
- standard_name :
- depth
- long_name :
- vertical coordinate of cell center
- units :
- m
- positive :
- down
Array Chunk Bytes 200 B 200 B Shape (50,) (50,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - Zp1(k_p1)>f4dask.array<chunksize=(51,), meta=np.ndarray>
- standard_name :
- depth_at_w_location
- long_name :
- vertical coordinate of cell interface
- units :
- m
- positive :
- down
Array Chunk Bytes 204 B 204 B Shape (51,) (51,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - Zu(k_u)>f4dask.array<chunksize=(50,), meta=np.ndarray>
- standard_name :
- depth_at_upper_w_location
- long_name :
- vertical coordinate of upper cell interface
- units :
- m
- positive :
- down
Array Chunk Bytes 200 B 200 B Shape (50,) (50,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - Zl(k_l)>f4dask.array<chunksize=(50,), meta=np.ndarray>
- standard_name :
- depth_at_lower_w_location
- long_name :
- vertical coordinate of lower cell interface
- units :
- m
- positive :
- down
Array Chunk Bytes 200 B 200 B Shape (50,) (50,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - rA(face, j, i)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- cell_area
- long_name :
- cell area
- units :
- m2
- coordinate :
- YC XC
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - dxG(face, j_g, i)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- cell_x_size_at_v_location
- long_name :
- cell x size
- units :
- m
- coordinate :
- YG XC
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - dyG(face, j, i_g)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- cell_y_size_at_u_location
- long_name :
- cell y size
- units :
- m
- coordinate :
- YC XG
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - Depth(face, j, i)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- ocean_depth
- long_name :
- ocean depth
- units :
- m
- coordinate :
- XC YC
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - rAz(face, j_g, i_g)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- cell_area_at_f_location
- long_name :
- cell area
- units :
- m
- coordinate :
- YG XG
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - dxC(face, j, i_g)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- cell_x_size_at_u_location
- long_name :
- cell x size
- units :
- m
- coordinate :
- YC XG
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - dyC(face, j_g, i)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- cell_y_size_at_v_location
- long_name :
- cell y size
- units :
- m
- coordinate :
- YG XC
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - rAw(face, j, i_g)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- cell_area_at_u_location
- long_name :
- cell area
- units :
- m2
- coordinate :
- YG XC
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - rAs(face, j_g, i)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- cell_area_at_v_location
- long_name :
- cell area
- units :
- m2
- coordinate :
- YG XC
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - drC(k_p1)>f4dask.array<chunksize=(51,), meta=np.ndarray>
- standard_name :
- cell_z_size_at_w_location
- long_name :
- cell z size
- units :
- m
Array Chunk Bytes 204 B 204 B Shape (51,) (51,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - drF(k)>f4dask.array<chunksize=(50,), meta=np.ndarray>
- standard_name :
- cell_z_size
- long_name :
- cell z size
- units :
- m
Array Chunk Bytes 200 B 200 B Shape (50,) (50,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - PHrefC(k)>f4dask.array<chunksize=(50,), meta=np.ndarray>
- standard_name :
- cell_reference_pressure
- long_name :
- Reference Hydrostatic Pressure
- units :
- m2 s-2
Array Chunk Bytes 200 B 200 B Shape (50,) (50,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - PHrefF(k_p1)>f4dask.array<chunksize=(51,), meta=np.ndarray>
- standard_name :
- cell_reference_pressure
- long_name :
- Reference Hydrostatic Pressure
- units :
- m2 s-2
Array Chunk Bytes 204 B 204 B Shape (51,) (51,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - hFacC(k, face, j, i)>f4dask.array<chunksize=(50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- cell_vertical_fraction
- long_name :
- vertical fraction of open cell
Array Chunk Bytes 20.08 MiB 1.54 MiB Shape (50, 13, 90, 90) (50, 1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - hFacW(k, face, j, i_g)>f4dask.array<chunksize=(50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- cell_vertical_fraction_at_u_location
- long_name :
- vertical fraction of open cell
Array Chunk Bytes 20.08 MiB 1.54 MiB Shape (50, 13, 90, 90) (50, 1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - hFacS(k, face, j_g, i)>f4dask.array<chunksize=(50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- cell_vertical_fraction_at_v_location
- long_name :
- vertical fraction of open cell
Array Chunk Bytes 20.08 MiB 1.54 MiB Shape (50, 13, 90, 90) (50, 1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - maskC(k, face, j, i)booldask.array<chunksize=(50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- sea_binary_mask_at_t_location
- long_name :
- mask denoting wet point at center
Array Chunk Bytes 5.02 MiB 395.51 kiB Shape (50, 13, 90, 90) (50, 1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type bool numpy.ndarray - maskW(k, face, j, i_g)booldask.array<chunksize=(50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- cell_vertical_fraction_at_u_location
- long_name :
- mask denoting wet point at interface
Array Chunk Bytes 5.02 MiB 395.51 kiB Shape (50, 13, 90, 90) (50, 1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type bool numpy.ndarray - maskS(k, face, j_g, i)booldask.array<chunksize=(50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- cell_vertical_fraction_at_v_location
- long_name :
- mask denoting wet point at interface
Array Chunk Bytes 5.02 MiB 395.51 kiB Shape (50, 13, 90, 90) (50, 1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type bool numpy.ndarray - maskCtrlC(k, face, j, i)booldask.array<chunksize=(50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- ctrl_vector_3d_mask
- long_name :
- CTRL 3D mask where ctrl vector is active at tracer location
- units :
Array Chunk Bytes 5.02 MiB 395.51 kiB Shape (50, 13, 90, 90) (50, 1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type bool numpy.ndarray - maskCtrlS(k, face, j_g, i)booldask.array<chunksize=(50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- ctrl_vector_3d_mask_at_v_location
- long_name :
- CTRL 3D mask where ctrl vector is active at v location
- units :
Array Chunk Bytes 5.02 MiB 395.51 kiB Shape (50, 13, 90, 90) (50, 1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type bool numpy.ndarray - rhoRef(k)>f4dask.array<chunksize=(50,), meta=np.ndarray>
- standard_name :
- reference_density_profile
- long_name :
- 1D, vertical reference density profile
- coordinate :
- Z
- units :
- kg m-3
Array Chunk Bytes 200 B 200 B Shape (50,) (50,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - maskCtrlW(k, face, j, i_g)booldask.array<chunksize=(50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- ctrl_vector_3d_mask_at_u_location
- long_name :
- CTRL 3D mask where ctrl vector is active at u location
- units :
Array Chunk Bytes 5.02 MiB 395.51 kiB Shape (50, 13, 90, 90) (50, 1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type bool numpy.ndarray - iter(time)float64dask.array<chunksize=(120,), meta=np.ndarray>
- standard_name :
- timestep
- long_name :
- model timestep number
Array Chunk Bytes 1.38 kiB 0.94 kiB Shape (176,) (120,) Dask graph 2 chunks in 7 graph layers Data type float64 numpy.ndarray - drC_kl(k_l)>f4dask.array<chunksize=(50,), meta=np.ndarray>
- standard_name :
- cell_z_size_at_w_location
- long_name :
- cell z size
- units :
- m
Array Chunk Bytes 200 B 200 B Shape (50,) (50,) Dask graph 1 chunks in 2 graph layers Data type >f4 numpy.ndarray
- GM_CHIXO(time, k, face, j, i_g)float32dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_CHIXO
- long_name :
- Redi Temp variance dissipation rate: X component
- units :
- degC^2/s
- mate :
- GM_CHIX
Array Chunk Bytes 3.45 GiB 185.39 MiB Shape (176, 50, 13, 90, 90) (120, 50, 1, 90, 90) Dask graph 26 chunks in 18 graph layers Data type float32 numpy.ndarray - GM_CHIZ(time, k_l, face, j, i)float32dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_CHIZ
- long_name :
- Redi Temp variance dissipation rate: Z component
- units :
- degC^2/s
Array Chunk Bytes 3.45 GiB 185.39 MiB Shape (176, 50, 13, 90, 90) (120, 50, 1, 90, 90) Dask graph 26 chunks in 1 graph layer Data type float32 numpy.ndarray - GM_ubT(time, k, face, j, i_g)float32dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_ubT
- long_name :
- Zonal Mass-Weight Bolus Transp of Pot Temp
- units :
- degC.m^3/s
- mate :
- GM_vbT
Array Chunk Bytes 3.45 GiB 185.39 MiB Shape (176, 50, 13, 90, 90) (120, 50, 1, 90, 90) Dask graph 26 chunks in 1 graph layer Data type float32 numpy.ndarray - GM_CHIX(time, k, face, j, i_g)float32dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_CHIX
- long_name :
- Redi Temp variance dissipation rate: X component
- units :
- degC^2/s
- mate :
- GM_KwzTz
Array Chunk Bytes 3.45 GiB 185.39 MiB Shape (176, 50, 13, 90, 90) (120, 50, 1, 90, 90) Dask graph 26 chunks in 1 graph layer Data type float32 numpy.ndarray - GM_CHIYO(time, k, face, j_g, i)float32dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_CHIYO
- long_name :
- Redi Temp variance dissipation rate: Y component
- units :
- degC^2/s
- mate :
- GM_CHIY
Array Chunk Bytes 3.45 GiB 185.39 MiB Shape (176, 50, 13, 90, 90) (120, 50, 1, 90, 90) Dask graph 26 chunks in 18 graph layers Data type float32 numpy.ndarray - GM_Kvz(time, k, face, j_g, i)float32dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_Kvz
- long_name :
- K_23 element (V.point, Z.dir) of GM-Redi tensor
- units :
- m^2/s
- mate :
- GM_Kuz
Array Chunk Bytes 3.45 GiB 185.39 MiB Shape (176, 50, 13, 90, 90) (120, 50, 1, 90, 90) Dask graph 26 chunks in 18 graph layers Data type float32 numpy.ndarray - GM_Kwz(time, k_l, face, j, i)float64dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_Kwz
- long_name :
- K_33 element (W.point, Z.dir) of GM-Redi tensor
- units :
- m^2/s
Array Chunk Bytes 6.90 GiB 370.79 MiB Shape (176, 50, 13, 90, 90) (120, 50, 1, 90, 90) Dask graph 26 chunks in 1 graph layer Data type float64 numpy.ndarray - THETA(time, k, face, j, i)float32dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- THETA
- long_name :
- Potential Temperature
- units :
- degC
Array Chunk Bytes 3.45 GiB 185.39 MiB Shape (176, 50, 13, 90, 90) (120, 50, 1, 90, 90) Dask graph 26 chunks in 18 graph layers Data type float32 numpy.ndarray - GM_Kvy(time, k, face, j_g, i)float32dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_Kvy
- long_name :
- K_22 element (V.point, Y.dir) of GM-Redi tensor
- units :
- m^2/s
- mate :
- GM_Kux
Array Chunk Bytes 3.45 GiB 185.39 MiB Shape (176, 50, 13, 90, 90) (120, 50, 1, 90, 90) Dask graph 26 chunks in 1 graph layer Data type float32 numpy.ndarray - GM_Kwy(time, k_l, face, j, i)float32dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_Kwy
- long_name :
- K_32 element (W.point, Y.dir) of GM-Redi tensor
- units :
- m^2/s
- mate :
- GM_Kwx
Array Chunk Bytes 3.45 GiB 185.39 MiB Shape (176, 50, 13, 90, 90) (120, 50, 1, 90, 90) Dask graph 26 chunks in 18 graph layers Data type float32 numpy.ndarray - GM_KwzTz(time, k_l, face, j, i)float32dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_KwzTz
- long_name :
- Redi main-diagonal vertical Temperature flux
- units :
- degC.m^3/s
Array Chunk Bytes 3.45 GiB 185.39 MiB Shape (176, 50, 13, 90, 90) (120, 50, 1, 90, 90) Dask graph 26 chunks in 1 graph layer Data type float32 numpy.ndarray - GM_Kux(time, k, face, j, i_g)float32dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_Kux
- long_name :
- K_11 element (U.point, X.dir) of GM-Redi tensor
- units :
- m^2/s
- mate :
- GM_Kvy
Array Chunk Bytes 3.45 GiB 185.39 MiB Shape (176, 50, 13, 90, 90) (120, 50, 1, 90, 90) Dask graph 26 chunks in 1 graph layer Data type float32 numpy.ndarray - GM_Kwx(time, k_l, face, j, i)float32dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_Kwx
- long_name :
- K_31 element (W.point, X.dir) of GM-Redi tensor
- units :
- m^2/s
- mate :
- GM_Kwy
Array Chunk Bytes 3.45 GiB 185.39 MiB Shape (176, 50, 13, 90, 90) (120, 50, 1, 90, 90) Dask graph 26 chunks in 18 graph layers Data type float32 numpy.ndarray - GM_CHIY(time, k, face, j_g, i)float32dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_CHIY
- long_name :
- Redi Temp variance dissipation rate: Y component
- units :
- degC^2/s
- mate :
- GM_CHIXO
Array Chunk Bytes 3.45 GiB 185.39 MiB Shape (176, 50, 13, 90, 90) (120, 50, 1, 90, 90) Dask graph 26 chunks in 1 graph layer Data type float32 numpy.ndarray - GM_Kuz(time, k, face, j, i_g)float32dask.array<chunksize=(120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_Kuz
- long_name :
- K_13 element (U.point, Z.dir) of GM-Redi tensor
- units :
- m^2/s
- mate :
- GM_Kvz
Array Chunk Bytes 3.45 GiB 185.39 MiB Shape (176, 50, 13, 90, 90) (120, 50, 1, 90, 90) Dask graph 26 chunks in 18 graph layers Data type float32 numpy.ndarray - Tx(time, k, face, j, i_g)float32dask.array<chunksize=(120, 50, 1, 90, 1), meta=np.ndarray>
Array Chunk Bytes 3.45 GiB 183.33 MiB Shape (176, 50, 13, 90, 90) (120, 50, 1, 90, 89) Dask graph 52 chunks in 27 graph layers Data type float32 numpy.ndarray - Ty(time, k, face, j_g, i)float32dask.array<chunksize=(120, 50, 1, 1, 90), meta=np.ndarray>
Array Chunk Bytes 3.45 GiB 183.33 MiB Shape (176, 50, 13, 90, 90) (120, 50, 1, 89, 90) Dask graph 52 chunks in 27 graph layers Data type float32 numpy.ndarray - Tz(time, k_l, face, j, i)float32dask.array<chunksize=(120, 1, 1, 90, 90), meta=np.ndarray>
Array Chunk Bytes 3.45 GiB 181.69 MiB Shape (176, 50, 13, 90, 90) (120, 49, 1, 90, 90) Dask graph 52 chunks in 30 graph layers Data type float32 numpy.ndarray - GM_CHI(time, k, face, j, i)float32dask.array<chunksize=(120, 49, 1, 89, 89), meta=np.ndarray>
Array Chunk Bytes 3.45 GiB 177.67 MiB Shape (176, 50, 13, 90, 90) (120, 49, 1, 89, 89) Dask graph 208 chunks in 50 graph layers Data type float32 numpy.ndarray
- iPandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89], dtype='int64', name='i')) - i_gPandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89], dtype='int64', name='i_g')) - jPandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89], dtype='int64', name='j')) - j_gPandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89], dtype='int64', name='j_g')) - kPandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49], dtype='int64', name='k')) - k_uPandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49], dtype='int64', name='k_u')) - k_lPandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49], dtype='int64', name='k_l')) - k_p1PandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50], dtype='int64', name='k_p1')) - facePandasIndex
PandasIndex(Int64Index([0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12], dtype='int64', name='face'))
- timePandasIndex
PandasIndex(TimedeltaIndex(['0 days 00:12:12', '0 days 00:23:48', '0 days 00:36:12', '0 days 00:48:12', '0 days 01:00:36', '0 days 01:12:36', '0 days 01:25:00', '0 days 01:37:24', '0 days 01:49:24', '0 days 02:01:48', ... '1 days 09:53:00', '1 days 10:05:24', '1 days 10:17:48', '1 days 10:29:00', '1 days 10:41:24', '1 days 10:53:24', '1 days 11:05:48', '1 days 11:17:48', '1 days 11:30:12', '1 days 11:42:36'], dtype='timedelta64[ns]', name='time', length=176, freq=None))
- Conventions :
- CF-1.6
- title :
- netCDF wrapper of MITgcm MDS binary data
- source :
- MITgcm
- history :
- Created by calling `open_mdsdataset(grid_dir='/glade/work/dcherian/mitgcm/ECCOV4/release4/run2/diags/../', iters=None, prefix=None, read_grid=True, delta_t=1, ref_date=None, calendar='gregorian', geometry='llc', grid_vars_to_coords=False, swap_dims=False, endian='>', chunks={'face': 1, 'k_l': -1, 'k': -1}, ignore_unknown_vars=False, default_dtype=<class 'numpy.float64'>, nx=None, ny=None, nz=None, llc_method='smallchunks', extra_metadata=None, extra_variables=None, data_dir='/glade/work/dcherian/mitgcm/ECCOV4/release4/run2/diags/GM_CHIXO', levels=None)`
concatted = xr.concat(
[adj, const], dim="node", compat="override", coords="minimal"
).assign_coords(node=["adj", "const"])
<xarray.Dataset>
Dimensions: (i: 90, i_g: 90, j: 90, j_g: 90, k: 50, k_u: 50, k_l: 50,
k_p1: 51, face: 13, time: 176, node: 2)
Coordinates: (12/46)
* i (i) int64 0 1 2 3 4 5 6 7 8 9 ... 80 81 82 83 84 85 86 87 88 89
* i_g (i_g) int64 0 1 2 3 4 5 6 7 8 9 ... 80 81 82 83 84 85 86 87 88 89
* j (j) int64 0 1 2 3 4 5 6 7 8 9 ... 80 81 82 83 84 85 86 87 88 89
* j_g (j_g) int64 0 1 2 3 4 5 6 7 8 9 ... 80 81 82 83 84 85 86 87 88 89
* k (k) int64 0 1 2 3 4 5 6 7 8 9 ... 40 41 42 43 44 45 46 47 48 49
* k_u (k_u) int64 0 1 2 3 4 5 6 7 8 9 ... 40 41 42 43 44 45 46 47 48 49
... ...
maskCtrlS (k, face, j_g, i) bool dask.array<chunksize=(50, 1, 90, 90), meta=np.ndarray>
rhoRef (k) >f4 dask.array<chunksize=(50,), meta=np.ndarray>
maskCtrlW (k, face, j, i_g) bool dask.array<chunksize=(50, 1, 90, 90), meta=np.ndarray>
iter (time) float64 dask.array<chunksize=(120,), meta=np.ndarray>
drC_kl (k_l) >f4 dask.array<chunksize=(50,), meta=np.ndarray>
* node (node) <U5 'adj' 'const'
Data variables: (12/19)
GM_CHIXO (node, time, k, face, j, i_g) float32 dask.array<chunksize=(1, 120, 50, 1, 90, 90), meta=np.ndarray>
GM_CHIZ (node, time, k_l, face, j, i) float32 dask.array<chunksize=(1, 120, 50, 1, 90, 90), meta=np.ndarray>
GM_ubT (node, time, k, face, j, i_g) float32 dask.array<chunksize=(1, 120, 50, 1, 90, 90), meta=np.ndarray>
GM_CHIX (node, time, k, face, j, i_g) float32 dask.array<chunksize=(1, 120, 50, 1, 90, 90), meta=np.ndarray>
GM_CHIYO (node, time, k, face, j_g, i) float32 dask.array<chunksize=(1, 120, 50, 1, 90, 90), meta=np.ndarray>
GM_Kvz (node, time, k, face, j_g, i) float32 dask.array<chunksize=(1, 120, 50, 1, 90, 90), meta=np.ndarray>
... ...
GM_CHIY (node, time, k, face, j_g, i) float32 dask.array<chunksize=(1, 120, 50, 1, 90, 90), meta=np.ndarray>
GM_Kuz (node, time, k, face, j, i_g) float32 dask.array<chunksize=(1, 120, 50, 1, 90, 90), meta=np.ndarray>
Tx (node, time, k, face, j, i_g) float32 dask.array<chunksize=(1, 120, 50, 1, 90, 1), meta=np.ndarray>
Ty (node, time, k, face, j_g, i) float32 dask.array<chunksize=(1, 120, 50, 1, 1, 90), meta=np.ndarray>
Tz (node, time, k_l, face, j, i) float32 dask.array<chunksize=(1, 120, 1, 1, 90, 90), meta=np.ndarray>
GM_CHI (node, time, k, face, j, i) float32 dask.array<chunksize=(1, 120, 49, 1, 89, 89), meta=np.ndarray>
Attributes:
Conventions: CF-1.6
title: netCDF wrapper of MITgcm MDS binary data
source: MITgcm
history: Created by calling `open_mdsdataset(grid_dir='/glade/work/d...xarray.Dataset
- i: 90
- i_g: 90
- j: 90
- j_g: 90
- k: 50
- k_u: 50
- k_l: 50
- k_p1: 51
- face: 13
- time: 176
- node: 2
- i(i)int640 1 2 3 4 5 6 ... 84 85 86 87 88 89
- standard_name :
- x_grid_index
- axis :
- X
- long_name :
- x-dimension of the t grid
- swap_dim :
- XC
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89]) - i_g(i_g)int640 1 2 3 4 5 6 ... 84 85 86 87 88 89
- standard_name :
- x_grid_index_at_u_location
- axis :
- X
- long_name :
- x-dimension of the u grid
- c_grid_axis_shift :
- -0.5
- swap_dim :
- XG
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89]) - j(j)int640 1 2 3 4 5 6 ... 84 85 86 87 88 89
- standard_name :
- y_grid_index
- axis :
- Y
- long_name :
- y-dimension of the t grid
- swap_dim :
- YC
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89]) - j_g(j_g)int640 1 2 3 4 5 6 ... 84 85 86 87 88 89
- standard_name :
- y_grid_index_at_v_location
- axis :
- Y
- long_name :
- y-dimension of the v grid
- c_grid_axis_shift :
- -0.5
- swap_dim :
- YG
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89]) - k(k)int640 1 2 3 4 5 6 ... 44 45 46 47 48 49
- standard_name :
- z_grid_index
- axis :
- Z
- long_name :
- z-dimension of the t grid
- swap_dim :
- Z
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49]) - k_u(k_u)int640 1 2 3 4 5 6 ... 44 45 46 47 48 49
- standard_name :
- z_grid_index_at_upper_w_location
- axis :
- Z
- long_name :
- z-dimension of the w grid
- c_grid_axis_shift :
- 0.5
- swap_dim :
- Zu
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49]) - k_l(k_l)int640 1 2 3 4 5 6 ... 44 45 46 47 48 49
- standard_name :
- z_grid_index_at_lower_w_location
- axis :
- Z
- long_name :
- z-dimension of the w grid
- c_grid_axis_shift :
- -0.5
- swap_dim :
- Zl
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49]) - k_p1(k_p1)int640 1 2 3 4 5 6 ... 45 46 47 48 49 50
- standard_name :
- z_grid_index_at_w_location
- axis :
- Z
- long_name :
- z-dimension of the w grid
- c_grid_axis_shift :
- (-0.5, 0.5)
- swap_dim :
- Zp1
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50]) - face(face)int640 1 2 3 4 5 6 7 8 9 10 11 12
- standard_name :
- face_index
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12])
- time(time)timedelta64[ns]00:12:12 ... 1 days 11:42:36
- standard_name :
- time
- long_name :
- Time
- axis :
- T
- calendar :
- gregorian
array([ 732000000000, 1428000000000, 2172000000000, 2892000000000, 3636000000000, 4356000000000, 5100000000000, 5844000000000, 6564000000000, 7308000000000, 8028000000000, 8772000000000, 9516000000000, 10188000000000, 10932000000000, 11652000000000, 12396000000000, 13116000000000, 13860000000000, 14604000000000, 15324000000000, 16068000000000, 16788000000000, 17532000000000, 18276000000000, 18948000000000, 19692000000000, 20412000000000, 21156000000000, 21876000000000, 22620000000000, 23364000000000, 24084000000000, 24828000000000, 25548000000000, 26292000000000, 27036000000000, 27708000000000, 28452000000000, 29172000000000, 29916000000000, 30636000000000, 31380000000000, 32124000000000, 32844000000000, 33588000000000, 34308000000000, 35052000000000, 35796000000000, 36492000000000, 37236000000000, 37956000000000, 38700000000000, 39420000000000, 40164000000000, 40908000000000, 41628000000000, 42372000000000, 43092000000000, 43836000000000, 44580000000000, 45252000000000, 45996000000000, 46716000000000, 47460000000000, 48180000000000, 48924000000000, 49668000000000, 50388000000000, 51132000000000, 51852000000000, 52596000000000, 53340000000000, 54012000000000, 54756000000000, 55476000000000, 56220000000000, 56940000000000, 57684000000000, 58428000000000, 59148000000000, 59892000000000, 60612000000000, 61356000000000, 62100000000000, 62772000000000, 63516000000000, 64236000000000, 64980000000000, 65700000000000, 66444000000000, 67188000000000, 67908000000000, 68652000000000, 69372000000000, 70116000000000, 70860000000000, 71556000000000, 72300000000000, 73020000000000, 73764000000000, 74484000000000, 75228000000000, 75972000000000, 76692000000000, 77436000000000, 78156000000000, 78900000000000, 79644000000000, 80316000000000, 81060000000000, 81780000000000, 82524000000000, 83244000000000, 83988000000000, 84732000000000, 85452000000000, 86196000000000, 86916000000000, 87660000000000, 88404000000000, 89076000000000, 89820000000000, 90540000000000, 91284000000000, 92004000000000, 92748000000000, 93492000000000, 94212000000000, 94956000000000, 95676000000000, 96420000000000, 97164000000000, 97836000000000, 98580000000000, 99300000000000, 100044000000000, 100764000000000, 101508000000000, 102252000000000, 102972000000000, 103716000000000, 104436000000000, 105180000000000, 105924000000000, 106620000000000, 107364000000000, 108084000000000, 108828000000000, 109548000000000, 110292000000000, 111036000000000, 111756000000000, 112500000000000, 113220000000000, 113964000000000, 114708000000000, 115380000000000, 116124000000000, 116844000000000, 117588000000000, 118308000000000, 119052000000000, 119796000000000, 120516000000000, 121260000000000, 121980000000000, 122724000000000, 123468000000000, 124140000000000, 124884000000000, 125604000000000, 126348000000000, 127068000000000, 127812000000000, 128556000000000], dtype='timedelta64[ns]') - XC(face, j, i)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- longitude
- long_name :
- longitude
- units :
- degrees_east
- coordinate :
- YC XC
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - YC(face, j, i)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- latitude
- long_name :
- latitude
- units :
- degrees_north
- coordinate :
- YC XC
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - XG(face, j_g, i_g)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- longitude_at_f_location
- long_name :
- longitude
- units :
- degrees_east
- coordinate :
- YG XG
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - YG(face, j_g, i_g)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- latitude_at_f_location
- long_name :
- latitude
- units :
- degrees_north
- coordinate :
- YG XG
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - CS(face, j, i)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- Cos of grid orientation angle
- long_name :
- AngleCS
- units :
- coordinate :
- YC XC
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - SN(face, j, i)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- Sin of grid orientation angle
- long_name :
- AngleSN
- units :
- coordinate :
- YC XC
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - Z(k)>f4dask.array<chunksize=(50,), meta=np.ndarray>
- standard_name :
- depth
- long_name :
- vertical coordinate of cell center
- units :
- m
- positive :
- down
Array Chunk Bytes 200 B 200 B Shape (50,) (50,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - Zp1(k_p1)>f4dask.array<chunksize=(51,), meta=np.ndarray>
- standard_name :
- depth_at_w_location
- long_name :
- vertical coordinate of cell interface
- units :
- m
- positive :
- down
Array Chunk Bytes 204 B 204 B Shape (51,) (51,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - Zu(k_u)>f4dask.array<chunksize=(50,), meta=np.ndarray>
- standard_name :
- depth_at_upper_w_location
- long_name :
- vertical coordinate of upper cell interface
- units :
- m
- positive :
- down
Array Chunk Bytes 200 B 200 B Shape (50,) (50,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - Zl(k_l)>f4dask.array<chunksize=(50,), meta=np.ndarray>
- standard_name :
- depth_at_lower_w_location
- long_name :
- vertical coordinate of lower cell interface
- units :
- m
- positive :
- down
Array Chunk Bytes 200 B 200 B Shape (50,) (50,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - rA(face, j, i)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- cell_area
- long_name :
- cell area
- units :
- m2
- coordinate :
- YC XC
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - dxG(face, j_g, i)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- cell_x_size_at_v_location
- long_name :
- cell x size
- units :
- m
- coordinate :
- YG XC
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - dyG(face, j, i_g)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- cell_y_size_at_u_location
- long_name :
- cell y size
- units :
- m
- coordinate :
- YC XG
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - Depth(face, j, i)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- ocean_depth
- long_name :
- ocean depth
- units :
- m
- coordinate :
- XC YC
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - rAz(face, j_g, i_g)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- cell_area_at_f_location
- long_name :
- cell area
- units :
- m
- coordinate :
- YG XG
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - dxC(face, j, i_g)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- cell_x_size_at_u_location
- long_name :
- cell x size
- units :
- m
- coordinate :
- YC XG
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - dyC(face, j_g, i)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- cell_y_size_at_v_location
- long_name :
- cell y size
- units :
- m
- coordinate :
- YG XC
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - rAw(face, j, i_g)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- cell_area_at_u_location
- long_name :
- cell area
- units :
- m2
- coordinate :
- YG XC
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - rAs(face, j_g, i)>f4dask.array<chunksize=(1, 90, 90), meta=np.ndarray>
- standard_name :
- cell_area_at_v_location
- long_name :
- cell area
- units :
- m2
- coordinate :
- YG XC
Array Chunk Bytes 411.33 kiB 31.64 kiB Shape (13, 90, 90) (1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - drC(k_p1)>f4dask.array<chunksize=(51,), meta=np.ndarray>
- standard_name :
- cell_z_size_at_w_location
- long_name :
- cell z size
- units :
- m
Array Chunk Bytes 204 B 204 B Shape (51,) (51,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - drF(k)>f4dask.array<chunksize=(50,), meta=np.ndarray>
- standard_name :
- cell_z_size
- long_name :
- cell z size
- units :
- m
Array Chunk Bytes 200 B 200 B Shape (50,) (50,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - PHrefC(k)>f4dask.array<chunksize=(50,), meta=np.ndarray>
- standard_name :
- cell_reference_pressure
- long_name :
- Reference Hydrostatic Pressure
- units :
- m2 s-2
Array Chunk Bytes 200 B 200 B Shape (50,) (50,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - PHrefF(k_p1)>f4dask.array<chunksize=(51,), meta=np.ndarray>
- standard_name :
- cell_reference_pressure
- long_name :
- Reference Hydrostatic Pressure
- units :
- m2 s-2
Array Chunk Bytes 204 B 204 B Shape (51,) (51,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - hFacC(k, face, j, i)>f4dask.array<chunksize=(50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- cell_vertical_fraction
- long_name :
- vertical fraction of open cell
Array Chunk Bytes 20.08 MiB 1.54 MiB Shape (50, 13, 90, 90) (50, 1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - hFacW(k, face, j, i_g)>f4dask.array<chunksize=(50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- cell_vertical_fraction_at_u_location
- long_name :
- vertical fraction of open cell
Array Chunk Bytes 20.08 MiB 1.54 MiB Shape (50, 13, 90, 90) (50, 1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - hFacS(k, face, j_g, i)>f4dask.array<chunksize=(50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- cell_vertical_fraction_at_v_location
- long_name :
- vertical fraction of open cell
Array Chunk Bytes 20.08 MiB 1.54 MiB Shape (50, 13, 90, 90) (50, 1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type >f4 numpy.ndarray - maskC(k, face, j, i)booldask.array<chunksize=(50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- sea_binary_mask_at_t_location
- long_name :
- mask denoting wet point at center
Array Chunk Bytes 5.02 MiB 395.51 kiB Shape (50, 13, 90, 90) (50, 1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type bool numpy.ndarray - maskW(k, face, j, i_g)booldask.array<chunksize=(50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- cell_vertical_fraction_at_u_location
- long_name :
- mask denoting wet point at interface
Array Chunk Bytes 5.02 MiB 395.51 kiB Shape (50, 13, 90, 90) (50, 1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type bool numpy.ndarray - maskS(k, face, j_g, i)booldask.array<chunksize=(50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- cell_vertical_fraction_at_v_location
- long_name :
- mask denoting wet point at interface
Array Chunk Bytes 5.02 MiB 395.51 kiB Shape (50, 13, 90, 90) (50, 1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type bool numpy.ndarray - maskCtrlC(k, face, j, i)booldask.array<chunksize=(50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- ctrl_vector_3d_mask
- long_name :
- CTRL 3D mask where ctrl vector is active at tracer location
- units :
Array Chunk Bytes 5.02 MiB 395.51 kiB Shape (50, 13, 90, 90) (50, 1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type bool numpy.ndarray - maskCtrlS(k, face, j_g, i)booldask.array<chunksize=(50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- ctrl_vector_3d_mask_at_v_location
- long_name :
- CTRL 3D mask where ctrl vector is active at v location
- units :
Array Chunk Bytes 5.02 MiB 395.51 kiB Shape (50, 13, 90, 90) (50, 1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type bool numpy.ndarray - rhoRef(k)>f4dask.array<chunksize=(50,), meta=np.ndarray>
- standard_name :
- reference_density_profile
- long_name :
- 1D, vertical reference density profile
- coordinate :
- Z
- units :
- kg m-3
Array Chunk Bytes 200 B 200 B Shape (50,) (50,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - maskCtrlW(k, face, j, i_g)booldask.array<chunksize=(50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- ctrl_vector_3d_mask_at_u_location
- long_name :
- CTRL 3D mask where ctrl vector is active at u location
- units :
Array Chunk Bytes 5.02 MiB 395.51 kiB Shape (50, 13, 90, 90) (50, 1, 90, 90) Dask graph 13 chunks in 1 graph layer Data type bool numpy.ndarray - iter(time)float64dask.array<chunksize=(120,), meta=np.ndarray>
- standard_name :
- timestep
- long_name :
- model timestep number
Array Chunk Bytes 1.38 kiB 0.94 kiB Shape (176,) (120,) Dask graph 2 chunks in 7 graph layers Data type float64 numpy.ndarray - drC_kl(k_l)>f4dask.array<chunksize=(50,), meta=np.ndarray>
- standard_name :
- cell_z_size_at_w_location
- long_name :
- cell z size
- units :
- m
Array Chunk Bytes 200 B 200 B Shape (50,) (50,) Dask graph 1 chunks in 2 graph layers Data type >f4 numpy.ndarray - node(node)<U5'adj' 'const'
array(['adj', 'const'], dtype='<U5')
- GM_CHIXO(node, time, k, face, j, i_g)float32dask.array<chunksize=(1, 120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_CHIXO
- long_name :
- Redi Temp variance dissipation rate: X component
- units :
- degC^2/s
- mate :
- GM_CHIX
Array Chunk Bytes 6.90 GiB 185.39 MiB Shape (2, 176, 50, 13, 90, 90) (1, 120, 50, 1, 90, 90) Dask graph 52 chunks in 24 graph layers Data type float32 numpy.ndarray - GM_CHIZ(node, time, k_l, face, j, i)float32dask.array<chunksize=(1, 120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_CHIZ
- long_name :
- Redi Temp variance dissipation rate: Z component
- units :
- degC^2/s
Array Chunk Bytes 6.90 GiB 185.39 MiB Shape (2, 176, 50, 13, 90, 90) (1, 120, 50, 1, 90, 90) Dask graph 52 chunks in 5 graph layers Data type float32 numpy.ndarray - GM_ubT(node, time, k, face, j, i_g)float32dask.array<chunksize=(1, 120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_ubT
- long_name :
- Zonal Mass-Weight Bolus Transp of Pot Temp
- units :
- degC.m^3/s
- mate :
- GM_vbT
Array Chunk Bytes 6.90 GiB 185.39 MiB Shape (2, 176, 50, 13, 90, 90) (1, 120, 50, 1, 90, 90) Dask graph 52 chunks in 5 graph layers Data type float32 numpy.ndarray - GM_CHIX(node, time, k, face, j, i_g)float32dask.array<chunksize=(1, 120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_CHIX
- long_name :
- Redi Temp variance dissipation rate: X component
- units :
- degC^2/s
- mate :
- GM_KwzTz
Array Chunk Bytes 6.90 GiB 185.39 MiB Shape (2, 176, 50, 13, 90, 90) (1, 120, 50, 1, 90, 90) Dask graph 52 chunks in 5 graph layers Data type float32 numpy.ndarray - GM_CHIYO(node, time, k, face, j_g, i)float32dask.array<chunksize=(1, 120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_CHIYO
- long_name :
- Redi Temp variance dissipation rate: Y component
- units :
- degC^2/s
- mate :
- GM_CHIY
Array Chunk Bytes 6.90 GiB 185.39 MiB Shape (2, 176, 50, 13, 90, 90) (1, 120, 50, 1, 90, 90) Dask graph 52 chunks in 24 graph layers Data type float32 numpy.ndarray - GM_Kvz(node, time, k, face, j_g, i)float32dask.array<chunksize=(1, 120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_Kvz
- long_name :
- K_23 element (V.point, Z.dir) of GM-Redi tensor
- units :
- m^2/s
- mate :
- GM_Kuz
Array Chunk Bytes 6.90 GiB 185.39 MiB Shape (2, 176, 50, 13, 90, 90) (1, 120, 50, 1, 90, 90) Dask graph 52 chunks in 24 graph layers Data type float32 numpy.ndarray - GM_Kwz(node, time, k_l, face, j, i)float64dask.array<chunksize=(1, 120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_Kwz
- long_name :
- K_33 element (W.point, Z.dir) of GM-Redi tensor
- units :
- m^2/s
Array Chunk Bytes 13.81 GiB 370.79 MiB Shape (2, 176, 50, 13, 90, 90) (1, 120, 50, 1, 90, 90) Dask graph 52 chunks in 6 graph layers Data type float64 numpy.ndarray - THETA(node, time, k, face, j, i)float32dask.array<chunksize=(1, 120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- THETA
- long_name :
- Potential Temperature
- units :
- degC
Array Chunk Bytes 6.90 GiB 185.39 MiB Shape (2, 176, 50, 13, 90, 90) (1, 120, 50, 1, 90, 90) Dask graph 52 chunks in 24 graph layers Data type float32 numpy.ndarray - GM_Kvy(node, time, k, face, j_g, i)float32dask.array<chunksize=(1, 120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_Kvy
- long_name :
- K_22 element (V.point, Y.dir) of GM-Redi tensor
- units :
- m^2/s
- mate :
- GM_Kux
Array Chunk Bytes 6.90 GiB 185.39 MiB Shape (2, 176, 50, 13, 90, 90) (1, 120, 50, 1, 90, 90) Dask graph 52 chunks in 5 graph layers Data type float32 numpy.ndarray - GM_Kwy(node, time, k_l, face, j, i)float32dask.array<chunksize=(1, 120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_Kwy
- long_name :
- K_32 element (W.point, Y.dir) of GM-Redi tensor
- units :
- m^2/s
- mate :
- GM_Kwx
Array Chunk Bytes 6.90 GiB 185.39 MiB Shape (2, 176, 50, 13, 90, 90) (1, 120, 50, 1, 90, 90) Dask graph 52 chunks in 24 graph layers Data type float32 numpy.ndarray - GM_KwzTz(node, time, k_l, face, j, i)float32dask.array<chunksize=(1, 120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_KwzTz
- long_name :
- Redi main-diagonal vertical Temperature flux
- units :
- degC.m^3/s
Array Chunk Bytes 6.90 GiB 185.39 MiB Shape (2, 176, 50, 13, 90, 90) (1, 120, 50, 1, 90, 90) Dask graph 52 chunks in 5 graph layers Data type float32 numpy.ndarray - GM_Kux(node, time, k, face, j, i_g)float32dask.array<chunksize=(1, 120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_Kux
- long_name :
- K_11 element (U.point, X.dir) of GM-Redi tensor
- units :
- m^2/s
- mate :
- GM_Kvy
Array Chunk Bytes 6.90 GiB 185.39 MiB Shape (2, 176, 50, 13, 90, 90) (1, 120, 50, 1, 90, 90) Dask graph 52 chunks in 5 graph layers Data type float32 numpy.ndarray - GM_Kwx(node, time, k_l, face, j, i)float32dask.array<chunksize=(1, 120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_Kwx
- long_name :
- K_31 element (W.point, X.dir) of GM-Redi tensor
- units :
- m^2/s
- mate :
- GM_Kwy
Array Chunk Bytes 6.90 GiB 185.39 MiB Shape (2, 176, 50, 13, 90, 90) (1, 120, 50, 1, 90, 90) Dask graph 52 chunks in 24 graph layers Data type float32 numpy.ndarray - GM_CHIY(node, time, k, face, j_g, i)float32dask.array<chunksize=(1, 120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_CHIY
- long_name :
- Redi Temp variance dissipation rate: Y component
- units :
- degC^2/s
- mate :
- GM_CHIXO
Array Chunk Bytes 6.90 GiB 185.39 MiB Shape (2, 176, 50, 13, 90, 90) (1, 120, 50, 1, 90, 90) Dask graph 52 chunks in 5 graph layers Data type float32 numpy.ndarray - GM_Kuz(node, time, k, face, j, i_g)float32dask.array<chunksize=(1, 120, 50, 1, 90, 90), meta=np.ndarray>
- standard_name :
- GM_Kuz
- long_name :
- K_13 element (U.point, Z.dir) of GM-Redi tensor
- units :
- m^2/s
- mate :
- GM_Kvz
Array Chunk Bytes 6.90 GiB 185.39 MiB Shape (2, 176, 50, 13, 90, 90) (1, 120, 50, 1, 90, 90) Dask graph 52 chunks in 24 graph layers Data type float32 numpy.ndarray - Tx(node, time, k, face, j, i_g)float32dask.array<chunksize=(1, 120, 50, 1, 90, 1), meta=np.ndarray>
Array Chunk Bytes 6.90 GiB 183.33 MiB Shape (2, 176, 50, 13, 90, 90) (1, 120, 50, 1, 90, 89) Dask graph 104 chunks in 41 graph layers Data type float32 numpy.ndarray - Ty(node, time, k, face, j_g, i)float32dask.array<chunksize=(1, 120, 50, 1, 1, 90), meta=np.ndarray>
Array Chunk Bytes 6.90 GiB 183.33 MiB Shape (2, 176, 50, 13, 90, 90) (1, 120, 50, 1, 89, 90) Dask graph 104 chunks in 41 graph layers Data type float32 numpy.ndarray - Tz(node, time, k_l, face, j, i)float32dask.array<chunksize=(1, 120, 1, 1, 90, 90), meta=np.ndarray>
Array Chunk Bytes 6.90 GiB 181.69 MiB Shape (2, 176, 50, 13, 90, 90) (1, 120, 49, 1, 90, 90) Dask graph 104 chunks in 42 graph layers Data type float32 numpy.ndarray - GM_CHI(node, time, k, face, j, i)float32dask.array<chunksize=(1, 120, 49, 1, 89, 89), meta=np.ndarray>
Array Chunk Bytes 6.90 GiB 177.67 MiB Shape (2, 176, 50, 13, 90, 90) (1, 120, 49, 1, 89, 89) Dask graph 416 chunks in 85 graph layers Data type float32 numpy.ndarray
- iPandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89], dtype='int64', name='i')) - i_gPandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89], dtype='int64', name='i_g')) - jPandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89], dtype='int64', name='j')) - j_gPandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89], dtype='int64', name='j_g')) - kPandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49], dtype='int64', name='k')) - k_uPandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49], dtype='int64', name='k_u')) - k_lPandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49], dtype='int64', name='k_l')) - k_p1PandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50], dtype='int64', name='k_p1')) - facePandasIndex
PandasIndex(Int64Index([0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12], dtype='int64', name='face'))
- timePandasIndex
PandasIndex(TimedeltaIndex(['0 days 00:12:12', '0 days 00:23:48', '0 days 00:36:12', '0 days 00:48:12', '0 days 01:00:36', '0 days 01:12:36', '0 days 01:25:00', '0 days 01:37:24', '0 days 01:49:24', '0 days 02:01:48', ... '1 days 09:53:00', '1 days 10:05:24', '1 days 10:17:48', '1 days 10:29:00', '1 days 10:41:24', '1 days 10:53:24', '1 days 11:05:48', '1 days 11:17:48', '1 days 11:30:12', '1 days 11:42:36'], dtype='timedelta64[ns]', name='time', length=176, freq=None)) - nodePandasIndex
PandasIndex(Index(['adj', 'const'], dtype='object', name='node'))
- Conventions :
- CF-1.6
- title :
- netCDF wrapper of MITgcm MDS binary data
- source :
- MITgcm
- history :
- Created by calling `open_mdsdataset(grid_dir='/glade/work/dcherian/mitgcm/ECCOV4/release4/run/diags/../', iters=None, prefix=None, read_grid=True, delta_t=1, ref_date=None, calendar='gregorian', geometry='llc', grid_vars_to_coords=False, swap_dims=False, endian='>', chunks={'face': 1, 'k_l': -1, 'k': -1}, ignore_unknown_vars=False, default_dtype=<class 'numpy.float64'>, nx=None, ny=None, nz=None, llc_method='smallchunks', extra_metadata=None, extra_variables=None, data_dir='/glade/work/dcherian/mitgcm/ECCOV4/release4/run/diags/GM_CHIXO', levels=None)`
natre = concatted.sel(face=2).isel(
i=slice(10, 18), j=slice(10, 20), i_g=slice(10, 18), j_g=slice(10, 20)
)
natre
<xarray.Dataset>
Dimensions: (i: 8, i_g: 8, j: 10, j_g: 10, k: 50, k_u: 50, k_l: 50,
k_p1: 51, time: 176, node: 2)
Coordinates: (12/46)
* i (i) int64 10 11 12 13 14 15 16 17
* i_g (i_g) int64 10 11 12 13 14 15 16 17
* j (j) int64 10 11 12 13 14 15 16 17 18 19
* j_g (j_g) int64 10 11 12 13 14 15 16 17 18 19
* k (k) int64 0 1 2 3 4 5 6 7 8 9 ... 40 41 42 43 44 45 46 47 48 49
* k_u (k_u) int64 0 1 2 3 4 5 6 7 8 9 ... 40 41 42 43 44 45 46 47 48 49
... ...
maskCtrlS (k, j_g, i) bool dask.array<chunksize=(50, 10, 8), meta=np.ndarray>
rhoRef (k) >f4 dask.array<chunksize=(50,), meta=np.ndarray>
maskCtrlW (k, j, i_g) bool dask.array<chunksize=(50, 10, 8), meta=np.ndarray>
iter (time) float64 dask.array<chunksize=(120,), meta=np.ndarray>
drC_kl (k_l) >f4 dask.array<chunksize=(50,), meta=np.ndarray>
* node (node) <U5 'adj' 'const'
Data variables: (12/19)
GM_CHIXO (node, time, k, j, i_g) float32 dask.array<chunksize=(1, 120, 50, 10, 8), meta=np.ndarray>
GM_CHIZ (node, time, k_l, j, i) float32 dask.array<chunksize=(1, 120, 50, 10, 8), meta=np.ndarray>
GM_ubT (node, time, k, j, i_g) float32 dask.array<chunksize=(1, 120, 50, 10, 8), meta=np.ndarray>
GM_CHIX (node, time, k, j, i_g) float32 dask.array<chunksize=(1, 120, 50, 10, 8), meta=np.ndarray>
GM_CHIYO (node, time, k, j_g, i) float32 dask.array<chunksize=(1, 120, 50, 10, 8), meta=np.ndarray>
GM_Kvz (node, time, k, j_g, i) float32 dask.array<chunksize=(1, 120, 50, 10, 8), meta=np.ndarray>
... ...
GM_CHIY (node, time, k, j_g, i) float32 dask.array<chunksize=(1, 120, 50, 10, 8), meta=np.ndarray>
GM_Kuz (node, time, k, j, i_g) float32 dask.array<chunksize=(1, 120, 50, 10, 8), meta=np.ndarray>
Tx (node, time, k, j, i_g) float32 dask.array<chunksize=(1, 120, 50, 10, 8), meta=np.ndarray>
Ty (node, time, k, j_g, i) float32 dask.array<chunksize=(1, 120, 50, 10, 8), meta=np.ndarray>
Tz (node, time, k_l, j, i) float32 dask.array<chunksize=(1, 120, 1, 10, 8), meta=np.ndarray>
GM_CHI (node, time, k, j, i) float32 dask.array<chunksize=(1, 120, 49, 10, 8), meta=np.ndarray>
Attributes:
Conventions: CF-1.6
title: netCDF wrapper of MITgcm MDS binary data
source: MITgcm
history: Created by calling `open_mdsdataset(grid_dir='/glade/work/d...xarray.Dataset
- i: 8
- i_g: 8
- j: 10
- j_g: 10
- k: 50
- k_u: 50
- k_l: 50
- k_p1: 51
- time: 176
- node: 2
- i(i)int6410 11 12 13 14 15 16 17
- standard_name :
- x_grid_index
- axis :
- X
- long_name :
- x-dimension of the t grid
- swap_dim :
- XC
array([10, 11, 12, 13, 14, 15, 16, 17])
- i_g(i_g)int6410 11 12 13 14 15 16 17
- standard_name :
- x_grid_index_at_u_location
- axis :
- X
- long_name :
- x-dimension of the u grid
- c_grid_axis_shift :
- -0.5
- swap_dim :
- XG
array([10, 11, 12, 13, 14, 15, 16, 17])
- j(j)int6410 11 12 13 14 15 16 17 18 19
- standard_name :
- y_grid_index
- axis :
- Y
- long_name :
- y-dimension of the t grid
- swap_dim :
- YC
array([10, 11, 12, 13, 14, 15, 16, 17, 18, 19])
- j_g(j_g)int6410 11 12 13 14 15 16 17 18 19
- standard_name :
- y_grid_index_at_v_location
- axis :
- Y
- long_name :
- y-dimension of the v grid
- c_grid_axis_shift :
- -0.5
- swap_dim :
- YG
array([10, 11, 12, 13, 14, 15, 16, 17, 18, 19])
- k(k)int640 1 2 3 4 5 6 ... 44 45 46 47 48 49
- standard_name :
- z_grid_index
- axis :
- Z
- long_name :
- z-dimension of the t grid
- swap_dim :
- Z
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49]) - k_u(k_u)int640 1 2 3 4 5 6 ... 44 45 46 47 48 49
- standard_name :
- z_grid_index_at_upper_w_location
- axis :
- Z
- long_name :
- z-dimension of the w grid
- c_grid_axis_shift :
- 0.5
- swap_dim :
- Zu
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49]) - k_l(k_l)int640 1 2 3 4 5 6 ... 44 45 46 47 48 49
- standard_name :
- z_grid_index_at_lower_w_location
- axis :
- Z
- long_name :
- z-dimension of the w grid
- c_grid_axis_shift :
- -0.5
- swap_dim :
- Zl
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49]) - k_p1(k_p1)int640 1 2 3 4 5 6 ... 45 46 47 48 49 50
- standard_name :
- z_grid_index_at_w_location
- axis :
- Z
- long_name :
- z-dimension of the w grid
- c_grid_axis_shift :
- (-0.5, 0.5)
- swap_dim :
- Zp1
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50]) - face()int642
- standard_name :
- face_index
array(2)
- time(time)timedelta64[ns]00:12:12 ... 1 days 11:42:36
- standard_name :
- time
- long_name :
- Time
- axis :
- T
- calendar :
- gregorian
array([ 732000000000, 1428000000000, 2172000000000, 2892000000000, 3636000000000, 4356000000000, 5100000000000, 5844000000000, 6564000000000, 7308000000000, 8028000000000, 8772000000000, 9516000000000, 10188000000000, 10932000000000, 11652000000000, 12396000000000, 13116000000000, 13860000000000, 14604000000000, 15324000000000, 16068000000000, 16788000000000, 17532000000000, 18276000000000, 18948000000000, 19692000000000, 20412000000000, 21156000000000, 21876000000000, 22620000000000, 23364000000000, 24084000000000, 24828000000000, 25548000000000, 26292000000000, 27036000000000, 27708000000000, 28452000000000, 29172000000000, 29916000000000, 30636000000000, 31380000000000, 32124000000000, 32844000000000, 33588000000000, 34308000000000, 35052000000000, 35796000000000, 36492000000000, 37236000000000, 37956000000000, 38700000000000, 39420000000000, 40164000000000, 40908000000000, 41628000000000, 42372000000000, 43092000000000, 43836000000000, 44580000000000, 45252000000000, 45996000000000, 46716000000000, 47460000000000, 48180000000000, 48924000000000, 49668000000000, 50388000000000, 51132000000000, 51852000000000, 52596000000000, 53340000000000, 54012000000000, 54756000000000, 55476000000000, 56220000000000, 56940000000000, 57684000000000, 58428000000000, 59148000000000, 59892000000000, 60612000000000, 61356000000000, 62100000000000, 62772000000000, 63516000000000, 64236000000000, 64980000000000, 65700000000000, 66444000000000, 67188000000000, 67908000000000, 68652000000000, 69372000000000, 70116000000000, 70860000000000, 71556000000000, 72300000000000, 73020000000000, 73764000000000, 74484000000000, 75228000000000, 75972000000000, 76692000000000, 77436000000000, 78156000000000, 78900000000000, 79644000000000, 80316000000000, 81060000000000, 81780000000000, 82524000000000, 83244000000000, 83988000000000, 84732000000000, 85452000000000, 86196000000000, 86916000000000, 87660000000000, 88404000000000, 89076000000000, 89820000000000, 90540000000000, 91284000000000, 92004000000000, 92748000000000, 93492000000000, 94212000000000, 94956000000000, 95676000000000, 96420000000000, 97164000000000, 97836000000000, 98580000000000, 99300000000000, 100044000000000, 100764000000000, 101508000000000, 102252000000000, 102972000000000, 103716000000000, 104436000000000, 105180000000000, 105924000000000, 106620000000000, 107364000000000, 108084000000000, 108828000000000, 109548000000000, 110292000000000, 111036000000000, 111756000000000, 112500000000000, 113220000000000, 113964000000000, 114708000000000, 115380000000000, 116124000000000, 116844000000000, 117588000000000, 118308000000000, 119052000000000, 119796000000000, 120516000000000, 121260000000000, 121980000000000, 122724000000000, 123468000000000, 124140000000000, 124884000000000, 125604000000000, 126348000000000, 127068000000000, 127812000000000, 128556000000000], dtype='timedelta64[ns]') - XC(j, i)>f4dask.array<chunksize=(10, 8), meta=np.ndarray>
- standard_name :
- longitude
- long_name :
- longitude
- units :
- degrees_east
- coordinate :
- YC XC
Array Chunk Bytes 320 B 320 B Shape (10, 8) (10, 8) Dask graph 1 chunks in 3 graph layers Data type >f4 numpy.ndarray - YC(j, i)>f4dask.array<chunksize=(10, 8), meta=np.ndarray>
- standard_name :
- latitude
- long_name :
- latitude
- units :
- degrees_north
- coordinate :
- YC XC
Array Chunk Bytes 320 B 320 B Shape (10, 8) (10, 8) Dask graph 1 chunks in 3 graph layers Data type >f4 numpy.ndarray - XG(j_g, i_g)>f4dask.array<chunksize=(10, 8), meta=np.ndarray>
- standard_name :
- longitude_at_f_location
- long_name :
- longitude
- units :
- degrees_east
- coordinate :
- YG XG
Array Chunk Bytes 320 B 320 B Shape (10, 8) (10, 8) Dask graph 1 chunks in 3 graph layers Data type >f4 numpy.ndarray - YG(j_g, i_g)>f4dask.array<chunksize=(10, 8), meta=np.ndarray>
- standard_name :
- latitude_at_f_location
- long_name :
- latitude
- units :
- degrees_north
- coordinate :
- YG XG
Array Chunk Bytes 320 B 320 B Shape (10, 8) (10, 8) Dask graph 1 chunks in 3 graph layers Data type >f4 numpy.ndarray - CS(j, i)>f4dask.array<chunksize=(10, 8), meta=np.ndarray>
- standard_name :
- Cos of grid orientation angle
- long_name :
- AngleCS
- units :
- coordinate :
- YC XC
Array Chunk Bytes 320 B 320 B Shape (10, 8) (10, 8) Dask graph 1 chunks in 3 graph layers Data type >f4 numpy.ndarray - SN(j, i)>f4dask.array<chunksize=(10, 8), meta=np.ndarray>
- standard_name :
- Sin of grid orientation angle
- long_name :
- AngleSN
- units :
- coordinate :
- YC XC
Array Chunk Bytes 320 B 320 B Shape (10, 8) (10, 8) Dask graph 1 chunks in 3 graph layers Data type >f4 numpy.ndarray - Z(k)>f4dask.array<chunksize=(50,), meta=np.ndarray>
- standard_name :
- depth
- long_name :
- vertical coordinate of cell center
- units :
- m
- positive :
- down
Array Chunk Bytes 200 B 200 B Shape (50,) (50,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - Zp1(k_p1)>f4dask.array<chunksize=(51,), meta=np.ndarray>
- standard_name :
- depth_at_w_location
- long_name :
- vertical coordinate of cell interface
- units :
- m
- positive :
- down
Array Chunk Bytes 204 B 204 B Shape (51,) (51,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - Zu(k_u)>f4dask.array<chunksize=(50,), meta=np.ndarray>
- standard_name :
- depth_at_upper_w_location
- long_name :
- vertical coordinate of upper cell interface
- units :
- m
- positive :
- down
Array Chunk Bytes 200 B 200 B Shape (50,) (50,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - Zl(k_l)>f4dask.array<chunksize=(50,), meta=np.ndarray>
- standard_name :
- depth_at_lower_w_location
- long_name :
- vertical coordinate of lower cell interface
- units :
- m
- positive :
- down
Array Chunk Bytes 200 B 200 B Shape (50,) (50,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - rA(j, i)>f4dask.array<chunksize=(10, 8), meta=np.ndarray>
- standard_name :
- cell_area
- long_name :
- cell area
- units :
- m2
- coordinate :
- YC XC
Array Chunk Bytes 320 B 320 B Shape (10, 8) (10, 8) Dask graph 1 chunks in 3 graph layers Data type >f4 numpy.ndarray - dxG(j_g, i)>f4dask.array<chunksize=(10, 8), meta=np.ndarray>
- standard_name :
- cell_x_size_at_v_location
- long_name :
- cell x size
- units :
- m
- coordinate :
- YG XC
Array Chunk Bytes 320 B 320 B Shape (10, 8) (10, 8) Dask graph 1 chunks in 3 graph layers Data type >f4 numpy.ndarray - dyG(j, i_g)>f4dask.array<chunksize=(10, 8), meta=np.ndarray>
- standard_name :
- cell_y_size_at_u_location
- long_name :
- cell y size
- units :
- m
- coordinate :
- YC XG
Array Chunk Bytes 320 B 320 B Shape (10, 8) (10, 8) Dask graph 1 chunks in 3 graph layers Data type >f4 numpy.ndarray - Depth(j, i)>f4dask.array<chunksize=(10, 8), meta=np.ndarray>
- standard_name :
- ocean_depth
- long_name :
- ocean depth
- units :
- m
- coordinate :
- XC YC
Array Chunk Bytes 320 B 320 B Shape (10, 8) (10, 8) Dask graph 1 chunks in 3 graph layers Data type >f4 numpy.ndarray - rAz(j_g, i_g)>f4dask.array<chunksize=(10, 8), meta=np.ndarray>
- standard_name :
- cell_area_at_f_location
- long_name :
- cell area
- units :
- m
- coordinate :
- YG XG
Array Chunk Bytes 320 B 320 B Shape (10, 8) (10, 8) Dask graph 1 chunks in 3 graph layers Data type >f4 numpy.ndarray - dxC(j, i_g)>f4dask.array<chunksize=(10, 8), meta=np.ndarray>
- standard_name :
- cell_x_size_at_u_location
- long_name :
- cell x size
- units :
- m
- coordinate :
- YC XG
Array Chunk Bytes 320 B 320 B Shape (10, 8) (10, 8) Dask graph 1 chunks in 3 graph layers Data type >f4 numpy.ndarray - dyC(j_g, i)>f4dask.array<chunksize=(10, 8), meta=np.ndarray>
- standard_name :
- cell_y_size_at_v_location
- long_name :
- cell y size
- units :
- m
- coordinate :
- YG XC
Array Chunk Bytes 320 B 320 B Shape (10, 8) (10, 8) Dask graph 1 chunks in 3 graph layers Data type >f4 numpy.ndarray - rAw(j, i_g)>f4dask.array<chunksize=(10, 8), meta=np.ndarray>
- standard_name :
- cell_area_at_u_location
- long_name :
- cell area
- units :
- m2
- coordinate :
- YG XC
Array Chunk Bytes 320 B 320 B Shape (10, 8) (10, 8) Dask graph 1 chunks in 3 graph layers Data type >f4 numpy.ndarray - rAs(j_g, i)>f4dask.array<chunksize=(10, 8), meta=np.ndarray>
- standard_name :
- cell_area_at_v_location
- long_name :
- cell area
- units :
- m2
- coordinate :
- YG XC
Array Chunk Bytes 320 B 320 B Shape (10, 8) (10, 8) Dask graph 1 chunks in 3 graph layers Data type >f4 numpy.ndarray - drC(k_p1)>f4dask.array<chunksize=(51,), meta=np.ndarray>
- standard_name :
- cell_z_size_at_w_location
- long_name :
- cell z size
- units :
- m
Array Chunk Bytes 204 B 204 B Shape (51,) (51,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - drF(k)>f4dask.array<chunksize=(50,), meta=np.ndarray>
- standard_name :
- cell_z_size
- long_name :
- cell z size
- units :
- m
Array Chunk Bytes 200 B 200 B Shape (50,) (50,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - PHrefC(k)>f4dask.array<chunksize=(50,), meta=np.ndarray>
- standard_name :
- cell_reference_pressure
- long_name :
- Reference Hydrostatic Pressure
- units :
- m2 s-2
Array Chunk Bytes 200 B 200 B Shape (50,) (50,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - PHrefF(k_p1)>f4dask.array<chunksize=(51,), meta=np.ndarray>
- standard_name :
- cell_reference_pressure
- long_name :
- Reference Hydrostatic Pressure
- units :
- m2 s-2
Array Chunk Bytes 204 B 204 B Shape (51,) (51,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - hFacC(k, j, i)>f4dask.array<chunksize=(50, 10, 8), meta=np.ndarray>
- standard_name :
- cell_vertical_fraction
- long_name :
- vertical fraction of open cell
Array Chunk Bytes 15.62 kiB 15.62 kiB Shape (50, 10, 8) (50, 10, 8) Dask graph 1 chunks in 3 graph layers Data type >f4 numpy.ndarray - hFacW(k, j, i_g)>f4dask.array<chunksize=(50, 10, 8), meta=np.ndarray>
- standard_name :
- cell_vertical_fraction_at_u_location
- long_name :
- vertical fraction of open cell
Array Chunk Bytes 15.62 kiB 15.62 kiB Shape (50, 10, 8) (50, 10, 8) Dask graph 1 chunks in 3 graph layers Data type >f4 numpy.ndarray - hFacS(k, j_g, i)>f4dask.array<chunksize=(50, 10, 8), meta=np.ndarray>
- standard_name :
- cell_vertical_fraction_at_v_location
- long_name :
- vertical fraction of open cell
Array Chunk Bytes 15.62 kiB 15.62 kiB Shape (50, 10, 8) (50, 10, 8) Dask graph 1 chunks in 3 graph layers Data type >f4 numpy.ndarray - maskC(k, j, i)booldask.array<chunksize=(50, 10, 8), meta=np.ndarray>
- standard_name :
- sea_binary_mask_at_t_location
- long_name :
- mask denoting wet point at center
Array Chunk Bytes 3.91 kiB 3.91 kiB Shape (50, 10, 8) (50, 10, 8) Dask graph 1 chunks in 3 graph layers Data type bool numpy.ndarray - maskW(k, j, i_g)booldask.array<chunksize=(50, 10, 8), meta=np.ndarray>
- standard_name :
- cell_vertical_fraction_at_u_location
- long_name :
- mask denoting wet point at interface
Array Chunk Bytes 3.91 kiB 3.91 kiB Shape (50, 10, 8) (50, 10, 8) Dask graph 1 chunks in 3 graph layers Data type bool numpy.ndarray - maskS(k, j_g, i)booldask.array<chunksize=(50, 10, 8), meta=np.ndarray>
- standard_name :
- cell_vertical_fraction_at_v_location
- long_name :
- mask denoting wet point at interface
Array Chunk Bytes 3.91 kiB 3.91 kiB Shape (50, 10, 8) (50, 10, 8) Dask graph 1 chunks in 3 graph layers Data type bool numpy.ndarray - maskCtrlC(k, j, i)booldask.array<chunksize=(50, 10, 8), meta=np.ndarray>
- standard_name :
- ctrl_vector_3d_mask
- long_name :
- CTRL 3D mask where ctrl vector is active at tracer location
- units :
Array Chunk Bytes 3.91 kiB 3.91 kiB Shape (50, 10, 8) (50, 10, 8) Dask graph 1 chunks in 3 graph layers Data type bool numpy.ndarray - maskCtrlS(k, j_g, i)booldask.array<chunksize=(50, 10, 8), meta=np.ndarray>
- standard_name :
- ctrl_vector_3d_mask_at_v_location
- long_name :
- CTRL 3D mask where ctrl vector is active at v location
- units :
Array Chunk Bytes 3.91 kiB 3.91 kiB Shape (50, 10, 8) (50, 10, 8) Dask graph 1 chunks in 3 graph layers Data type bool numpy.ndarray - rhoRef(k)>f4dask.array<chunksize=(50,), meta=np.ndarray>
- standard_name :
- reference_density_profile
- long_name :
- 1D, vertical reference density profile
- coordinate :
- Z
- units :
- kg m-3
Array Chunk Bytes 200 B 200 B Shape (50,) (50,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - maskCtrlW(k, j, i_g)booldask.array<chunksize=(50, 10, 8), meta=np.ndarray>
- standard_name :
- ctrl_vector_3d_mask_at_u_location
- long_name :
- CTRL 3D mask where ctrl vector is active at u location
- units :
Array Chunk Bytes 3.91 kiB 3.91 kiB Shape (50, 10, 8) (50, 10, 8) Dask graph 1 chunks in 3 graph layers Data type bool numpy.ndarray - iter(time)float64dask.array<chunksize=(120,), meta=np.ndarray>
- standard_name :
- timestep
- long_name :
- model timestep number
Array Chunk Bytes 1.38 kiB 0.94 kiB Shape (176,) (120,) Dask graph 2 chunks in 7 graph layers Data type float64 numpy.ndarray - drC_kl(k_l)>f4dask.array<chunksize=(50,), meta=np.ndarray>
- standard_name :
- cell_z_size_at_w_location
- long_name :
- cell z size
- units :
- m
Array Chunk Bytes 200 B 200 B Shape (50,) (50,) Dask graph 1 chunks in 2 graph layers Data type >f4 numpy.ndarray - node(node)<U5'adj' 'const'
array(['adj', 'const'], dtype='<U5')
- GM_CHIXO(node, time, k, j, i_g)float32dask.array<chunksize=(1, 120, 50, 10, 8), meta=np.ndarray>
- standard_name :
- GM_CHIXO
- long_name :
- Redi Temp variance dissipation rate: X component
- units :
- degC^2/s
- mate :
- GM_CHIX
Array Chunk Bytes 5.37 MiB 1.83 MiB Shape (2, 176, 50, 10, 8) (1, 120, 50, 10, 8) Dask graph 4 chunks in 26 graph layers Data type float32 numpy.ndarray - GM_CHIZ(node, time, k_l, j, i)float32dask.array<chunksize=(1, 120, 50, 10, 8), meta=np.ndarray>
- standard_name :
- GM_CHIZ
- long_name :
- Redi Temp variance dissipation rate: Z component
- units :
- degC^2/s
Array Chunk Bytes 5.37 MiB 1.83 MiB Shape (2, 176, 50, 10, 8) (1, 120, 50, 10, 8) Dask graph 4 chunks in 7 graph layers Data type float32 numpy.ndarray - GM_ubT(node, time, k, j, i_g)float32dask.array<chunksize=(1, 120, 50, 10, 8), meta=np.ndarray>
- standard_name :
- GM_ubT
- long_name :
- Zonal Mass-Weight Bolus Transp of Pot Temp
- units :
- degC.m^3/s
- mate :
- GM_vbT
Array Chunk Bytes 5.37 MiB 1.83 MiB Shape (2, 176, 50, 10, 8) (1, 120, 50, 10, 8) Dask graph 4 chunks in 7 graph layers Data type float32 numpy.ndarray - GM_CHIX(node, time, k, j, i_g)float32dask.array<chunksize=(1, 120, 50, 10, 8), meta=np.ndarray>
- standard_name :
- GM_CHIX
- long_name :
- Redi Temp variance dissipation rate: X component
- units :
- degC^2/s
- mate :
- GM_KwzTz
Array Chunk Bytes 5.37 MiB 1.83 MiB Shape (2, 176, 50, 10, 8) (1, 120, 50, 10, 8) Dask graph 4 chunks in 7 graph layers Data type float32 numpy.ndarray - GM_CHIYO(node, time, k, j_g, i)float32dask.array<chunksize=(1, 120, 50, 10, 8), meta=np.ndarray>
- standard_name :
- GM_CHIYO
- long_name :
- Redi Temp variance dissipation rate: Y component
- units :
- degC^2/s
- mate :
- GM_CHIY
Array Chunk Bytes 5.37 MiB 1.83 MiB Shape (2, 176, 50, 10, 8) (1, 120, 50, 10, 8) Dask graph 4 chunks in 26 graph layers Data type float32 numpy.ndarray - GM_Kvz(node, time, k, j_g, i)float32dask.array<chunksize=(1, 120, 50, 10, 8), meta=np.ndarray>
- standard_name :
- GM_Kvz
- long_name :
- K_23 element (V.point, Z.dir) of GM-Redi tensor
- units :
- m^2/s
- mate :
- GM_Kuz
Array Chunk Bytes 5.37 MiB 1.83 MiB Shape (2, 176, 50, 10, 8) (1, 120, 50, 10, 8) Dask graph 4 chunks in 26 graph layers Data type float32 numpy.ndarray - GM_Kwz(node, time, k_l, j, i)float64dask.array<chunksize=(1, 120, 50, 10, 8), meta=np.ndarray>
- standard_name :
- GM_Kwz
- long_name :
- K_33 element (W.point, Z.dir) of GM-Redi tensor
- units :
- m^2/s
Array Chunk Bytes 10.74 MiB 3.66 MiB Shape (2, 176, 50, 10, 8) (1, 120, 50, 10, 8) Dask graph 4 chunks in 8 graph layers Data type float64 numpy.ndarray - THETA(node, time, k, j, i)float32dask.array<chunksize=(1, 120, 50, 10, 8), meta=np.ndarray>
- standard_name :
- THETA
- long_name :
- Potential Temperature
- units :
- degC
Array Chunk Bytes 5.37 MiB 1.83 MiB Shape (2, 176, 50, 10, 8) (1, 120, 50, 10, 8) Dask graph 4 chunks in 26 graph layers Data type float32 numpy.ndarray - GM_Kvy(node, time, k, j_g, i)float32dask.array<chunksize=(1, 120, 50, 10, 8), meta=np.ndarray>
- standard_name :
- GM_Kvy
- long_name :
- K_22 element (V.point, Y.dir) of GM-Redi tensor
- units :
- m^2/s
- mate :
- GM_Kux
Array Chunk Bytes 5.37 MiB 1.83 MiB Shape (2, 176, 50, 10, 8) (1, 120, 50, 10, 8) Dask graph 4 chunks in 7 graph layers Data type float32 numpy.ndarray - GM_Kwy(node, time, k_l, j, i)float32dask.array<chunksize=(1, 120, 50, 10, 8), meta=np.ndarray>
- standard_name :
- GM_Kwy
- long_name :
- K_32 element (W.point, Y.dir) of GM-Redi tensor
- units :
- m^2/s
- mate :
- GM_Kwx
Array Chunk Bytes 5.37 MiB 1.83 MiB Shape (2, 176, 50, 10, 8) (1, 120, 50, 10, 8) Dask graph 4 chunks in 26 graph layers Data type float32 numpy.ndarray - GM_KwzTz(node, time, k_l, j, i)float32dask.array<chunksize=(1, 120, 50, 10, 8), meta=np.ndarray>
- standard_name :
- GM_KwzTz
- long_name :
- Redi main-diagonal vertical Temperature flux
- units :
- degC.m^3/s
Array Chunk Bytes 5.37 MiB 1.83 MiB Shape (2, 176, 50, 10, 8) (1, 120, 50, 10, 8) Dask graph 4 chunks in 7 graph layers Data type float32 numpy.ndarray - GM_Kux(node, time, k, j, i_g)float32dask.array<chunksize=(1, 120, 50, 10, 8), meta=np.ndarray>
- standard_name :
- GM_Kux
- long_name :
- K_11 element (U.point, X.dir) of GM-Redi tensor
- units :
- m^2/s
- mate :
- GM_Kvy
Array Chunk Bytes 5.37 MiB 1.83 MiB Shape (2, 176, 50, 10, 8) (1, 120, 50, 10, 8) Dask graph 4 chunks in 7 graph layers Data type float32 numpy.ndarray - GM_Kwx(node, time, k_l, j, i)float32dask.array<chunksize=(1, 120, 50, 10, 8), meta=np.ndarray>
- standard_name :
- GM_Kwx
- long_name :
- K_31 element (W.point, X.dir) of GM-Redi tensor
- units :
- m^2/s
- mate :
- GM_Kwy
Array Chunk Bytes 5.37 MiB 1.83 MiB Shape (2, 176, 50, 10, 8) (1, 120, 50, 10, 8) Dask graph 4 chunks in 26 graph layers Data type float32 numpy.ndarray - GM_CHIY(node, time, k, j_g, i)float32dask.array<chunksize=(1, 120, 50, 10, 8), meta=np.ndarray>
- standard_name :
- GM_CHIY
- long_name :
- Redi Temp variance dissipation rate: Y component
- units :
- degC^2/s
- mate :
- GM_CHIXO
Array Chunk Bytes 5.37 MiB 1.83 MiB Shape (2, 176, 50, 10, 8) (1, 120, 50, 10, 8) Dask graph 4 chunks in 7 graph layers Data type float32 numpy.ndarray - GM_Kuz(node, time, k, j, i_g)float32dask.array<chunksize=(1, 120, 50, 10, 8), meta=np.ndarray>
- standard_name :
- GM_Kuz
- long_name :
- K_13 element (U.point, Z.dir) of GM-Redi tensor
- units :
- m^2/s
- mate :
- GM_Kvz
Array Chunk Bytes 5.37 MiB 1.83 MiB Shape (2, 176, 50, 10, 8) (1, 120, 50, 10, 8) Dask graph 4 chunks in 26 graph layers Data type float32 numpy.ndarray - Tx(node, time, k, j, i_g)float32dask.array<chunksize=(1, 120, 50, 10, 8), meta=np.ndarray>
Array Chunk Bytes 5.37 MiB 1.83 MiB Shape (2, 176, 50, 10, 8) (1, 120, 50, 10, 8) Dask graph 4 chunks in 43 graph layers Data type float32 numpy.ndarray - Ty(node, time, k, j_g, i)float32dask.array<chunksize=(1, 120, 50, 10, 8), meta=np.ndarray>
Array Chunk Bytes 5.37 MiB 1.83 MiB Shape (2, 176, 50, 10, 8) (1, 120, 50, 10, 8) Dask graph 4 chunks in 43 graph layers Data type float32 numpy.ndarray - Tz(node, time, k_l, j, i)float32dask.array<chunksize=(1, 120, 1, 10, 8), meta=np.ndarray>
Array Chunk Bytes 5.37 MiB 1.79 MiB Shape (2, 176, 50, 10, 8) (1, 120, 49, 10, 8) Dask graph 8 chunks in 44 graph layers Data type float32 numpy.ndarray - GM_CHI(node, time, k, j, i)float32dask.array<chunksize=(1, 120, 49, 10, 8), meta=np.ndarray>
Array Chunk Bytes 5.37 MiB 1.79 MiB Shape (2, 176, 50, 10, 8) (1, 120, 49, 10, 8) Dask graph 8 chunks in 87 graph layers Data type float32 numpy.ndarray
- iPandasIndex
PandasIndex(Int64Index([10, 11, 12, 13, 14, 15, 16, 17], dtype='int64', name='i'))
- i_gPandasIndex
PandasIndex(Int64Index([10, 11, 12, 13, 14, 15, 16, 17], dtype='int64', name='i_g'))
- jPandasIndex
PandasIndex(Int64Index([10, 11, 12, 13, 14, 15, 16, 17, 18, 19], dtype='int64', name='j'))
- j_gPandasIndex
PandasIndex(Int64Index([10, 11, 12, 13, 14, 15, 16, 17, 18, 19], dtype='int64', name='j_g'))
- kPandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49], dtype='int64', name='k')) - k_uPandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49], dtype='int64', name='k_u')) - k_lPandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49], dtype='int64', name='k_l')) - k_p1PandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50], dtype='int64', name='k_p1')) - timePandasIndex
PandasIndex(TimedeltaIndex(['0 days 00:12:12', '0 days 00:23:48', '0 days 00:36:12', '0 days 00:48:12', '0 days 01:00:36', '0 days 01:12:36', '0 days 01:25:00', '0 days 01:37:24', '0 days 01:49:24', '0 days 02:01:48', ... '1 days 09:53:00', '1 days 10:05:24', '1 days 10:17:48', '1 days 10:29:00', '1 days 10:41:24', '1 days 10:53:24', '1 days 11:05:48', '1 days 11:17:48', '1 days 11:30:12', '1 days 11:42:36'], dtype='timedelta64[ns]', name='time', length=176, freq=None)) - nodePandasIndex
PandasIndex(Index(['adj', 'const'], dtype='object', name='node'))
- Conventions :
- CF-1.6
- title :
- netCDF wrapper of MITgcm MDS binary data
- source :
- MITgcm
- history :
- Created by calling `open_mdsdataset(grid_dir='/glade/work/dcherian/mitgcm/ECCOV4/release4/run/diags/../', iters=None, prefix=None, read_grid=True, delta_t=1, ref_date=None, calendar='gregorian', geometry='llc', grid_vars_to_coords=False, swap_dims=False, endian='>', chunks={'face': 1, 'k_l': -1, 'k': -1}, ignore_unknown_vars=False, default_dtype=<class 'numpy.float64'>, nx=None, ny=None, nz=None, llc_method='smallchunks', extra_metadata=None, extra_variables=None, data_dir='/glade/work/dcherian/mitgcm/ECCOV4/release4/run/diags/GM_CHIXO', levels=None)`
(natre,) = dask.optimize(natre)
natre_mean = natre.cf.mean(["X", "Y"]).load()
/glade/u/home/dcherian/miniconda3/envs/eddydiff/lib/python3.10/site-packages/dask_jobqueue/core.py:255: FutureWarning: job_extra has been renamed to job_extra_directives. You are still using it (even if only set to []; please also check config files). If you did not set job_extra_directives yet, job_extra will be respected for now, but it will be removed in a future release. If you already set job_extra_directives, job_extra is ignored and you can remove it.
warnings.warn(warn, FutureWarning)
/glade/u/home/dcherian/miniconda3/envs/eddydiff/lib/python3.10/site-packages/dask_jobqueue/core.py:274: FutureWarning: env_extra has been renamed to job_script_prologue. You are still using it (even if only set to []; please also check config files). If you did not set job_script_prologue yet, env_extra will be respected for now, but it will be removed in a future release. If you already set job_script_prologue, env_extra is ignored and you can remove it.
warnings.warn(warn, FutureWarning)
natre_mean.to_netcdf("../datasets/ecco-chi.nc")
natre_mean["GM_CHIX"].mean("time").cf.plot(hue="node")
[<matplotlib.lines.Line2D at 0x2ac11e83d180>,
<matplotlib.lines.Line2D at 0x2ac11e83d030>]
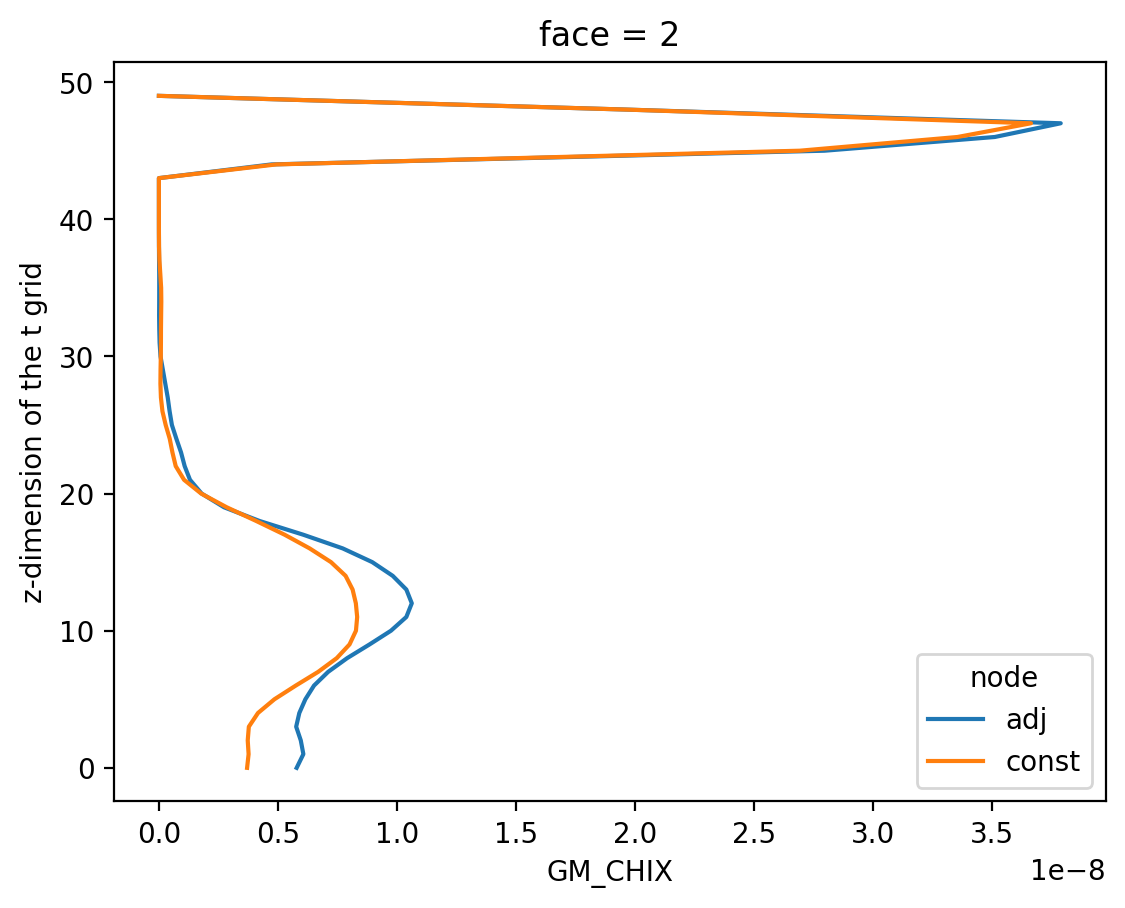
natre_mean["GM_Kux"].mean("time").cf.plot(hue="node")
natre_mean["GM_Kvy"].mean("time").cf.plot(hue="node")
[<matplotlib.lines.Line2D at 0x2abef70a2380>,
<matplotlib.lines.Line2D at 0x2abef70a38b0>]
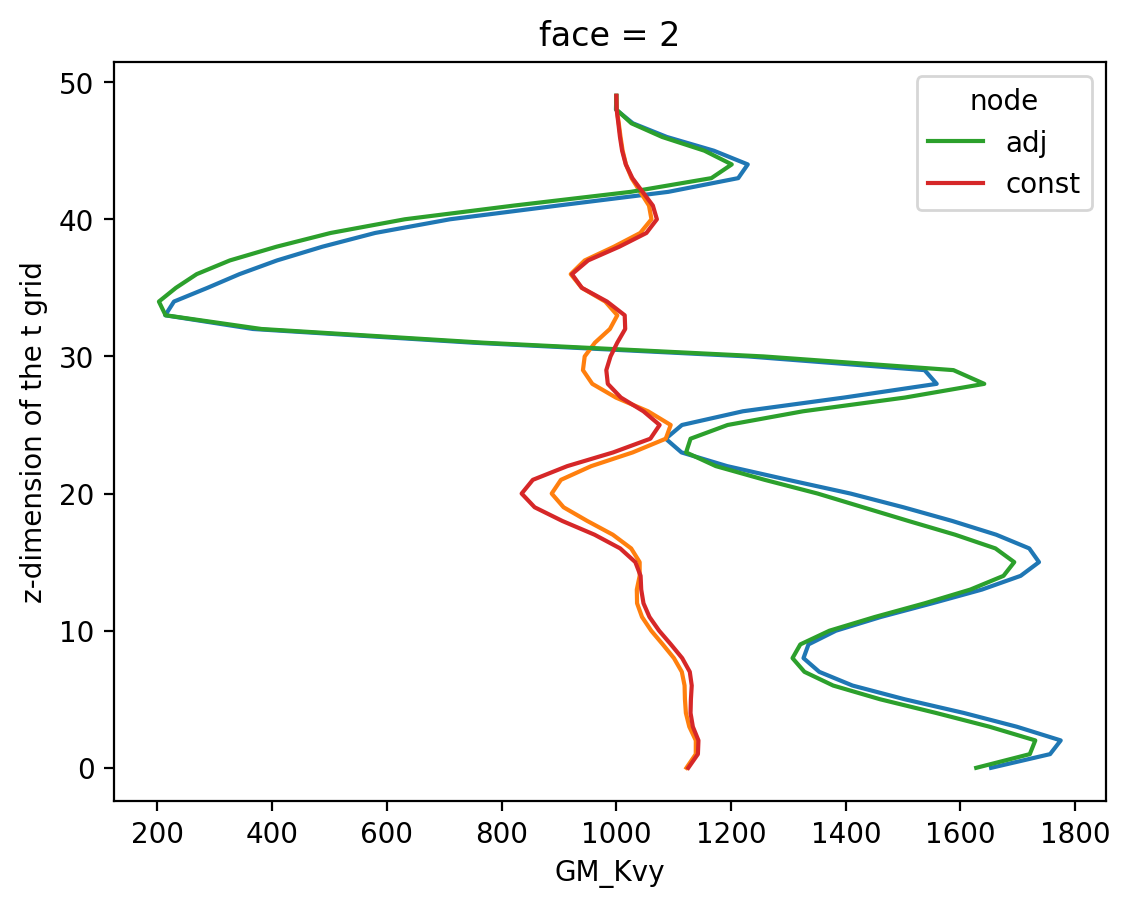
natre_mean.GM_Kvy.mean("time").cf.plot(hue="node")
[<matplotlib.lines.Line2D at 0x2abe4ea55c90>,
<matplotlib.lines.Line2D at 0x2abe4ea57160>]
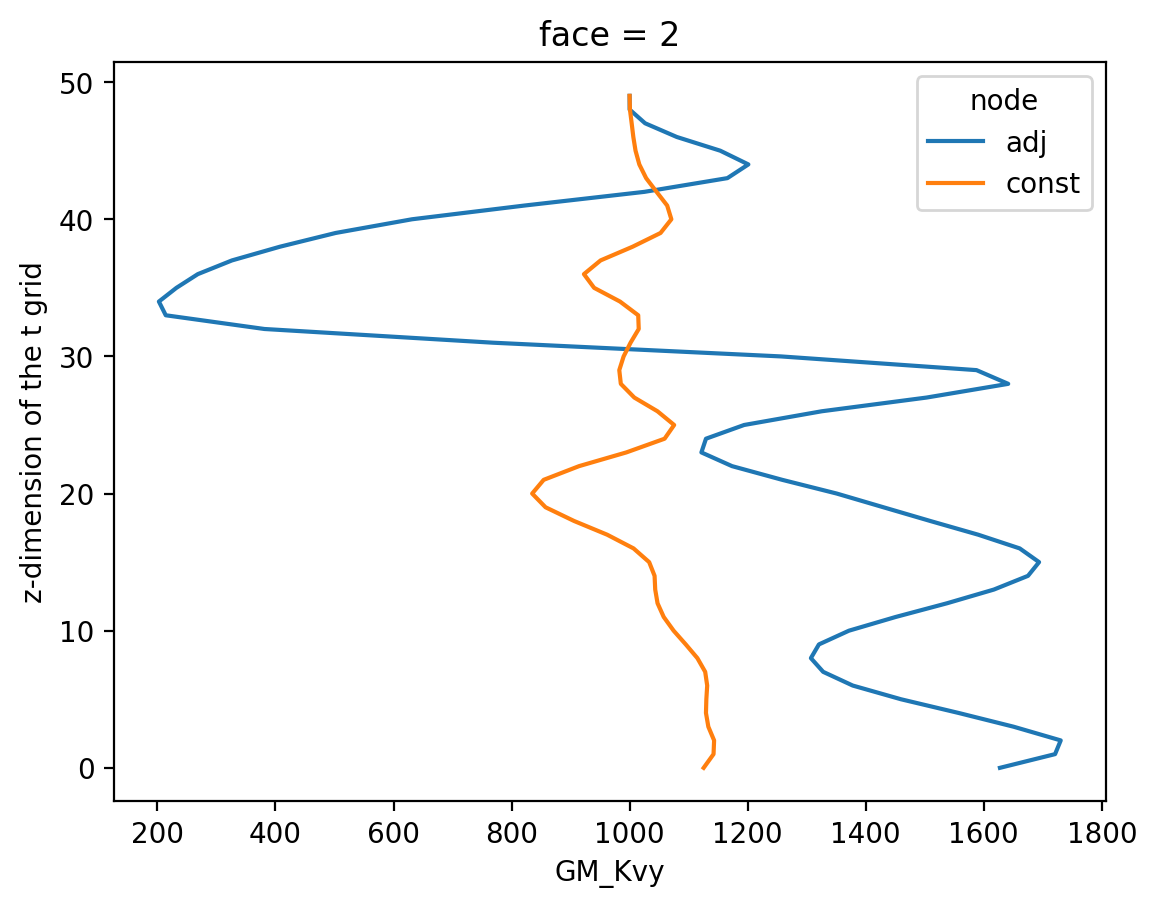
Dev#
dsfull, grid = open_directory("/glade/work/dcherian/mitgcm/ECCOV4/release4/run/diags/")
dsfull
ds = dsfull.sel(face=2).load()
/glade/u/home/dcherian/miniconda3/envs/eddydiff/lib/python3.10/site-packages/dask_jobqueue/core.py:255: FutureWarning: job_extra has been renamed to job_extra_directives. You are still using it (even if only set to []; please also check config files). If you did not set job_extra_directives yet, job_extra will be respected for now, but it will be removed in a future release. If you already set job_extra_directives, job_extra is ignored and you can remove it.
warnings.warn(warn, FutureWarning)
/glade/u/home/dcherian/miniconda3/envs/eddydiff/lib/python3.10/site-packages/dask_jobqueue/core.py:274: FutureWarning: env_extra has been renamed to job_script_prologue. You are still using it (even if only set to []; please also check config files). If you did not set job_script_prologue yet, env_extra will be respected for now, but it will be removed in a future release. If you already set job_script_prologue, env_extra is ignored and you can remove it.
warnings.warn(warn, FutureWarning)
NATRE region#
natre = ds.isel(i=slice(10, 18), j=slice(10, 20), i_g=slice(10, 18), j_g=slice(10, 20))
natre.load(scheduler=client)
<xarray.Dataset>
Dimensions: (i: 8, i_g: 8, j: 10, j_g: 10, k: 50, k_u: 50, k_l: 50,
k_p1: 51, time: 24)
Coordinates: (12/45)
* i (i) int64 10 11 12 13 14 15 16 17
* i_g (i_g) int64 10 11 12 13 14 15 16 17
* j (j) int64 10 11 12 13 14 15 16 17 18 19
* j_g (j_g) int64 10 11 12 13 14 15 16 17 18 19
* k (k) int64 0 1 2 3 4 5 6 7 8 9 ... 40 41 42 43 44 45 46 47 48 49
* k_u (k_u) int64 0 1 2 3 4 5 6 7 8 9 ... 40 41 42 43 44 45 46 47 48 49
... ...
maskCtrlS (k, j_g, i) bool True True True True ... False False False False
rhoRef (k) >f4 1.024e+03 1.024e+03 1.024e+03 ... 1.052e+03 1.054e+03
maskCtrlW (k, j, i_g) bool True True True True ... False False False False
iter (time) int64 1 2 3 4 5 6 7 8 9 10 ... 16 17 18 19 20 21 22 23 24
* time (time) timedelta64[ns] 00:00:01 00:00:02 ... 00:00:23 00:00:24
drC_kl (k_l) >f4 10.0 10.0 10.0 10.0 10.0 ... 399.0 422.0 445.0 228.2
Data variables: (12/19)
GM_CHIZ (time, k_l, j, i) float32 0.0 0.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0
GM_ubT (time, k, j, i_g) float32 1.363e+06 1.334e+06 ... 0.0 0.0
GM_CHIX (time, k, j, i_g) float32 1.571e-09 3.038e-09 ... 0.0 0.0
GM_Kvz (time, k, j_g, i) float32 3.523 3.411 3.31 3.206 ... 0.0 0.0 0.0
GM_Kwz (time, k_l, j, i) float32 0.0 0.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0
THETA (time, k, j, i) float32 23.1 22.98 22.81 22.6 ... 0.0 0.0 0.0 0.0
... ...
Tx (time, k, j, i_g) float32 -8.891e-07 -1.213e-06 ... 0.0 0.0
Ty (time, k, j_g, i) float32 -2.024e-06 -1.976e-06 ... 0.0 0.0
Tz (time, k_l, j, i) float32 nan nan nan nan nan ... 0.0 0.0 0.0 0.0
chix_ (time, k, j, i_g) float32 1.433e-09 2.585e-09 ... 0.0 0.0
chiy_ (time, k, j_g, i) float32 7.264e-09 6.708e-09 ... 0.0 0.0
GM_CHI (time, k, j, i) float32 1.467e-08 2.126e-08 2.9e-08 ... nan nan
Attributes:
Conventions: CF-1.6
title: netCDF wrapper of MITgcm MDS binary data
source: MITgcm
history: Created by calling `open_mdsdataset(grid_dir='/glade/work/d...xarray.Dataset
- i: 8
- i_g: 8
- j: 10
- j_g: 10
- k: 50
- k_u: 50
- k_l: 50
- k_p1: 51
- time: 24
- i(i)int6410 11 12 13 14 15 16 17
- standard_name :
- x_grid_index
- axis :
- X
- long_name :
- x-dimension of the t grid
- swap_dim :
- XC
array([10, 11, 12, 13, 14, 15, 16, 17])
- i_g(i_g)int6410 11 12 13 14 15 16 17
- standard_name :
- x_grid_index_at_u_location
- axis :
- X
- long_name :
- x-dimension of the u grid
- c_grid_axis_shift :
- -0.5
- swap_dim :
- XG
array([10, 11, 12, 13, 14, 15, 16, 17])
- j(j)int6410 11 12 13 14 15 16 17 18 19
- standard_name :
- y_grid_index
- axis :
- Y
- long_name :
- y-dimension of the t grid
- swap_dim :
- YC
array([10, 11, 12, 13, 14, 15, 16, 17, 18, 19])
- j_g(j_g)int6410 11 12 13 14 15 16 17 18 19
- standard_name :
- y_grid_index_at_v_location
- axis :
- Y
- long_name :
- y-dimension of the v grid
- c_grid_axis_shift :
- -0.5
- swap_dim :
- YG
array([10, 11, 12, 13, 14, 15, 16, 17, 18, 19])
- k(k)int640 1 2 3 4 5 6 ... 44 45 46 47 48 49
- standard_name :
- z_grid_index
- axis :
- Z
- long_name :
- z-dimension of the t grid
- swap_dim :
- Z
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49]) - k_u(k_u)int640 1 2 3 4 5 6 ... 44 45 46 47 48 49
- standard_name :
- z_grid_index_at_upper_w_location
- axis :
- Z
- long_name :
- z-dimension of the w grid
- c_grid_axis_shift :
- 0.5
- swap_dim :
- Zu
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49]) - k_l(k_l)int640 1 2 3 4 5 6 ... 44 45 46 47 48 49
- standard_name :
- z_grid_index_at_lower_w_location
- axis :
- Z
- long_name :
- z-dimension of the w grid
- c_grid_axis_shift :
- -0.5
- swap_dim :
- Zl
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49]) - k_p1(k_p1)int640 1 2 3 4 5 6 ... 45 46 47 48 49 50
- standard_name :
- z_grid_index_at_w_location
- axis :
- Z
- long_name :
- z-dimension of the w grid
- c_grid_axis_shift :
- (-0.5, 0.5)
- swap_dim :
- Zp1
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50]) - face()int642
- standard_name :
- face_index
array(2)
- XC(j, i)>f4-27.5 -26.5 -25.5 ... -21.5 -20.5
- standard_name :
- longitude
- long_name :
- longitude
- units :
- degrees_east
- coordinate :
- YC XC
array([[-27.5, -26.5, -25.5, -24.5, -23.5, -22.5, -21.5, -20.5], [-27.5, -26.5, -25.5, -24.5, -23.5, -22.5, -21.5, -20.5], [-27.5, -26.5, -25.5, -24.5, -23.5, -22.5, -21.5, -20.5], [-27.5, -26.5, -25.5, -24.5, -23.5, -22.5, -21.5, -20.5], [-27.5, -26.5, -25.5, -24.5, -23.5, -22.5, -21.5, -20.5], [-27.5, -26.5, -25.5, -24.5, -23.5, -22.5, -21.5, -20.5], [-27.5, -26.5, -25.5, -24.5, -23.5, -22.5, -21.5, -20.5], [-27.5, -26.5, -25.5, -24.5, -23.5, -22.5, -21.5, -20.5], [-27.5, -26.5, -25.5, -24.5, -23.5, -22.5, -21.5, -20.5], [-27.5, -26.5, -25.5, -24.5, -23.5, -22.5, -21.5, -20.5]], dtype=float32) - YC(j, i)>f420.12 20.12 20.12 ... 28.36 28.36
- standard_name :
- latitude
- long_name :
- latitude
- units :
- degrees_north
- coordinate :
- YC XC
array([[20.1248 , 20.1248 , 20.1248 , 20.1248 , 20.1248 , 20.1248 , 20.1248 , 20.1248 ], [21.065521, 21.065521, 21.065521, 21.065521, 21.065521, 21.065521, 21.065521, 21.065521], [22.00034 , 22.00034 , 22.00034 , 22.00034 , 22.00034 , 22.00034 , 22.00034 , 22.00034 ], [22.929024, 22.929024, 22.929024, 22.929024, 22.929024, 22.929024, 22.929024, 22.929024], [23.851347, 23.851347, 23.851347, 23.851347, 23.851347, 23.851347, 23.851347, 23.851347], [24.767094, 24.767094, 24.767094, 24.767094, 24.767094, 24.767094, 24.767094, 24.767094], [25.676054, 25.676054, 25.676054, 25.676054, 25.676054, 25.676054, 25.676054, 25.676054], [26.578028, 26.578028, 26.578028, 26.578028, 26.578028, 26.578028, 26.578028, 26.578028], [27.472822, 27.472822, 27.472822, 27.472822, 27.472822, 27.472822, 27.472822, 27.472822], [28.360258, 28.360258, 28.360258, 28.360258, 28.360258, 28.360258, 28.360258, 28.360258]], dtype=float32) - XG(j_g, i_g)>f4-28.0 -27.0 -26.0 ... -22.0 -21.0
- standard_name :
- longitude_at_f_location
- long_name :
- longitude
- units :
- degrees_east
- coordinate :
- YG XG
array([[-28., -27., -26., -25., -24., -23., -22., -21.], [-28., -27., -26., -25., -24., -23., -22., -21.], [-28., -27., -26., -25., -24., -23., -22., -21.], [-28., -27., -26., -25., -24., -23., -22., -21.], [-28., -27., -26., -25., -24., -23., -22., -21.], [-28., -27., -26., -25., -24., -23., -22., -21.], [-28., -27., -26., -25., -24., -23., -22., -21.], [-28., -27., -26., -25., -24., -23., -22., -21.], [-28., -27., -26., -25., -24., -23., -22., -21.], [-28., -27., -26., -25., -24., -23., -22., -21.]], dtype=float32) - YG(j_g, i_g)>f419.65 19.65 19.65 ... 27.92 27.92
- standard_name :
- latitude_at_f_location
- long_name :
- latitude
- units :
- degrees_north
- coordinate :
- YG XG
array([[19.652302, 19.652302, 19.652302, 19.652302, 19.652302, 19.652302, 19.652302, 19.652302], [20.595882, 20.595882, 20.595882, 20.595882, 20.595882, 20.595882, 20.595882, 20.595882], [21.533682, 21.533682, 21.533682, 21.533682, 21.533682, 21.533682, 21.533682, 21.533682], [22.465464, 22.465464, 22.465464, 22.465464, 22.465464, 22.465464, 22.465464, 22.465464], [23.390995, 23.390995, 23.390995, 23.390995, 23.390995, 23.390995, 23.390995, 23.390995], [24.310057, 24.310057, 24.310057, 24.310057, 24.310057, 24.310057, 24.310057, 24.310057], [25.222435, 25.222435, 25.222435, 25.222435, 25.222435, 25.222435, 25.222435, 25.222435], [26.127926, 26.127926, 26.127926, 26.127926, 26.127926, 26.127926, 26.127926, 26.127926], [27.026335, 27.026335, 27.026335, 27.026335, 27.026335, 27.026335, 27.026335, 27.026335], [27.91747 , 27.91747 , 27.91747 , 27.91747 , 27.91747 , 27.91747 , 27.91747 , 27.91747 ]], dtype=float32) - CS(j, i)>f41.0 1.0 1.0 1.0 ... 1.0 1.0 1.0 1.0
- standard_name :
- Cos of grid orientation angle
- long_name :
- AngleCS
- units :
- coordinate :
- YC XC
array([[1., 1., 1., 1., 1., 1., 1., 1.], [1., 1., 1., 1., 1., 1., 1., 1.], [1., 1., 1., 1., 1., 1., 1., 1.], [1., 1., 1., 1., 1., 1., 1., 1.], [1., 1., 1., 1., 1., 1., 1., 1.], [1., 1., 1., 1., 1., 1., 1., 1.], [1., 1., 1., 1., 1., 1., 1., 1.], [1., 1., 1., 1., 1., 1., 1., 1.], [1., 1., 1., 1., 1., 1., 1., 1.], [1., 1., 1., 1., 1., 1., 1., 1.]], dtype=float32) - SN(j, i)>f4-0.0 -0.0 ... 2.01e-15 -0.0
- standard_name :
- Sin of grid orientation angle
- long_name :
- AngleSN
- units :
- coordinate :
- YC XC
array([[-0.0000000e+00, -0.0000000e+00, 1.8976452e-15, -3.7724560e-15, 3.7724560e-15, -0.0000000e+00, -0.0000000e+00, -1.8976452e-15], [-0.0000000e+00, -0.0000000e+00, 1.8976452e-15, -0.0000000e+00, -0.0000000e+00, -1.9096464e-15, -0.0000000e+00, 1.2001251e-17], [-0.0000000e+00, -0.0000000e+00, -0.0000000e+00, -0.0000000e+00, -0.0000000e+00, -1.9096464e-15, -0.0000000e+00, 1.9096464e-15], [-0.0000000e+00, -0.0000000e+00, -0.0000000e+00, -0.0000000e+00, -0.0000000e+00, -0.0000000e+00, -0.0000000e+00, -0.0000000e+00], [-0.0000000e+00, -0.0000000e+00, -0.0000000e+00, -0.0000000e+00, -0.0000000e+00, -0.0000000e+00, -0.0000000e+00, -0.0000000e+00], [-0.0000000e+00, 1.9635623e-15, -1.9635623e-15, -0.0000000e+00, -0.0000000e+00, -0.0000000e+00, -0.0000000e+00, -0.0000000e+00], [-1.9785377e-15, 1.9635623e-15, -1.9635623e-15, -0.0000000e+00, -0.0000000e+00, -0.0000000e+00, -0.0000000e+00, -0.0000000e+00], [ 2.0097007e-15, -3.9882381e-15, 3.9882381e-15, -0.0000000e+00, -7.9764762e-15, -0.0000000e+00, 3.9882381e-15, -0.0000000e+00], [ 3.9882381e-15, -3.9882381e-15, 3.9882381e-15, -0.0000000e+00, -7.9764762e-15, -2.0103109e-15, 5.9985493e-15, -0.0000000e+00], [-0.0000000e+00, -0.0000000e+00, -0.0000000e+00, 2.0271176e-15, -2.0271176e-15, -2.0103109e-15, 2.0103109e-15, -0.0000000e+00]], dtype=float32) - Z(k)>f4-5.0 -15.0 ... -5.906e+03
- standard_name :
- depth
- long_name :
- vertical coordinate of cell center
- units :
- m
- positive :
- down
array([-5.000000e+00, -1.500000e+01, -2.500000e+01, -3.500000e+01, -4.500000e+01, -5.500000e+01, -6.500000e+01, -7.500500e+01, -8.502500e+01, -9.509500e+01, -1.053100e+02, -1.158700e+02, -1.271500e+02, -1.397400e+02, -1.544700e+02, -1.724000e+02, -1.947350e+02, -2.227100e+02, -2.574700e+02, -2.999300e+02, -3.506800e+02, -4.099300e+02, -4.774700e+02, -5.527100e+02, -6.347350e+02, -7.224000e+02, -8.144700e+02, -9.097400e+02, -1.007155e+03, -1.105905e+03, -1.205535e+03, -1.306205e+03, -1.409150e+03, -1.517095e+03, -1.634175e+03, -1.765135e+03, -1.914150e+03, -2.084035e+03, -2.276225e+03, -2.491250e+03, -2.729250e+03, -2.990250e+03, -3.274250e+03, -3.581250e+03, -3.911250e+03, -4.264250e+03, -4.640250e+03, -5.039250e+03, -5.461250e+03, -5.906250e+03], dtype=float32) - Zp1(k_p1)>f40.0 -10.0 ... -5.678e+03 -6.134e+03
- standard_name :
- depth_at_w_location
- long_name :
- vertical coordinate of cell interface
- units :
- m
- positive :
- down
array([ 0. , -10. , -20. , -30. , -40. , -50. , -60. , -70. , -80.01, -90.04, -100.15, -110.47, -121.27, -133.03, -146.45, -162.49, -182.31, -207.16, -238.26, -276.68, -323.18, -378.18, -441.68, -513.26, -592.16, -677.31, -767.49, -861.45, -958.03, -1056.28, -1155.53, -1255.54, -1356.87, -1461.43, -1572.76, -1695.59, -1834.68, -1993.62, -2174.45, -2378. , -2604.5 , -2854. , -3126.5 , -3422. , -3740.5 , -4082. , -4446.5 , -4834. , -5244.5 , -5678. , -6134.5 ], dtype=float32) - Zu(k_u)>f4-10.0 -20.0 ... -6.134e+03
- standard_name :
- depth_at_upper_w_location
- long_name :
- vertical coordinate of upper cell interface
- units :
- m
- positive :
- down
array([ -10. , -20. , -30. , -40. , -50. , -60. , -70. , -80.01, -90.04, -100.15, -110.47, -121.27, -133.03, -146.45, -162.49, -182.31, -207.16, -238.26, -276.68, -323.18, -378.18, -441.68, -513.26, -592.16, -677.31, -767.49, -861.45, -958.03, -1056.28, -1155.53, -1255.54, -1356.87, -1461.43, -1572.76, -1695.59, -1834.68, -1993.62, -2174.45, -2378. , -2604.5 , -2854. , -3126.5 , -3422. , -3740.5 , -4082. , -4446.5 , -4834. , -5244.5 , -5678. , -6134.5 ], dtype=float32) - Zl(k_l)>f40.0 -10.0 ... -5.244e+03 -5.678e+03
- standard_name :
- depth_at_lower_w_location
- long_name :
- vertical coordinate of lower cell interface
- units :
- m
- positive :
- down
array([ 0. , -10. , -20. , -30. , -40. , -50. , -60. , -70. , -80.01, -90.04, -100.15, -110.47, -121.27, -133.03, -146.45, -162.49, -182.31, -207.16, -238.26, -276.68, -323.18, -378.18, -441.68, -513.26, -592.16, -677.31, -767.49, -861.45, -958.03, -1056.28, -1155.53, -1255.54, -1356.87, -1461.43, -1572.76, -1695.59, -1834.68, -1993.62, -2174.45, -2378. , -2604.5 , -2854. , -3126.5 , -3422. , -3740.5 , -4082. , -4446.5 , -4834. , -5244.5 , -5678. ], dtype=float32) - rA(j, i)>f41.095e+10 1.095e+10 ... 9.612e+09
- standard_name :
- cell_area
- long_name :
- cell area
- units :
- m2
- coordinate :
- YC XC
array([[1.0950906e+10, 1.0950906e+10, 1.0950906e+10, 1.0950906e+10, 1.0950906e+10, 1.0950906e+10, 1.0950906e+10, 1.0950906e+10], [1.0816871e+10, 1.0816871e+10, 1.0816871e+10, 1.0816871e+10, 1.0816871e+10, 1.0816871e+10, 1.0816871e+10, 1.0816871e+10], [1.0678484e+10, 1.0678484e+10, 1.0678484e+10, 1.0678484e+10, 1.0678484e+10, 1.0678484e+10, 1.0678484e+10, 1.0678484e+10], [1.0536025e+10, 1.0536025e+10, 1.0536025e+10, 1.0536025e+10, 1.0536025e+10, 1.0536025e+10, 1.0536025e+10, 1.0536025e+10], [1.0389782e+10, 1.0389782e+10, 1.0389782e+10, 1.0389782e+10, 1.0389782e+10, 1.0389782e+10, 1.0389782e+10, 1.0389782e+10], [1.0240039e+10, 1.0240039e+10, 1.0240039e+10, 1.0240039e+10, 1.0240039e+10, 1.0240039e+10, 1.0240039e+10, 1.0240039e+10], [1.0087084e+10, 1.0087084e+10, 1.0087084e+10, 1.0087084e+10, 1.0087084e+10, 1.0087084e+10, 1.0087084e+10, 1.0087084e+10], [9.9312026e+09, 9.9312026e+09, 9.9312026e+09, 9.9312026e+09, 9.9312026e+09, 9.9312026e+09, 9.9312026e+09, 9.9312026e+09], [9.7726781e+09, 9.7726781e+09, 9.7726781e+09, 9.7726781e+09, 9.7726781e+09, 9.7726781e+09, 9.7726781e+09, 9.7726781e+09], [9.6117924e+09, 9.6117924e+09, 9.6117924e+09, 9.6117924e+09, 9.6117924e+09, 9.6117924e+09, 9.6117924e+09, 9.6117924e+09]], dtype=float32) - dxG(j_g, i)>f41.047e+05 1.047e+05 ... 9.824e+04
- standard_name :
- cell_x_size_at_v_location
- long_name :
- cell x size
- units :
- m
- coordinate :
- YG XC
array([[104701.48 , 104701.48 , 104701.48 , 104701.48 , 104701.48 , 104701.48 , 104701.48 , 104701.48 ], [104071.55 , 104071.55 , 104071.55 , 104071.55 , 104071.55 , 104071.55 , 104071.55 , 104071.55 ], [103417.5 , 103417.5 , 103417.5 , 103417.5 , 103417.5 , 103417.5 , 103417.5 , 103417.5 ], [102740.22 , 102740.22 , 102740.22 , 102740.22 , 102740.22 , 102740.22 , 102740.22 , 102740.22 ], [102040.58 , 102040.58 , 102040.58 , 102040.58 , 102040.58 , 102040.58 , 102040.58 , 102040.58 ], [101319.484, 101319.484, 101319.484, 101319.484, 101319.484, 101319.484, 101319.484, 101319.484], [100577.84 , 100577.84 , 100577.84 , 100577.84 , 100577.84 , 100577.84 , 100577.84 , 100577.84 ], [ 99816.586, 99816.586, 99816.586, 99816.586, 99816.586, 99816.586, 99816.586, 99816.586], [ 99036.65 , 99036.65 , 99036.65 , 99036.65 , 99036.65 , 99036.65 , 99036.65 , 99036.65 ], [ 98238.96 , 98238.96 , 98238.96 , 98238.96 , 98238.96 , 98238.96 , 98238.96 , 98238.96 ]], dtype=float32) - dyG(j, i_g)>f41.049e+05 1.049e+05 ... 9.825e+04
- standard_name :
- cell_y_size_at_u_location
- long_name :
- cell y size
- units :
- m
- coordinate :
- YC XG
array([[104904.92 , 104904.92 , 104904.92 , 104904.92 , 104904.92 , 104904.92 , 104904.92 , 104904.92 ], [104262.18 , 104262.18 , 104262.18 , 104262.18 , 104262.18 , 104262.18 , 104262.18 , 104262.18 ], [103593. , 103593. , 103593. , 103593. , 103593. , 103593. , 103593. , 103593. ], [102898.28 , 102898.28 , 102898.28 , 102898.28 , 102898.28 , 102898.28 , 102898.28 , 102898.28 ], [102178.95 , 102178.95 , 102178.95 , 102178.95 , 102178.95 , 102178.95 , 102178.95 , 102178.95 ], [101435.945, 101435.945, 101435.945, 101435.945, 101435.945, 101435.945, 101435.945, 101435.945], [100670.21 , 100670.21 , 100670.21 , 100670.21 , 100670.21 , 100670.21 , 100670.21 , 100670.21 ], [ 99882.7 , 99882.7 , 99882.7 , 99882.7 , 99882.7 , 99882.7 , 99882.7 , 99882.7 ], [ 99074.41 , 99074.41 , 99074.41 , 99074.41 , 99074.41 , 99074.41 , 99074.41 , 99074.41 ], [ 98246.27 , 98246.27 , 98246.27 , 98246.27 , 98246.27 , 98246.27 , 98246.27 , 98246.27 ]], dtype=float32) - Depth(j, i)>f44.834e+03 4.755e+03 ... 4.639e+03
- standard_name :
- ocean_depth
- long_name :
- ocean depth
- units :
- m
- coordinate :
- XC YC
array([[4834. , 4755.442 , 4592.7134, 4446.5 , 4316.972 , 4082. , 3867.3867, 3656.9805], [5119.3086, 5045.3174, 4916.97 , 4797.1206, 4644.9146, 4441.4663, 4213.616 , 3948.209 ], [5353.368 , 5244.5 , 5145.085 , 5022.9175, 4834. , 4634.7456, 4395.123 , 4082. ], [5490.0015, 5373.423 , 5244.5 , 5125.006 , 4945.592 , 4719.215 , 4443.8135, 4082. ], [5537.9336, 5417.104 , 5331.2 , 5156.198 , 4984.341 , 4764.6167, 4446.5 , 3940.8796], [5484.422 , 5372.304 , 5244.5 , 5166.3994, 5016.643 , 4814.217 , 4533.3965, 4081.4624], [5331.2 , 5176.734 , 5146.98 , 5145.518 , 5038.288 , 4834. , 4629.338 , 4207.3945], [5157.1235, 5103.156 , 5145.147 , 5137.3247, 5038.491 , 4916.1 , 4711.077 , 4375.7227], [5154.945 , 5169.684 , 5202.6797, 5164.4927, 5048.946 , 4917.9297, 4771.9316, 4542.9316], [5099.8784, 5193.563 , 5244.5 , 5216.8755, 5086.396 , 4948.9214, 4815.4067, 4638.548 ]], dtype=float32) - rAz(j_g, i_g)>f41.102e+10 1.102e+10 ... 9.693e+09
- standard_name :
- cell_area_at_f_location
- long_name :
- cell area
- units :
- m
- coordinate :
- YG XG
array([[1.1016203e+10, 1.1016203e+10, 1.1016203e+10, 1.1016203e+10, 1.1016203e+10, 1.1016203e+10, 1.1016203e+10, 1.1016203e+10], [1.0884450e+10, 1.0884450e+10, 1.0884450e+10, 1.0884450e+10, 1.0884450e+10, 1.0884450e+10, 1.0884450e+10, 1.0884450e+10], [1.0748204e+10, 1.0748204e+10, 1.0748204e+10, 1.0748204e+10, 1.0748204e+10, 1.0748204e+10, 1.0748204e+10, 1.0748204e+10], [1.0607745e+10, 1.0607745e+10, 1.0607745e+10, 1.0607745e+10, 1.0607745e+10, 1.0607745e+10, 1.0607745e+10, 1.0607745e+10], [1.0463359e+10, 1.0463359e+10, 1.0463359e+10, 1.0463359e+10, 1.0463359e+10, 1.0463359e+10, 1.0463359e+10, 1.0463359e+10], [1.0315330e+10, 1.0315330e+10, 1.0315330e+10, 1.0315330e+10, 1.0315330e+10, 1.0315330e+10, 1.0315330e+10, 1.0315330e+10], [1.0163945e+10, 1.0163945e+10, 1.0163945e+10, 1.0163945e+10, 1.0163945e+10, 1.0163945e+10, 1.0163945e+10, 1.0163945e+10], [1.0009491e+10, 1.0009491e+10, 1.0009491e+10, 1.0009491e+10, 1.0009491e+10, 1.0009491e+10, 1.0009491e+10, 1.0009491e+10], [9.8522532e+09, 9.8522532e+09, 9.8522532e+09, 9.8522532e+09, 9.8522532e+09, 9.8522532e+09, 9.8522532e+09, 9.8522532e+09], [9.6925133e+09, 9.6925133e+09, 9.6925133e+09, 9.6925133e+09, 9.6925133e+09, 9.6925133e+09, 9.6925133e+09, 9.6925133e+09]], dtype=float32) - dxC(j, i_g)>f41.044e+05 1.044e+05 ... 9.783e+04
- standard_name :
- cell_x_size_at_u_location
- long_name :
- cell x size
- units :
- m
- coordinate :
- YC XG
array([[104389.58 , 104389.58 , 104389.58 , 104389.58 , 104389.58 , 104389.58 , 104389.58 , 104389.58 ], [103747.484, 103747.484, 103747.484, 103747.484, 103747.484, 103747.484, 103747.484, 103747.484], [103081.71 , 103081.71 , 103081.71 , 103081.71 , 103081.71 , 103081.71 , 103081.71 , 103081.71 ], [102393.14 , 102393.14 , 102393.14 , 102393.14 , 102393.14 , 102393.14 , 102393.14 , 102393.14 ], [101682.66 , 101682.66 , 101682.66 , 101682.66 , 101682.66 , 101682.66 , 101682.66 , 101682.66 ], [100951.17 , 100951.17 , 100951.17 , 100951.17 , 100951.17 , 100951.17 , 100951.17 , 100951.17 ], [100199.61 , 100199.61 , 100199.61 , 100199.61 , 100199.61 , 100199.61 , 100199.61 , 100199.61 ], [ 99428.89 , 99428.89 , 99428.89 , 99428.89 , 99428.89 , 99428.89 , 99428.89 , 99428.89 ], [ 98639.96 , 98639.96 , 98639.96 , 98639.96 , 98639.96 , 98639.96 , 98639.96 , 98639.96 ], [ 97833.76 , 97833.76 , 97833.76 , 97833.76 , 97833.76 , 97833.76 , 97833.76 , 97833.76 ]], dtype=float32) - dyC(j_g, i)>f41.052e+05 1.052e+05 ... 9.866e+04
- standard_name :
- cell_y_size_at_v_location
- long_name :
- cell y size
- units :
- m
- coordinate :
- YG XC
array([[105216.09 , 105216.09 , 105216.09 , 105216.09 , 105216.09 , 105216.09 , 105216.09 , 105216.09 ], [104586.914, 104586.914, 104586.914, 104586.914, 104586.914, 104586.914, 104586.914, 104586.914], [103930.836, 103930.836, 103930.836, 103930.836, 103930.836, 103930.836, 103930.836, 103930.836], [103248.77 , 103248.77 , 103248.77 , 103248.77 , 103248.77 , 103248.77 , 103248.77 , 103248.77 ], [102541.63 , 102541.63 , 102541.63 , 102541.63 , 102541.63 , 102541.63 , 102541.63 , 102541.63 ], [101810.35 , 101810.35 , 101810.35 , 101810.35 , 101810.35 , 101810.35 , 101810.35 , 101810.35 ], [101055.86 , 101055.86 , 101055.86 , 101055.86 , 101055.86 , 101055.86 , 101055.86 , 101055.86 ], [100279.12 , 100279.12 , 100279.12 , 100279.12 , 100279.12 , 100279.12 , 100279.12 , 100279.12 ], [ 99481.09 , 99481.09 , 99481.09 , 99481.09 , 99481.09 , 99481.09 , 99481.09 , 99481.09 ], [ 98662.76 , 98662.76 , 98662.76 , 98662.76 , 98662.76 , 98662.76 , 98662.76 , 98662.76 ]], dtype=float32) - rAw(j, i_g)>f41.095e+10 1.095e+10 ... 9.612e+09
- standard_name :
- cell_area_at_u_location
- long_name :
- cell area
- units :
- m2
- coordinate :
- YG XC
array([[1.0950906e+10, 1.0950906e+10, 1.0950906e+10, 1.0950906e+10, 1.0950906e+10, 1.0950906e+10, 1.0950906e+10, 1.0950906e+10], [1.0816871e+10, 1.0816871e+10, 1.0816871e+10, 1.0816871e+10, 1.0816871e+10, 1.0816871e+10, 1.0816871e+10, 1.0816871e+10], [1.0678484e+10, 1.0678484e+10, 1.0678484e+10, 1.0678484e+10, 1.0678484e+10, 1.0678484e+10, 1.0678484e+10, 1.0678484e+10], [1.0536025e+10, 1.0536025e+10, 1.0536025e+10, 1.0536025e+10, 1.0536025e+10, 1.0536025e+10, 1.0536025e+10, 1.0536025e+10], [1.0389782e+10, 1.0389782e+10, 1.0389782e+10, 1.0389782e+10, 1.0389782e+10, 1.0389782e+10, 1.0389782e+10, 1.0389782e+10], [1.0240039e+10, 1.0240039e+10, 1.0240039e+10, 1.0240039e+10, 1.0240039e+10, 1.0240039e+10, 1.0240039e+10, 1.0240039e+10], [1.0087084e+10, 1.0087084e+10, 1.0087084e+10, 1.0087084e+10, 1.0087084e+10, 1.0087084e+10, 1.0087084e+10, 1.0087084e+10], [9.9312026e+09, 9.9312026e+09, 9.9312026e+09, 9.9312026e+09, 9.9312026e+09, 9.9312026e+09, 9.9312026e+09, 9.9312026e+09], [9.7726781e+09, 9.7726781e+09, 9.7726781e+09, 9.7726781e+09, 9.7726781e+09, 9.7726781e+09, 9.7726781e+09, 9.7726781e+09], [9.6117924e+09, 9.6117924e+09, 9.6117924e+09, 9.6117924e+09, 9.6117924e+09, 9.6117924e+09, 9.6117924e+09, 9.6117924e+09]], dtype=float32) - rAs(j_g, i)>f41.102e+10 1.102e+10 ... 9.693e+09
- standard_name :
- cell_area_at_v_location
- long_name :
- cell area
- units :
- m2
- coordinate :
- YG XC
array([[1.1016203e+10, 1.1016203e+10, 1.1016203e+10, 1.1016203e+10, 1.1016203e+10, 1.1016203e+10, 1.1016203e+10, 1.1016203e+10], [1.0884450e+10, 1.0884450e+10, 1.0884450e+10, 1.0884450e+10, 1.0884450e+10, 1.0884450e+10, 1.0884450e+10, 1.0884450e+10], [1.0748204e+10, 1.0748204e+10, 1.0748204e+10, 1.0748204e+10, 1.0748204e+10, 1.0748204e+10, 1.0748204e+10, 1.0748204e+10], [1.0607745e+10, 1.0607745e+10, 1.0607745e+10, 1.0607745e+10, 1.0607745e+10, 1.0607745e+10, 1.0607745e+10, 1.0607745e+10], [1.0463359e+10, 1.0463359e+10, 1.0463359e+10, 1.0463359e+10, 1.0463359e+10, 1.0463359e+10, 1.0463359e+10, 1.0463359e+10], [1.0315330e+10, 1.0315330e+10, 1.0315330e+10, 1.0315330e+10, 1.0315330e+10, 1.0315330e+10, 1.0315330e+10, 1.0315330e+10], [1.0163945e+10, 1.0163945e+10, 1.0163945e+10, 1.0163945e+10, 1.0163945e+10, 1.0163945e+10, 1.0163945e+10, 1.0163945e+10], [1.0009491e+10, 1.0009491e+10, 1.0009491e+10, 1.0009491e+10, 1.0009491e+10, 1.0009491e+10, 1.0009491e+10, 1.0009491e+10], [9.8522532e+09, 9.8522532e+09, 9.8522532e+09, 9.8522532e+09, 9.8522532e+09, 9.8522532e+09, 9.8522532e+09, 9.8522532e+09], [9.6925133e+09, 9.6925133e+09, 9.6925133e+09, 9.6925133e+09, 9.6925133e+09, 9.6925133e+09, 9.6925133e+09, 9.6925133e+09]], dtype=float32) - drC(k_p1)>f45.0 10.0 10.0 ... 422.0 445.0 228.2
- standard_name :
- cell_z_size_at_w_location
- long_name :
- cell z size
- units :
- m
array([ 5. , 10. , 10. , 10. , 10. , 10. , 10. , 10.005, 10.02 , 10.07 , 10.215, 10.56 , 11.28 , 12.59 , 14.73 , 17.93 , 22.335, 27.975, 34.76 , 42.46 , 50.75 , 59.25 , 67.54 , 75.24 , 82.025, 87.665, 92.07 , 95.27 , 97.415, 98.75 , 99.63 , 100.67 , 102.945, 107.945, 117.08 , 130.96 , 149.015, 169.885, 192.19 , 215.025, 238. , 261. , 284. , 307. , 330. , 353. , 376. , 399. , 422. , 445. , 228.25 ], dtype=float32) - drF(k)>f410.0 10.0 10.0 ... 433.5 456.5
- standard_name :
- cell_z_size
- long_name :
- cell z size
- units :
- m
array([ 10. , 10. , 10. , 10. , 10. , 10. , 10. , 10.01, 10.03, 10.11, 10.32, 10.8 , 11.76, 13.42, 16.04, 19.82, 24.85, 31.1 , 38.42, 46.5 , 55. , 63.5 , 71.58, 78.9 , 85.15, 90.18, 93.96, 96.58, 98.25, 99.25, 100.01, 101.33, 104.56, 111.33, 122.83, 139.09, 158.94, 180.83, 203.55, 226.5 , 249.5 , 272.5 , 295.5 , 318.5 , 341.5 , 364.5 , 387.5 , 410.5 , 433.5 , 456.5 ], dtype=float32) - PHrefC(k)>f449.05 147.1 ... 5.357e+04 5.794e+04
- standard_name :
- cell_reference_pressure
- long_name :
- Reference Hydrostatic Pressure
- units :
- m2 s-2
array([4.9049999e+01, 1.4714999e+02, 2.4525000e+02, 3.4335001e+02, 4.4145001e+02, 5.3954999e+02, 6.3765002e+02, 7.3579907e+02, 8.3409528e+02, 9.3288196e+02, 1.0330911e+03, 1.1366847e+03, 1.2473416e+03, 1.3708494e+03, 1.5153507e+03, 1.6912440e+03, 1.9103503e+03, 2.1847852e+03, 2.5257808e+03, 2.9423132e+03, 3.4401709e+03, 4.0214133e+03, 4.6839805e+03, 5.4220850e+03, 6.2267505e+03, 7.0867441e+03, 7.9899507e+03, 8.9245498e+03, 9.8801904e+03, 1.0848928e+04, 1.1826299e+04, 1.2813871e+04, 1.3823762e+04, 1.4882702e+04, 1.6031257e+04, 1.7315975e+04, 1.8777811e+04, 2.0444383e+04, 2.2329768e+04, 2.4439162e+04, 2.6773943e+04, 2.9334352e+04, 3.2120393e+04, 3.5132062e+04, 3.8369363e+04, 4.1832293e+04, 4.5520852e+04, 4.9435043e+04, 5.3574863e+04, 5.7940312e+04], dtype=float32) - PHrefF(k_p1)>f40.0 98.1 ... 5.57e+04 6.018e+04
- standard_name :
- cell_reference_pressure
- long_name :
- Reference Hydrostatic Pressure
- units :
- m2 s-2
array([ 0. , 98.1 , 196.2 , 294.3 , 392.4 , 490.5 , 588.6 , 686.7 , 784.8981, 883.2924, 982.4715, 1083.7107, 1189.6587, 1305.0243, 1436.6746, 1594.0269, 1788.461 , 2032.2396, 2337.3306, 2714.2307, 3170.3958, 3709.9458, 4332.881 , 5035.0806, 5809.09 , 6644.411 , 7529.0767, 8450.824 , 9398.274 , 10362.106 , 11335.749 , 12316.848 , 13310.895 , 14336.628 , 15428.775 , 16633.738 , 17998.21 , 19557.412 , 21331.355 , 23328.18 , 25550.145 , 27997.74 , 30670.965 , 33569.82 , 36694.305 , 40044.42 , 43620.164 , 47421.54 , 51448.547 , 55701.18 , 60179.445 ], dtype=float32) - hFacC(k, j, i)>f41.0 1.0 1.0 1.0 ... 0.0 0.0 0.0 0.0
- standard_name :
- cell_vertical_fraction
- long_name :
- vertical fraction of open cell
array([[[1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ], ..., [1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ]], [[1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ], ... [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ]], [[0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], ..., [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ]]], dtype=float32) - hFacW(k, j, i_g)>f41.0 1.0 1.0 1.0 ... 0.0 0.0 0.0 0.0
- standard_name :
- cell_vertical_fraction_at_u_location
- long_name :
- vertical fraction of open cell
array([[[1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ], ..., [1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ]], [[1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ], ... [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ]], [[0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], ..., [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ]]], dtype=float32) - hFacS(k, j_g, i)>f41.0 1.0 1.0 1.0 ... 0.0 0.0 0.0 0.0
- standard_name :
- cell_vertical_fraction_at_v_location
- long_name :
- vertical fraction of open cell
array([[[1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ], ..., [1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ]], [[1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ], ... [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ]], [[0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], ..., [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ]]], dtype=float32) - maskC(k, j, i)boolTrue True True ... False False
- standard_name :
- sea_binary_mask_at_t_location
- long_name :
- mask denoting wet point at center
array([[[ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], ..., [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True]], [[ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], ..., [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True]], [[ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], ..., ... ..., [ True, True, True, ..., True, False, False], [ True, True, True, ..., True, False, False], [ True, True, True, ..., True, False, False]], [[False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [ True, False, False, ..., False, False, False], ..., [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False]], [[False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], ..., [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False]]]) - maskW(k, j, i_g)boolTrue True True ... False False
- standard_name :
- cell_vertical_fraction_at_u_location
- long_name :
- mask denoting wet point at interface
array([[[ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], ..., [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True]], [[ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], ..., [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True]], [[ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], ..., ... ..., [ True, True, True, ..., True, False, False], [ True, True, True, ..., True, False, False], [ True, True, True, ..., True, False, False]], [[False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [ True, False, False, ..., False, False, False], ..., [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False]], [[False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], ..., [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False]]]) - maskS(k, j_g, i)boolTrue True True ... False False
- standard_name :
- cell_vertical_fraction_at_v_location
- long_name :
- mask denoting wet point at interface
array([[[ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], ..., [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True]], [[ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], ..., [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True]], [[ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], ..., ... ..., [ True, True, True, ..., False, False, False], [ True, True, True, ..., True, False, False], [ True, True, True, ..., True, False, False]], [[False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], ..., [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False]], [[False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], ..., [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False]]]) - maskCtrlC(k, j, i)boolTrue True True ... False False
- standard_name :
- ctrl_vector_3d_mask
- long_name :
- CTRL 3D mask where ctrl vector is active at tracer location
- units :
array([[[ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], ..., [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True]], [[ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], ..., [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True]], [[ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], ..., ... ..., [ True, True, True, ..., True, False, False], [ True, True, True, ..., True, False, False], [ True, True, True, ..., True, False, False]], [[False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [ True, False, False, ..., False, False, False], ..., [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False]], [[False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], ..., [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False]]]) - maskCtrlS(k, j_g, i)boolTrue True True ... False False
- standard_name :
- ctrl_vector_3d_mask_at_v_location
- long_name :
- CTRL 3D mask where ctrl vector is active at v location
- units :
array([[[ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], ..., [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True]], [[ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], ..., [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True]], [[ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], ..., ... ..., [ True, True, True, ..., False, False, False], [ True, True, True, ..., True, False, False], [ True, True, True, ..., True, False, False]], [[False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], ..., [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False]], [[False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], ..., [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False]]]) - rhoRef(k)>f41.024e+03 1.024e+03 ... 1.054e+03
- standard_name :
- reference_density_profile
- long_name :
- 1D, vertical reference density profile
- coordinate :
- Z
- units :
- kg m-3
array([1023.5776 , 1023.6208 , 1023.66394, 1023.991 , 1024.0343 , 1024.0776 , 1024.3971 , 1024.7086 , 1024.7523 , 1025.0563 , 1025.3527 , 1025.3989 , 1025.6919 , 1025.9822 , 1026.0472 , 1026.3529 , 1026.6698 , 1027.0033 , 1027.3589 , 1027.7407 , 1028.3246 , 1028.5933 , 1029.0645 , 1029.5631 , 1030.0851 , 1030.4861 , 1031.0481 , 1031.4844 , 1032.0652 , 1032.518 , 1032.9736 , 1033.5654 , 1034.0369 , 1034.53 , 1035.1935 , 1035.7921 , 1036.471 , 1037.2421 , 1038.2471 , 1039.2203 , 1040.2919 , 1041.4601 , 1042.7234 , 1044.0795 , 1045.5265 , 1047.0621 , 1048.6838 , 1050.3894 , 1052.1759 , 1054.0406 ], dtype=float32) - maskCtrlW(k, j, i_g)boolTrue True True ... False False
- standard_name :
- ctrl_vector_3d_mask_at_u_location
- long_name :
- CTRL 3D mask where ctrl vector is active at u location
- units :
array([[[ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], ..., [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True]], [[ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], ..., [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True]], [[ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], ..., ... ..., [ True, True, True, ..., True, False, False], [ True, True, True, ..., True, False, False], [ True, True, True, ..., True, False, False]], [[False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [ True, False, False, ..., False, False, False], ..., [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False]], [[False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], ..., [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False]]]) - iter(time)int641 2 3 4 5 6 7 ... 19 20 21 22 23 24
- standard_name :
- timestep
- long_name :
- model timestep number
array([ 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24]) - time(time)timedelta64[ns]00:00:01 00:00:02 ... 00:00:24
- standard_name :
- time
- long_name :
- Time
- axis :
- T
- calendar :
- gregorian
array([ 1000000000, 2000000000, 3000000000, 4000000000, 5000000000, 6000000000, 7000000000, 8000000000, 9000000000, 10000000000, 11000000000, 12000000000, 13000000000, 14000000000, 15000000000, 16000000000, 17000000000, 18000000000, 19000000000, 20000000000, 21000000000, 22000000000, 23000000000, 24000000000], dtype='timedelta64[ns]') - drC_kl(k_l)>f410.0 10.0 10.0 ... 445.0 228.2
- standard_name :
- cell_z_size_at_w_location
- long_name :
- cell z size
- units :
- m
array([ 10. , 10. , 10. , 10. , 10. , 10. , 10.005, 10.02 , 10.07 , 10.215, 10.56 , 11.28 , 12.59 , 14.73 , 17.93 , 22.335, 27.975, 34.76 , 42.46 , 50.75 , 59.25 , 67.54 , 75.24 , 82.025, 87.665, 92.07 , 95.27 , 97.415, 98.75 , 99.63 , 100.67 , 102.945, 107.945, 117.08 , 130.96 , 149.015, 169.885, 192.19 , 215.025, 238. , 261. , 284. , 307. , 330. , 353. , 376. , 399. , 422. , 445. , 228.25 ], dtype=float32)
- GM_CHIZ(time, k_l, j, i)float320.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0
- standard_name :
- GM_CHIZ
- long_name :
- Redi Temp variance dissipation rate: Z component
- units :
- degC^2/s
array([[[[0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], ..., [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00]], [[8.96746544e-13, 1.74780904e-12, 2.99178694e-12, ..., 6.53042898e-12, 6.58428477e-12, 6.73465537e-12], [1.61002445e-09, 2.05971613e-08, 1.36162739e-12, ..., 3.96390819e-12, 4.39305927e-12, 3.34672611e-12], [1.66915592e-10, 1.18665799e-09, 1.57606941e-08, ..., 2.20751304e-12, 2.43433550e-12, 1.74343537e-12], ... 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00]], [[0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], ..., [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00]]]], dtype=float32) - GM_ubT(time, k, j, i_g)float321.363e+06 1.334e+06 ... 0.0 0.0
- standard_name :
- GM_ubT
- long_name :
- Zonal Mass-Weight Bolus Transp of Pot Temp
- units :
- degC.m^3/s
- mate :
- GM_vbT
array([[[[ 1362986.2 , 1334435.9 , 1308713.1 , ..., 1250643.4 , 1246402.9 , 1252541.9 ], [ 1321090.9 , 1320729.5 , 1296473.9 , ..., 1246273.6 , 1237552.4 , 1232376.6 ], [ 213291.34 , 201504.22 , 448878.4 , ..., 1219688.8 , 1209364.6 , 1198211. ], ..., [ 999204.3 , 987998.5 , 730655.3 , ..., -102951.016 , -138122.64 , -36039.906 ], [ 973194.44 , 967057.94 , 965877.06 , ..., 203785.48 , 63781.824 , 28307.82 ], [ 970884.06 , 967982.1 , 969262.7 , ..., 664230.3 , 436563.97 , 360388.2 ]], [[ 106500.13 , 97531.35 , 84569.46 , ..., 7919.1636, -25316.396 , -57220.918 ], [ 122870.7 , 82541.77 , 70514.75 , ..., 3268.195 , -28708. , -60971.043 ], [ -60300.562 , -290542.1 , 863342.94 , ..., -10604.97 , -40564.496 , -71992.29 ], ... [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ]], [[ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], ..., [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ]]]], dtype=float32) - GM_CHIX(time, k, j, i_g)float321.571e-09 3.038e-09 ... 0.0 0.0
- standard_name :
- GM_CHIX
- long_name :
- Redi Temp variance dissipation rate: X component
- units :
- degC^2/s
- mate :
- GM_KwzTz
array([[[[ 1.57073354e-09, 3.03783954e-09, 5.24529664e-09, ..., 2.16047233e-08, 3.91388397e-08, 6.02352230e-08], [ 1.69642900e-09, 2.48811260e-09, 3.93770172e-09, ..., 2.76667400e-08, 4.94266494e-08, 7.34814236e-08], [ 1.88299598e-09, 2.30834374e-09, 2.90634006e-09, ..., 1.53638222e-08, 3.26456018e-08, 5.64273996e-08], ..., [-1.51910184e-09, -1.04408793e-09, -3.61996044e-10, ..., 2.05752193e-09, 2.46603826e-09, 2.09546003e-09], [-2.41093456e-09, -1.86043358e-09, -1.34118749e-09, ..., 9.81380532e-10, 1.55818902e-09, 1.98274885e-09], [-3.06394954e-09, -2.54115373e-09, -2.23318142e-09, ..., -2.35179903e-10, 4.75803397e-10, 1.00445785e-09]], [[ 1.55616176e-09, 3.04793701e-09, 5.15399634e-09, ..., 2.82235266e-08, 5.83077018e-08, 9.72858558e-08], [ 1.62642921e-09, 2.39364528e-09, 3.79050302e-09, ..., 4.05912388e-08, 7.62354588e-08, 1.12844020e-07], [ 1.92149541e-09, 2.34271091e-09, 2.88671709e-09, ..., 2.17586766e-08, 4.80109463e-08, 8.01088973e-08], ... 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00]], [[ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], ..., [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00]]]], dtype=float32) - GM_Kvz(time, k, j_g, i)float323.523 3.411 3.31 ... 0.0 0.0 0.0
- standard_name :
- GM_Kvz
- long_name :
- K_23 element (V.point, Z.dir) of GM-Redi tensor
- units :
- m^2/s
- mate :
- GM_Kuz
array([[[[ 3.5234537 , 3.4112551 , 3.3099675 , ..., 2.958492 , 2.8169467 , 2.7696686 ], [ 3.5952473 , 3.4734573 , 3.3450308 , ..., 2.8282216 , 2.6789253 , 2.627114 ], [ 3.6615362 , 3.5277123 , 3.3912249 , ..., 2.6320338 , 2.425822 , 2.3633146 ], ..., [ 1.2037884 , 1.2813897 , 1.3859956 , ..., 1.4870073 , 1.4610972 , 1.2866143 ], [ 0.5592338 , 0.6360955 , 0.73997337, ..., 1.0285405 , 1.0867356 , 1.1140348 ], [ 0.0519005 , 0.11499736, 0.20792365, ..., 0.6229226 , 0.73785686, 0.8338319 ]], [[ 3.7016644 , 3.581409 , 3.4601724 , ..., 3.0268915 , 2.862994 , 2.788607 ], [ 3.791752 , 3.6512969 , 3.503654 , ..., 2.9126794 , 2.7285984 , 2.640672 ], [ 3.8879032 , 3.729321 , 3.5709429 , ..., 2.7448974 , 2.499652 , 2.3952727 ], ... [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ]], [[ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], ..., [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ]]]], dtype=float32) - GM_Kwz(time, k_l, j, i)float320.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0
- standard_name :
- GM_Kwz
- long_name :
- K_33 element (W.point, Z.dir) of GM-Redi tensor
- units :
- m^2/s
array([[[[0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], ..., [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00]], [[7.32346531e-03, 7.07426155e-03, 6.81576831e-03, ..., 6.03489019e-03, 5.84230386e-03, 5.72020188e-03], [7.47355679e-03, 7.19466945e-03, 6.92162011e-03, ..., 6.08533947e-03, 5.86504815e-03, 5.69197629e-03], [3.75665119e-03, 5.07736765e-03, 7.03470875e-03, ..., 6.20458368e-03, 5.97196259e-03, 5.75759541e-03], ... 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00]], [[0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], ..., [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00]]]], dtype=float32) - THETA(time, k, j, i)float3223.1 22.98 22.81 ... 0.0 0.0 0.0
- standard_name :
- THETA
- long_name :
- Potential Temperature
- units :
- degC
array([[[[23.102392 , 22.97575 , 22.808664 , ..., 22.021645 , 21.61812 , 21.110315 ], [22.90873 , 22.78377 , 22.627527 , ..., 21.821398 , 21.354494 , 20.753613 ], [22.73793 , 22.616974 , 22.47819 , ..., 21.770494 , 21.326824 , 20.72706 ], ..., [21.73688 , 21.63554 , 21.541092 , ..., 21.240301 , 21.127367 , 21.015366 ], [21.430742 , 21.323229 , 21.22359 , ..., 20.940674 , 20.838423 , 20.726461 ], [21.147064 , 21.040262 , 20.942566 , ..., 20.633923 , 20.53154 , 20.418625 ]], [[23.110973 , 22.99283 , 22.835894 , ..., 22.075857 , 21.675074 , 21.161968 ], [22.898602 , 22.78074 , 22.633213 , ..., 21.851357 , 21.386576 , 20.781723 ], [22.708565 , 22.593296 , 22.461658 , ..., 21.77499 , 21.333527 , 20.731936 ], ... [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ]], [[ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], ..., [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ]]]], dtype=float32) - GM_Kvy(time, k, j_g, i)float321.773e+03 1.718e+03 ... 1e+03 1e+03
- standard_name :
- GM_Kvy
- long_name :
- K_22 element (V.point, Y.dir) of GM-Redi tensor
- units :
- m^2/s
- mate :
- GM_Kux
array([[[[1773.2377 , 1718.4346 , 1661.4255 , ..., 1496.4053 , 1457.339 , 1437.1597 ], [1801.3541 , 1740.1859 , 1679.3044 , ..., 1498.5139 , 1453.9059 , 1423.7748 ], [1831.5298 , 1764.724 , 1700.8589 , ..., 1516.3419 , 1467.9069 , 1427.3201 ], ..., [1713.6536 , 1671.4559 , 1636.3822 , ..., 1591.4851 , 1582.5416 , 1576.6406 ], [1702.2253 , 1668.5106 , 1643.9369 , ..., 1616.0554 , 1605.5177 , 1599.4583 ], [1735.4988 , 1705.3276 , 1685.2504 , ..., 1652.2048 , 1631.7039 , 1615.5544 ]], [[1859.0394 , 1798.7805 , 1732.9152 , ..., 1527.8093 , 1478.4789 , 1448.44 ], [1897.9014 , 1827.0468 , 1755.0426 , ..., 1531.5435 , 1472.9321 , 1429.2697 ], [1944.1934 , 1864.8488 , 1788.2233 , ..., 1556.1389 , 1491.3458 , 1435.0729 ], ... [ 992.4003 , 1000. , 1000. , ..., 1000. , 1000. , 1000. ], [1000. , 1000. , 1000. , ..., 1000. , 1000. , 1000. ], [1000. , 1000. , 1000. , ..., 1000. , 1000. , 1000. ]], [[1000. , 1000. , 1000. , ..., 1000. , 1000. , 1000. ], [1000. , 1000. , 1000. , ..., 1000. , 1000. , 1000. ], [1000. , 1000. , 1000. , ..., 1000. , 1000. , 1000. ], ..., [1000. , 1000. , 1000. , ..., 1000. , 1000. , 1000. ], [1000. , 1000. , 1000. , ..., 1000. , 1000. , 1000. ], [1000. , 1000. , 1000. , ..., 1000. , 1000. , 1000. ]]]], dtype=float32) - GM_Kwy(time, k_l, j, i)float320.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0
- standard_name :
- GM_Kwy
- long_name :
- K_32 element (W.point, Y.dir) of GM-Redi tensor
- units :
- m^2/s
- mate :
- GM_Kwx
array([[[[ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], ..., [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00]], [[ 3.65245175e+00, 3.52934194e+00, 3.39922357e+00, ..., 2.94598818e+00, 2.77828979e+00, 2.70381212e+00], [ 3.73470831e+00, 3.59557581e+00, 3.45126987e+00, ..., 2.76040745e+00, 2.57247281e+00, 2.50619626e+00], [ 2.67661858e+00, 3.04864812e+00, 3.51111817e+00, ..., 2.52925396e+00, 2.17401552e+00, 1.99876392e+00], ... 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00]], [[ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], ..., [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00]]]], dtype=float32) - GM_KwzTz(time, k_l, j, i)float320.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0
- standard_name :
- GM_KwzTz
- long_name :
- Redi main-diagonal vertical Temperature flux
- units :
- degC.m^3/s
array([[[[ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], ..., [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00]], [[ 8.87448669e+02, 1.21769238e+03, 1.56376978e+03, ..., 2.17398022e+03, 2.14781274e+03, 2.14938110e+03], [ 3.75216172e+04, 1.31677266e+05, 1.05010974e+03, ..., 1.67998682e+03, 1.73628503e+03, 1.49294324e+03], [-8.45587305e+03, 2.62114883e+04, 1.12439867e+05, ..., 1.24973389e+03, 1.28753296e+03, 1.06987512e+03], ... 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00]], [[ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], ..., [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00]]]], dtype=float32) - GM_Kux(time, k, j, i_g)float321.813e+03 1.757e+03 ... 1e+03 1e+03
- standard_name :
- GM_Kux
- long_name :
- K_11 element (U.point, X.dir) of GM-Redi tensor
- units :
- m^2/s
- mate :
- GM_Kvy
array([[[[1813.0138 , 1756.5457 , 1697.5741 , ..., 1520.3167 , 1472.3077 , 1439.0079 ], [1850.2822 , 1784.9944 , 1721.9163 , ..., 1532.594 , 1480.1122 , 1438.6727 ], [1883.9384 , 1811.2594 , 1743.6666 , ..., 1556.6793 , 1504.1367 , 1456.5541 ], ..., [1719.0902 , 1678.9803 , 1646.4723 , ..., 1604.3131 , 1596.5332 , 1591.3633 ], [1727.1248 , 1691.7556 , 1665.9753 , ..., 1637.8092 , 1625.0399 , 1613.6127 ], [1782.4603 , 1749.0708 , 1724.6027 , ..., 1683.324 , 1658.8688 , 1633.6456 ]], [[1906.1835 , 1842.886 , 1774.9333 , ..., 1557.7323 , 1496.9907 , 1451.6184 ], [1959.1198 , 1882.0621 , 1807.1561 , ..., 1575.4631 , 1507.4847 , 1450.5834 ], [2014.1537 , 1926.98 , 1845.916 , ..., 1609.519 , 1540. , 1475.8354 ], ... [1000. , 1000. , 1000. , ..., 1000. , 1000. , 1000. ], [1000. , 1000. , 1000. , ..., 1000. , 1000. , 1000. ], [1000. , 1000. , 1000. , ..., 1000. , 1000. , 1000. ]], [[1000. , 1000. , 1000. , ..., 1000. , 1000. , 1000. ], [1000. , 1000. , 1000. , ..., 1000. , 1000. , 1000. ], [1000. , 1000. , 1000. , ..., 1000. , 1000. , 1000. ], ..., [1000. , 1000. , 1000. , ..., 1000. , 1000. , 1000. ], [1000. , 1000. , 1000. , ..., 1000. , 1000. , 1000. ], [1000. , 1000. , 1000. , ..., 1000. , 1000. , 1000. ]]]], dtype=float32) - GM_Kwx(time, k_l, j, i)float320.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0
- standard_name :
- GM_Kwx
- long_name :
- K_31 element (W.point, X.dir) of GM-Redi tensor
- units :
- m^2/s
- mate :
- GM_Kwy
array([[[[ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], ..., [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00]], [[ 2.60540068e-01, 2.34605163e-01, 2.42803559e-01, ..., 6.52785242e-01, 9.02349234e-01, 9.32511389e-01], [ 1.24364711e-01, 1.12481669e-01, 2.56792873e-01, ..., 1.27983963e+00, 1.40786433e+00, 1.34856522e+00], [ 3.00220922e-02, 4.38793413e-02, 2.09357426e-01, ..., 1.79641020e+00, 2.04688573e+00, 2.07181549e+00], ... 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00]], [[ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], ..., [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00]]]], dtype=float32) - GM_CHIY(time, k, j_g, i)float321.906e-08 2.389e-08 ... 0.0 0.0
- standard_name :
- GM_CHIY
- long_name :
- Redi Temp variance dissipation rate: Y component
- units :
- degC^2/s
- mate :
- GM_CHIX
array([[[[ 1.90624192e-08, 2.38941720e-08, 2.93394748e-08, ..., 9.22692749e-08, 1.08465798e-07, 1.35906717e-07], [ 5.66162583e-09, 1.03428865e-08, 1.45716239e-08, ..., 2.82827735e-08, 3.92981470e-08, 5.22955865e-08], [-6.93589408e-09, -3.01445646e-09, 8.69445016e-10, ..., 2.58451816e-09, 1.35648348e-09, 1.08896103e-09], ..., [-1.30068214e-08, -1.55594684e-08, -1.80896631e-08, ..., -1.91424245e-08, -1.82408684e-08, -1.28363808e-08], [ 2.23010854e-09, 2.96149050e-10, -2.37851006e-09, ..., -1.15426690e-08, -1.29760398e-08, -1.33164217e-08], [ 1.31179485e-08, 1.13270726e-08, 8.80011441e-09, ..., -1.11021348e-09, -4.31439240e-09, -7.05089720e-09]], [[ 3.39417490e-08, 4.63411531e-08, 5.99678103e-08, ..., 1.79574002e-07, 2.15385626e-07, 2.74676296e-07], [ 1.06769855e-08, 2.16716263e-08, 3.24281260e-08, ..., 6.62833202e-08, 8.91267504e-08, 1.15742104e-07], [-1.20703643e-08, -3.64434150e-09, 5.15297449e-09, ..., 1.02737490e-08, 7.06276237e-09, 5.89086113e-09], ... 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00]], [[ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], ..., [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00]]]], dtype=float32) - GM_Kuz(time, k, j, i_g)float320.29 0.2908 0.2527 ... 0.0 0.0 0.0
- standard_name :
- GM_Kuz
- long_name :
- K_13 element (U.point, Z.dir) of GM-Redi tensor
- units :
- m^2/s
- mate :
- GM_Kvz
array([[[[ 2.89955556e-01, 2.90787369e-01, 2.52726793e-01, ..., 5.24166942e-01, 7.97675312e-01, 9.91192639e-01], [ 1.70158729e-01, 1.28065750e-01, 1.62317991e-01, ..., 1.15654528e+00, 1.39321506e+00, 1.44695711e+00], [ 6.34963661e-02, 5.96320555e-02, 9.39691141e-02, ..., 1.58118975e+00, 1.98407197e+00, 2.11331797e+00], ..., [ 4.15094078e-01, 3.41445595e-01, 2.41583496e-01, ..., -3.59266996e-02, -4.36268933e-02, -8.35403614e-03], [ 5.09507298e-01, 4.35916156e-01, 3.70185673e-01, ..., 6.48831576e-02, 2.09460016e-02, 9.85645410e-03], [ 5.75008690e-01, 5.11762679e-01, 4.77979153e-01, ..., 2.23056570e-01, 1.43688723e-01, 1.13735698e-01]], [[ 2.43646011e-01, 2.22522706e-01, 1.64260954e-01, ..., 4.71758395e-01, 7.69009650e-01, 9.88173664e-01], [ 1.29131779e-01, 6.95698410e-02, 8.79006013e-02, ..., 1.10139918e+00, 1.35940289e+00, 1.42814076e+00], [ 3.01836841e-02, 1.65591687e-02, 3.89116369e-02, ..., 1.53717041e+00, 1.95709538e+00, 2.08987164e+00], ... 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00]], [[ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], ..., [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00]]]], dtype=float32) - Tx(time, k, j, i_g)float32-8.891e-07 -1.213e-06 ... 0.0 0.0
array([[[[-8.89052046e-07, -1.21316930e-06, -1.60059699e-06, ..., -2.99717499e-06, -3.86556212e-06, -4.86451700e-06], [-1.01828232e-06, -1.20446236e-06, -1.50598703e-06, ..., -3.35499271e-06, -4.50038578e-06, -5.79177004e-06], [-1.05442734e-06, -1.17340335e-06, -1.34635332e-06, ..., -3.05265075e-06, -4.30406408e-06, -5.81833456e-06], ..., [-1.03584591e-06, -1.01921421e-06, -9.49905882e-07, ..., -1.04443995e-06, -1.13582792e-06, -1.12644739e-06], [-1.14118041e-06, -1.08995812e-06, -1.01011790e-06, ..., -9.64812557e-07, -1.03660886e-06, -1.13505075e-06], [-1.19914841e-06, -1.09166808e-06, -9.98595056e-07, ..., -1.05078527e-06, -1.04649621e-06, -1.15415207e-06]], [[-8.26983978e-07, -1.13175167e-06, -1.50337462e-06, ..., -2.92776190e-06, -3.83930592e-06, -4.91529318e-06], [-9.66401103e-07, -1.13603517e-06, -1.42198815e-06, ..., -3.29494878e-06, -4.47992352e-06, -5.83004658e-06], [-1.01111118e-06, -1.11822658e-06, -1.27702162e-06, ..., -2.99989802e-06, -4.28265548e-06, -5.83606061e-06], ... 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00]], [[ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], ..., [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00]]]], dtype=float32) - Ty(time, k, j_g, i)float32-2.024e-06 -1.976e-06 ... 0.0 0.0
array([[[[-2.02392766e-06, -1.97576173e-06, -1.93450251e-06, ..., -3.78693994e-06, -4.33204650e-06, -5.48596108e-06], [-1.85169097e-06, -1.83560599e-06, -1.73192882e-06, ..., -1.91464505e-06, -2.52064137e-06, -3.41058740e-06], [-1.64339349e-06, -1.60487241e-06, -1.43689567e-06, ..., -4.89780746e-07, -2.66233855e-07, -2.55479534e-07], ..., [-3.02616422e-06, -3.07592154e-06, -3.06799006e-06, ..., -2.60054480e-06, -2.48621313e-06, -1.92106017e-06], [-3.07733944e-06, -3.13940222e-06, -3.19157198e-06, ..., -3.01190198e-06, -2.90451408e-06, -2.90411162e-06], [-2.87522926e-06, -2.86801856e-06, -2.84833845e-06, ..., -3.10908854e-06, -3.11042254e-06, -3.12008842e-06]], [[-2.17531374e-06, -2.13579506e-06, -2.10330973e-06, ..., -3.95781399e-06, -4.51298138e-06, -5.66325207e-06], [-2.03057743e-06, -2.02787828e-06, -1.93791539e-06, ..., -2.14654642e-06, -2.75845150e-06, -3.63568620e-06], [-1.82849271e-06, -1.80355221e-06, -1.65066092e-06, ..., -7.34781224e-07, -5.10426844e-07, -4.79044729e-07], ... 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00]], [[ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], ..., [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00]]]], dtype=float32) - Tz(time, k_l, j, i)float32nan nan nan nan ... 0.0 0.0 0.0 0.0
array([[[[ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], ..., [ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan]], [[ 8.58116138e-04, 1.70803070e-03, 2.72293086e-03, ..., 5.42125711e-03, 5.69534302e-03, 5.16529102e-03], [-1.01280212e-03, -3.02886969e-04, 5.68580639e-04, ..., 2.99587240e-03, 3.20816040e-03, 2.81105051e-03], [-2.93655391e-03, -2.36778264e-03, -1.65309908e-03, ..., 4.49562067e-04, 6.70242298e-04, 4.87518322e-04], ... -4.06887056e-03, 0.00000000e+00, 0.00000000e+00], [-4.06723469e-03, -4.06506099e-03, -4.06806776e-03, ..., -4.05515358e-03, 0.00000000e+00, 0.00000000e+00], [-4.09392221e-03, -4.08582808e-03, -4.08058101e-03, ..., -4.05411376e-03, 0.00000000e+00, 0.00000000e+00]], [[ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [-7.82281067e-03, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], ..., [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00]]]], dtype=float32) - chix_(time, k, j, i_g)float321.433e-09 2.585e-09 ... 0.0 0.0
array([[[[1.43303058e-09, 2.58524824e-09, 4.34903313e-09, ..., 1.36570932e-08, 2.20000604e-08, 3.40520003e-08], [1.91855554e-09, 2.58954413e-09, 3.90530097e-09, ..., 1.72508408e-08, 2.99774108e-08, 4.82597038e-08], [2.09459472e-09, 2.49387844e-09, 3.16068727e-09, ..., 1.45061918e-08, 2.78640844e-08, 4.93087491e-08], ..., [1.84454385e-09, 1.74412085e-09, 1.48564683e-09, ..., 1.75007275e-09, 2.05969553e-09, 2.01925499e-09], [2.24922192e-09, 2.00982031e-09, 1.69985837e-09, ..., 1.52457635e-09, 1.74619963e-09, 2.07888218e-09], [2.56310129e-09, 2.08443618e-09, 1.71976022e-09, ..., 1.85864157e-09, 1.81671733e-09, 2.17612550e-09]], [[1.30364364e-09, 2.36048225e-09, 4.01158928e-09, ..., 1.33525537e-08, 2.20660468e-08, 3.50712561e-08], [1.82968285e-09, 2.42894416e-09, 3.65416053e-09, ..., 1.71043109e-08, 3.02547889e-08, 4.93045214e-08], [2.05916173e-09, 2.40955478e-09, 3.01029091e-09, ..., 1.44846863e-08, 2.82453527e-08, 5.02663688e-08], ... 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00]], [[0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], ..., [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00]]]], dtype=float32) - chiy_(time, k, j_g, i)float327.264e-09 6.708e-09 ... 0.0 0.0
array([[[[7.26368388e-09, 6.70814027e-09, 6.21755269e-09, ..., 2.14598188e-08, 2.73493370e-08, 4.32524274e-08], [6.17641005e-09, 5.86346793e-09, 5.03720354e-09, ..., 5.49335111e-09, 9.23758403e-09, 1.65615006e-08], [4.94648988e-09, 4.54525040e-09, 3.51171092e-09, ..., 3.63747948e-10, 1.04045918e-10, 9.31608818e-11], ..., [1.56930735e-08, 1.58141358e-08, 1.54025503e-08, ..., 1.07629488e-08, 9.78209425e-09, 5.81854875e-09], [1.61201044e-08, 1.64445844e-08, 1.67453571e-08, ..., 1.46601327e-08, 1.35444722e-08, 1.34896148e-08], [1.43472700e-08, 1.40272247e-08, 1.36724898e-08, ..., 1.59709259e-08, 1.57862914e-08, 1.57273448e-08]], [[8.79695605e-09, 8.20535462e-09, 7.66626407e-09, ..., 2.39320492e-08, 3.01121830e-08, 4.64549821e-08], [7.82551179e-09, 7.51334639e-09, 6.59109034e-09, ..., 7.05683378e-09, 1.12076215e-08, 1.88923917e-08], [6.50018839e-09, 6.06598105e-09, 4.87233853e-09, ..., 8.40164771e-10, 3.88548638e-10, 3.29326066e-10], ... 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00]], [[0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], ..., [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00]]]], dtype=float32) - GM_CHI(time, k, j, i)float321.467e-08 2.126e-08 ... nan nan
array([[[[ 1.46667576e-08, 2.12609699e-08, 2.89983415e-08, ..., 9.06510706e-08, 1.23572306e-07, 1.52458412e-07], [ 2.26014896e-09, 1.71757026e-08, 1.31287567e-08, ..., 5.39823226e-08, 8.17835470e-08, 1.05452770e-07], [-9.69373826e-09, -4.79596274e-09, 7.23206783e-09, ..., 2.55406025e-08, 4.65365844e-08, 7.37664649e-08], ..., [-5.68891778e-09, -6.71004541e-09, -8.08713896e-09, ..., -1.03007043e-08, -1.04933218e-08, -8.27528623e-09], [ 5.61306068e-09, 4.74314943e-09, 2.90452395e-09, ..., -3.59815822e-09, -4.92300689e-09, -6.05926820e-09], [ 1.52130148e-08, 1.32342919e-08, 9.90639659e-09, ..., 2.68633693e-09, 8.58029259e-10, -1.16215693e-09]], [[ 2.46134952e-08, 3.81109437e-08, 5.33822089e-08, ..., 1.66205268e-07, 2.30063804e-07, 2.91313910e-07], [ 1.09292371e-08, 2.24053256e-08, 2.44300953e-08, ..., 9.66992033e-08, 1.42642548e-07, 1.80024244e-07], [ 3.94496702e-09, 6.66413058e-09, 6.04141270e-09, ..., 4.00267233e-08, 6.77672389e-08, 9.97493927e-08], ... 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00]], [[ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], ..., [ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan]]]], dtype=float32)
- iPandasIndex
PandasIndex(Int64Index([10, 11, 12, 13, 14, 15, 16, 17], dtype='int64', name='i'))
- i_gPandasIndex
PandasIndex(Int64Index([10, 11, 12, 13, 14, 15, 16, 17], dtype='int64', name='i_g'))
- jPandasIndex
PandasIndex(Int64Index([10, 11, 12, 13, 14, 15, 16, 17, 18, 19], dtype='int64', name='j'))
- j_gPandasIndex
PandasIndex(Int64Index([10, 11, 12, 13, 14, 15, 16, 17, 18, 19], dtype='int64', name='j_g'))
- kPandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49], dtype='int64', name='k')) - k_uPandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49], dtype='int64', name='k_u')) - k_lPandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49], dtype='int64', name='k_l')) - k_p1PandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50], dtype='int64', name='k_p1')) - timePandasIndex
PandasIndex(TimedeltaIndex(['0 days 00:00:01', '0 days 00:00:02', '0 days 00:00:03', '0 days 00:00:04', '0 days 00:00:05', '0 days 00:00:06', '0 days 00:00:07', '0 days 00:00:08', '0 days 00:00:09', '0 days 00:00:10', '0 days 00:00:11', '0 days 00:00:12', '0 days 00:00:13', '0 days 00:00:14', '0 days 00:00:15', '0 days 00:00:16', '0 days 00:00:17', '0 days 00:00:18', '0 days 00:00:19', '0 days 00:00:20', '0 days 00:00:21', '0 days 00:00:22', '0 days 00:00:23', '0 days 00:00:24'], dtype='timedelta64[ns]', name='time', freq=None))
- Conventions :
- CF-1.6
- title :
- netCDF wrapper of MITgcm MDS binary data
- source :
- MITgcm
- history :
- Created by calling `open_mdsdataset(grid_dir='/glade/work/dcherian/mitgcm/ECCOV4/release4/run/diags/../', iters=None, prefix=None, read_grid=True, delta_t=1, ref_date=None, calendar='gregorian', geometry='llc', grid_vars_to_coords=False, swap_dims=False, endian='>', chunks={'face': 1, 'k_l': -1, 'k': -1}, ignore_unknown_vars=False, default_dtype=None, nx=None, ny=None, nz=None, llc_method='smallchunks', extra_metadata=None, extra_variables=None, data_dir='/glade/work/dcherian/mitgcm/ECCOV4/release4/run/diags/GM_CHIZ', levels=None)`
natre.GM_CHI.mean(["time", "j", "i"]).plot(y="Z")
[<matplotlib.lines.Line2D at 0x2b4fc22cb430>]
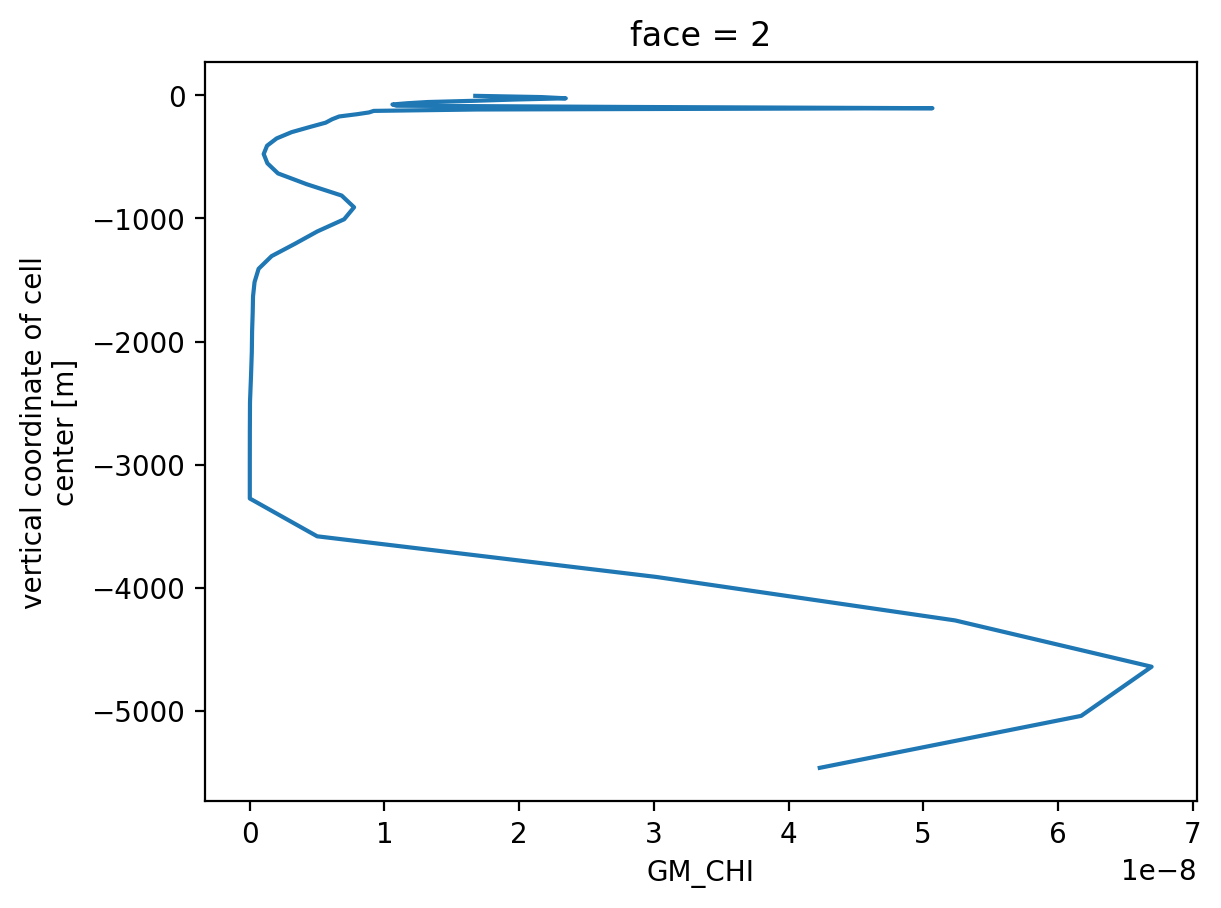
Development#
Checking CHIX, CHIXO, CHIY, CHIYO#
ds["chix_"] = ds.GM_Kux * ds.Tx**2
ds["chixo_"] = 2 * ds.GM_Kuz * ds.Tx * grid.interp(ds.Tz, axis=["X", "Z"])
ds["chiy_"] = ds.GM_Kvy * ds.Ty**2
ds["chiyo_"] = 2 * ds.GM_Kvz * ds.Ty * grid.interp(ds.Tz, axis=["Y", "Z"])
ds[["GM_Kvz"]].isel(time=5, j_g=20, i=20).to_array().plot(hue="variable")
ds[["GM_Kuz"]].isel(time=5, j=20, i_g=20).to_array().plot(hue="variable")
[<matplotlib.lines.Line2D at 0x2ab5eb6bc550>]
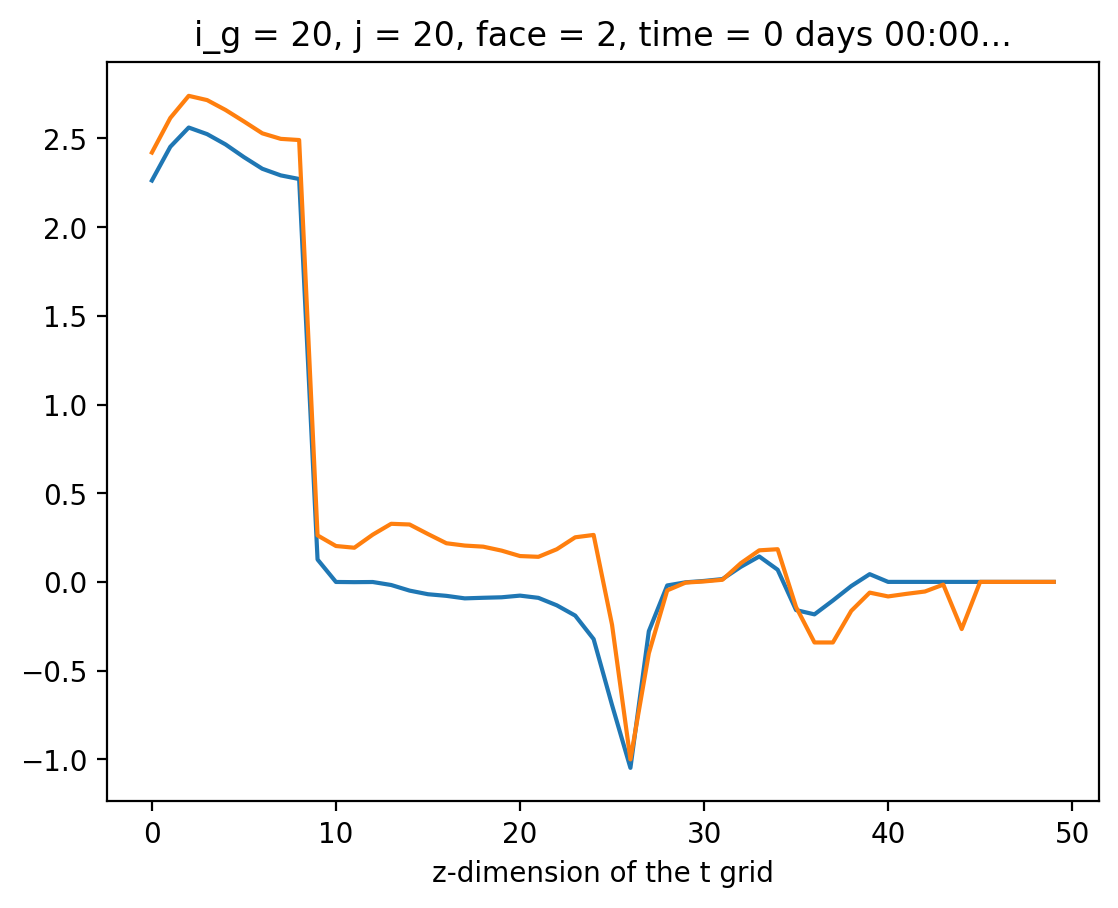
ds[["chix_", "GM_CHIX"]].isel(time=5, j=20, i_g=20).to_array().plot(
hue="variable", yscale="log"
)
[<matplotlib.lines.Line2D at 0x2ab5f2fcf730>,
<matplotlib.lines.Line2D at 0x2ab5f2fcf700>]
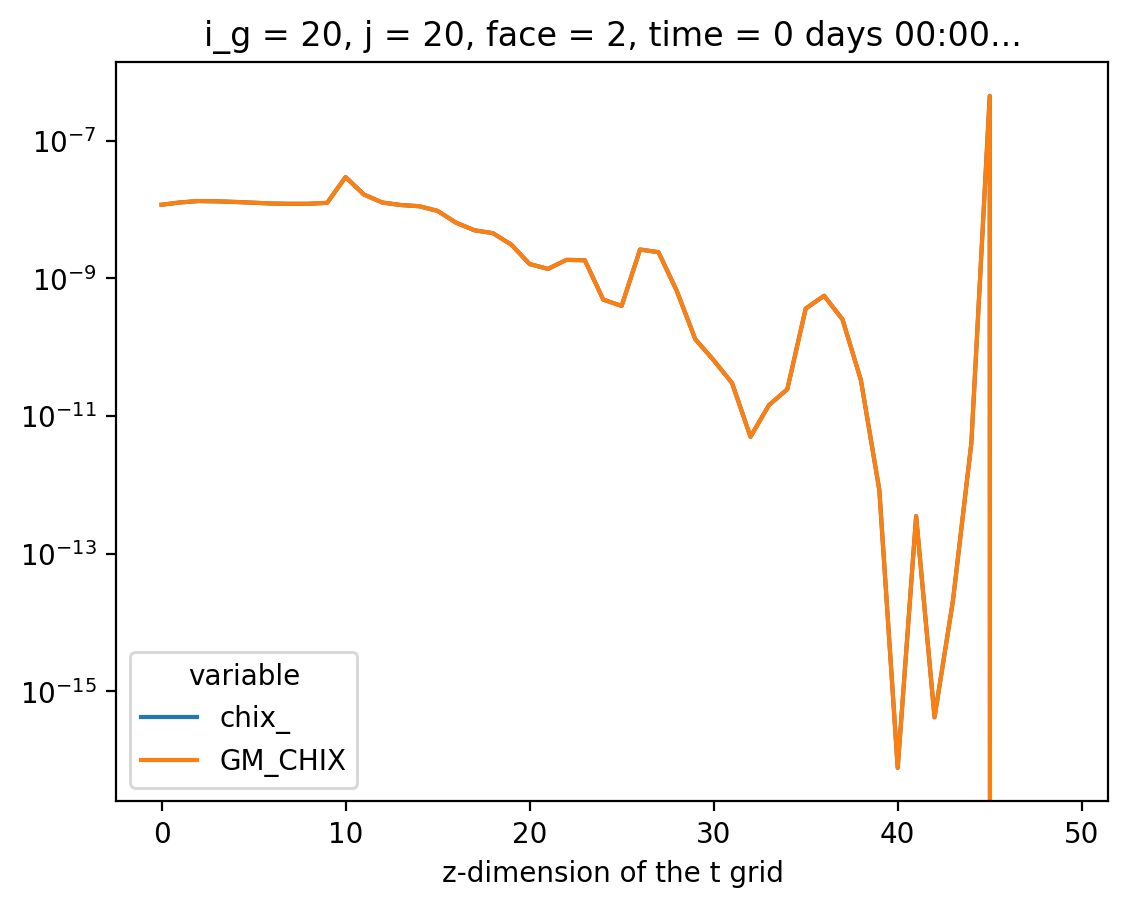
ds[["chixo_", "GM_CHIXO"]].isel(time=5, j=20, i_g=20).to_array().plot(hue="variable")
[<matplotlib.lines.Line2D at 0x2ab5eee5f400>,
<matplotlib.lines.Line2D at 0x2ab5eee5e1a0>]
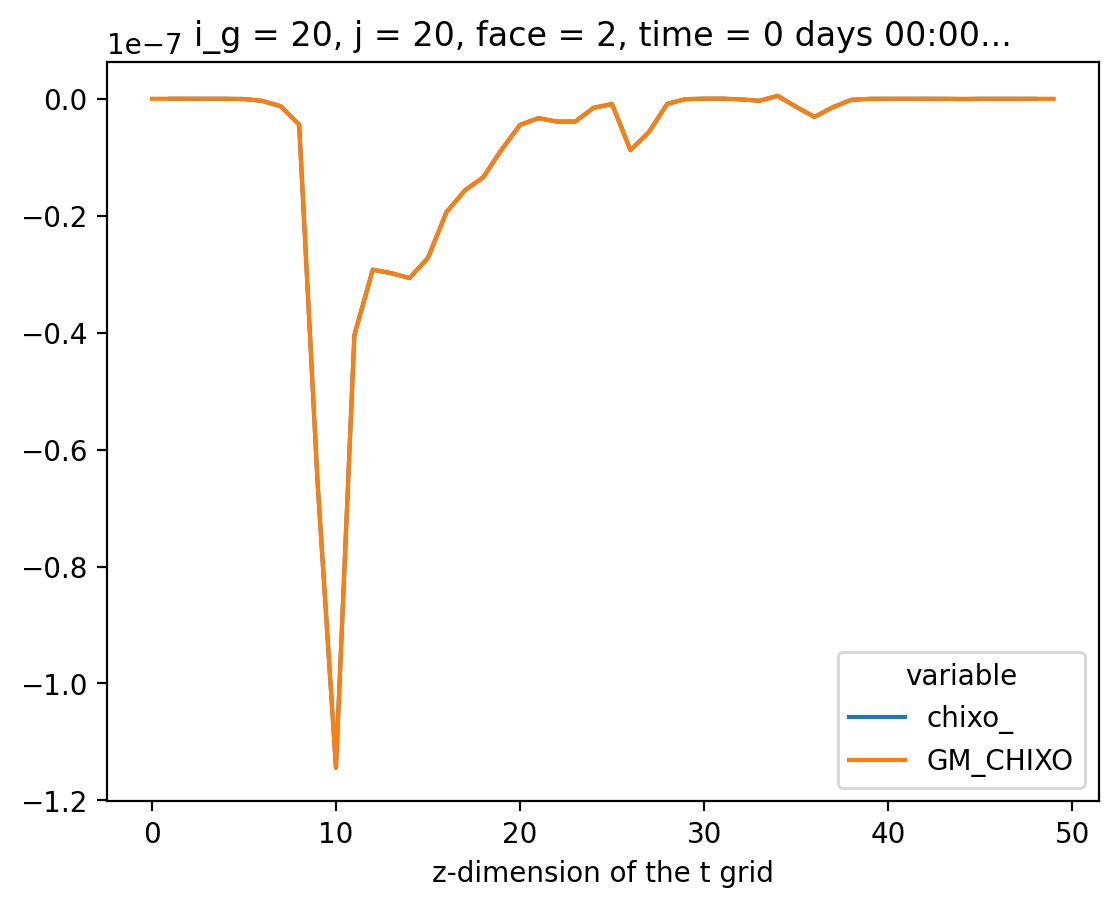
ds[["chiy_", "GM_CHIY"]].isel(time=5, j_g=20, i=20).to_array().plot(
hue="variable", yscale="log"
)
[<matplotlib.lines.Line2D at 0x2ab60a3ef8e0>,
<matplotlib.lines.Line2D at 0x2ab60a3ee0e0>]
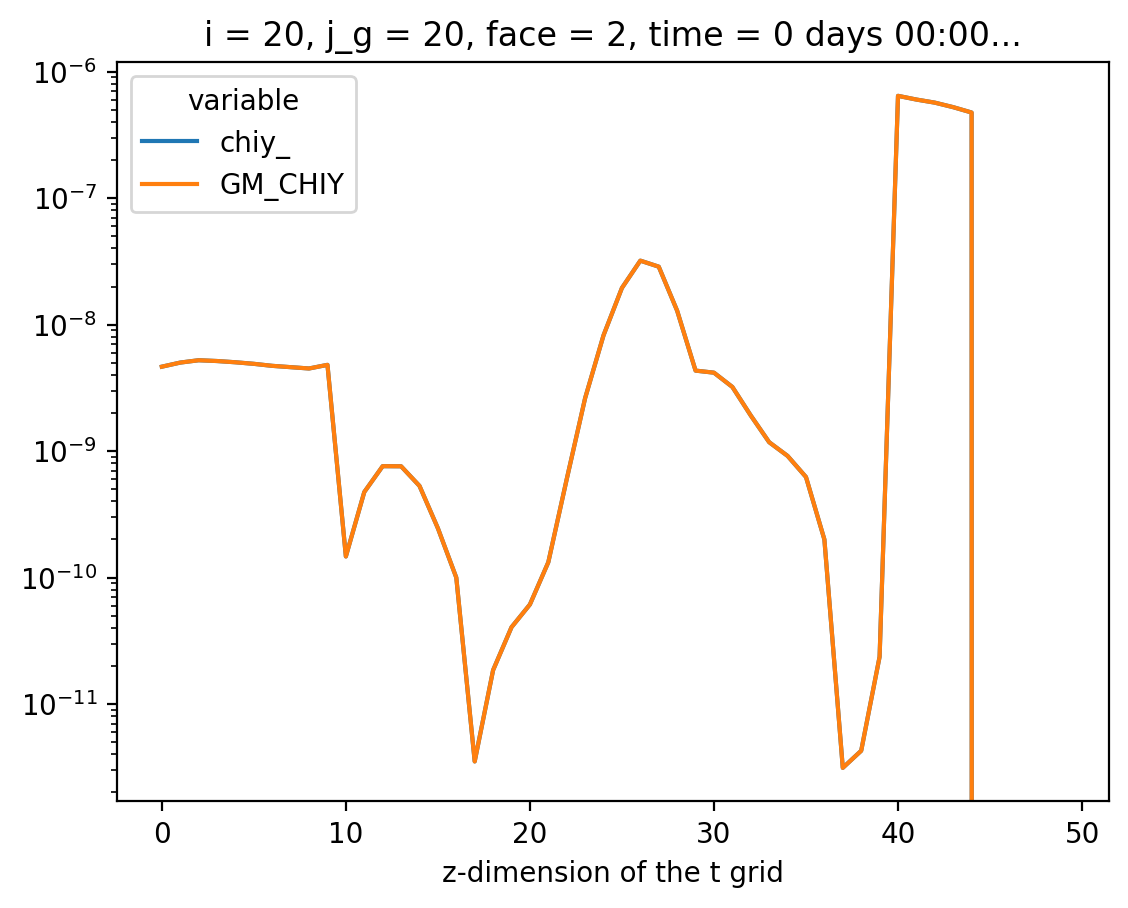
ds[["chiyo_", "GM_CHIYO"]].isel(time=5, j_g=20, i=20).to_array().plot(
hue="variable",
)
[<matplotlib.lines.Line2D at 0x2ab5f33cbac0>,
<matplotlib.lines.Line2D at 0x2ab5f33cba90>]
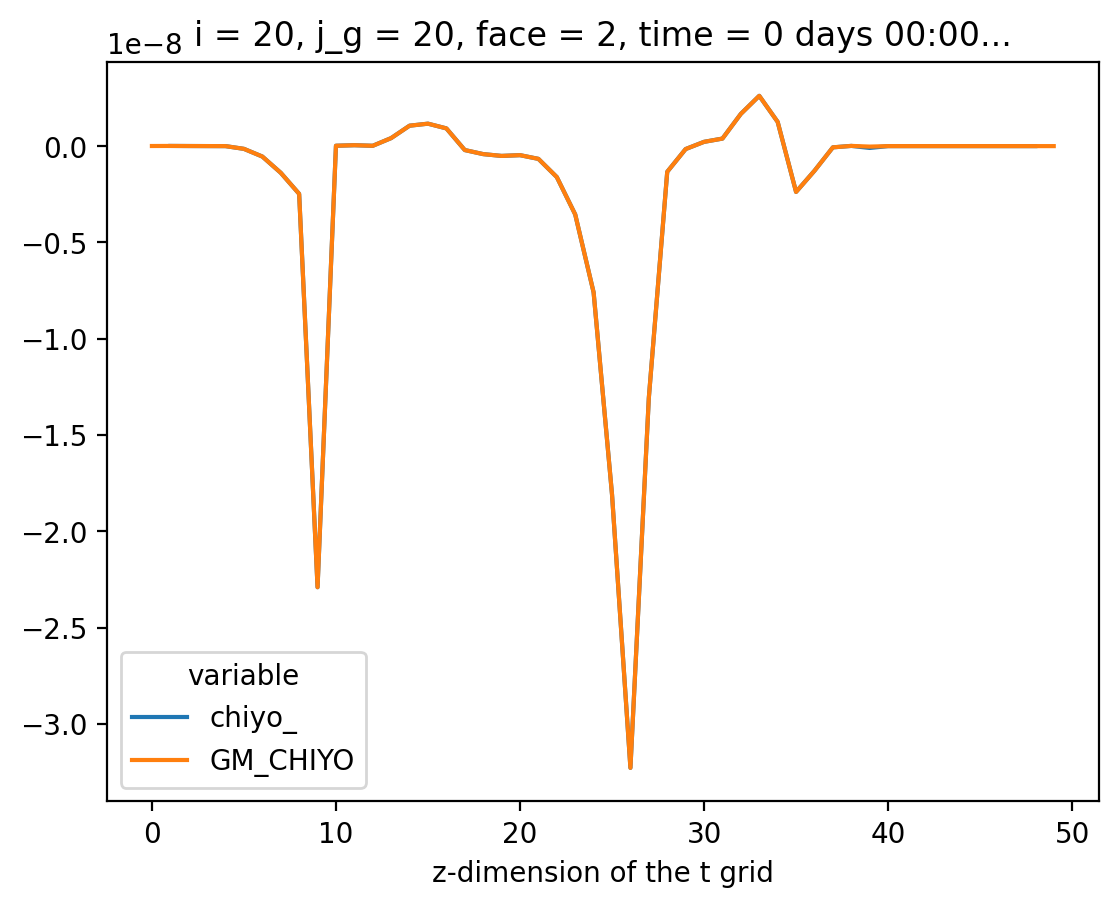
Checking CHIZ#
subset = ds.isel(time=4, i_g=4, i=4, j=39, j_g=39)
subset["GM_Tz"] = subset["GM_KwzTz"] / subset.GM_Kwz / subset.rA
# subset["THETA"] = ds.THETA.isel(time=5,i=4, j=39)
subset = subset.load()
subset.drC.isel(k_p1=slice(1, None)) still seems right
subset["Tz_2"] = (
-grid.diff(subset.THETA, "Z")
/ subset.drC.isel(k_p1=slice(1, None)).rename({"k_p1": "k_l"}).variable
)
# (subset["GM_KwzTz"] * subset["GM_KwzTz"] / subset.GM_Kwz / subset.rA**2).plot()
np.sqrt(subset.GM_CHIZ / subset.GM_Kwz).plot(marker=".")
np.abs(subset.Tz).plot()
np.abs(subset.Tz_2).plot()
[<matplotlib.lines.Line2D at 0x2ab52a7568c0>]
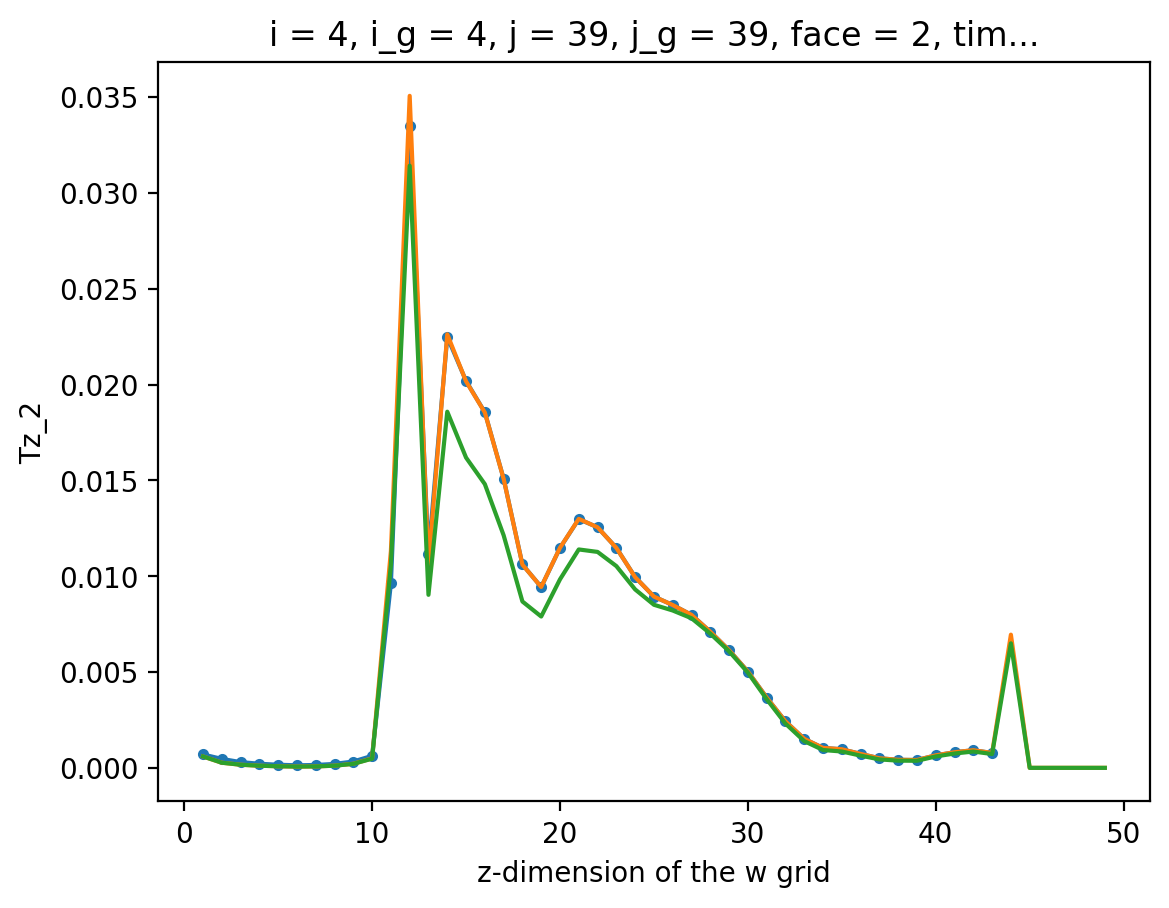
(subset["GM_KwzTz"]).plot(marker=".")
(-subset.GM_Kwz * subset.Tz * subset.rA).plot()
(-subset.GM_Kwz * subset.Tz_2 * subset.rA).plot()
[<matplotlib.lines.Line2D at 0x2ab5f4135660>]
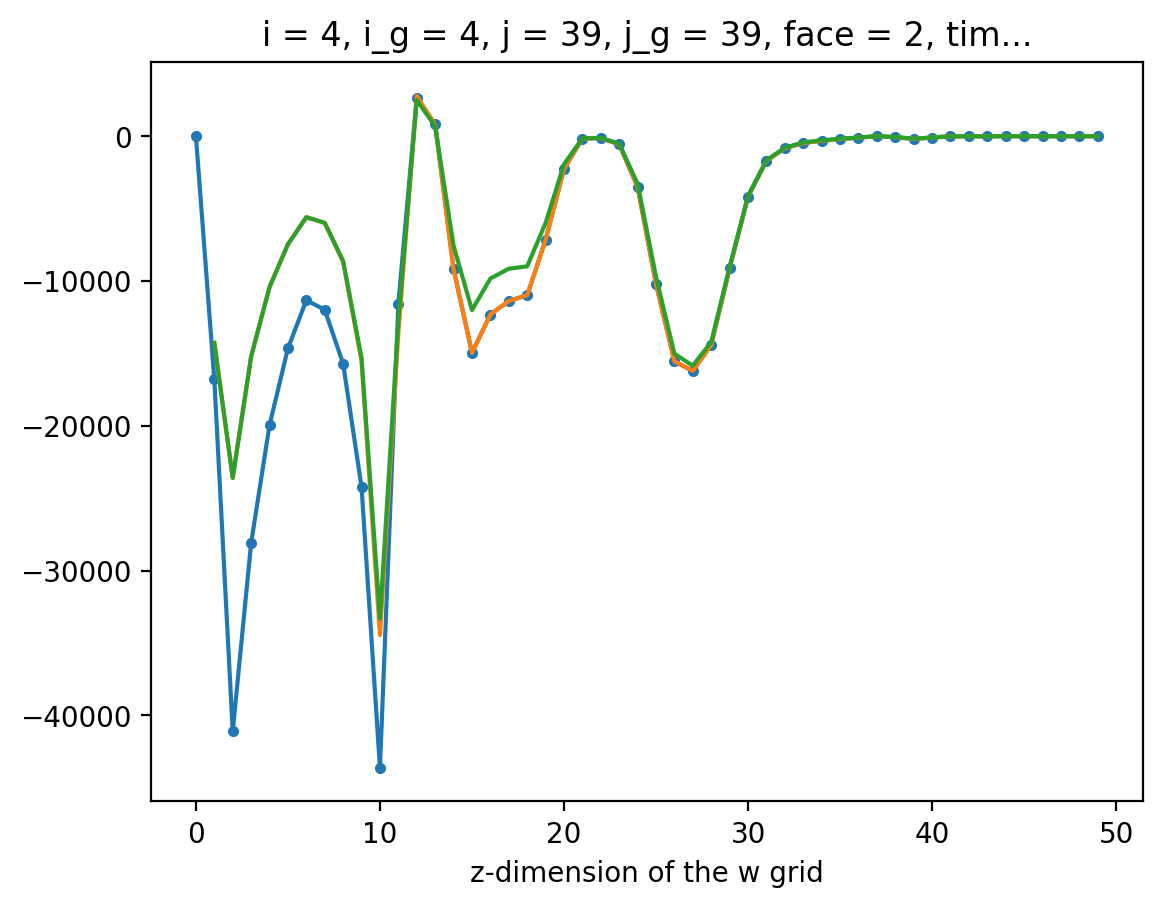
Is Kvz == Kwy?#
Seems close, and errors from averaging probably?
grid.interp(ds.GM_Kvz, axis="Z").isel(j_g=20, i=20, time=1).plot()
grid.interp(ds.GM_Kwy, axis="Y").isel(j_g=20, i=20, time=1).plot()
[<matplotlib.lines.Line2D at 0x2b4fc513cc70>]
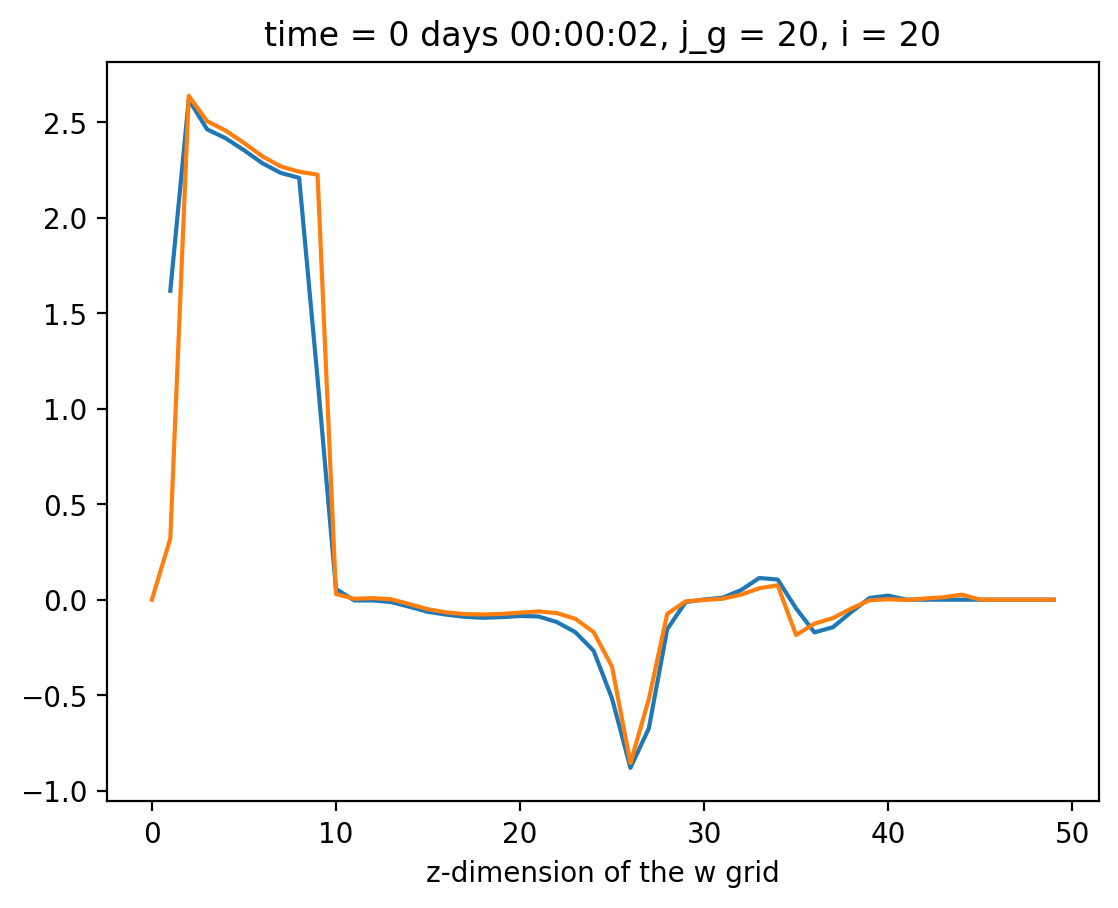
File reading experiments#
ecco.read_llc_to_tiles(
dirname + "GM_CHIZ_INST/", "GM_CHIZ.0000000001.data", nk=50, nl=1, use_xmitgcm=True
)
read_llc_to_tiles: full_filename: /glade/work/dcherian/mitgcm/ECCOV4/release4/run/diags/GM_CHIZ_INST//GM_CHIZ.0000000001.data
|
||||||||||||||||
import xmitgcm
xmitgcm.open_mdsdataset(dirname + "GM_CHIZ_INST/", dirname + "../")
<xarray.Dataset>
Dimensions: (XC: 90, YC: 1170, XG: 90, YG: 1170, Z: 50, Zp1: 51, Zu: 50,
Zl: 50, time: 8)
Coordinates: (12/33)
* XC (XC) >f4 0.0 0.0 0.0 0.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0 0.0 0.0
* YC (YC) >f4 0.0 0.0 0.0 0.0 0.0 ... 9.482 -57.27 67.47 9.482 -57.27
* XG (XG) >f4 0.0 0.0 0.0 0.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0 0.0 0.0
* YG (YG) >f4 0.0 0.0 0.0 0.0 0.0 ... 9.97 -57.01 67.56 9.97 -57.01
* Z (Z) >f4 -5.0 -15.0 -25.0 ... -5.039e+03 -5.461e+03 -5.906e+03
* Zp1 (Zp1) >f4 0.0 -10.0 -20.0 ... -5.244e+03 -5.678e+03 -6.134e+03
... ...
maskCtrlC (Z, YC, XC) bool dask.array<chunksize=(50, 1170, 90), meta=np.ndarray>
maskCtrlS (Z, YG, XC) bool dask.array<chunksize=(50, 1170, 90), meta=np.ndarray>
rhoRef (Z) >f4 dask.array<chunksize=(50,), meta=np.ndarray>
maskCtrlW (Z, YC, XG) bool dask.array<chunksize=(50, 1170, 90), meta=np.ndarray>
iter (time) int64 dask.array<chunksize=(1,), meta=np.ndarray>
* time (time) timedelta64[ns] 00:00:01 00:00:02 ... 00:00:07 00:00:08
Data variables:
GM_CHIZ (time, Z, YC, XC) float32 dask.array<chunksize=(1, 50, 1170, 90), meta=np.ndarray>
Attributes:
Conventions: CF-1.6
title: netCDF wrapper of MITgcm MDS binary data
source: MITgcm
history: Created by calling `open_mdsdataset(grid_dir='/glade/work/d...xarray.Dataset
- XC: 90
- YC: 1170
- XG: 90
- YG: 1170
- Z: 50
- Zp1: 51
- Zu: 50
- Zl: 50
- time: 8
- XC(XC)>f40.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0
- standard_name :
- longitude
- long_name :
- longitude
- units :
- degrees_east
- coordinate :
- YC XC
- axis :
- X
array([0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0.], dtype=float32) - YC(YC)>f40.0 0.0 0.0 ... 67.47 9.482 -57.27
- standard_name :
- latitude
- long_name :
- latitude
- units :
- degrees_north
- coordinate :
- YC XC
- axis :
- Y
array([ 0. , 0. , 0. , ..., 67.47211 , 9.482398, -57.271408], dtype=float32) - XG(XG)>f40.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0
- standard_name :
- longitude_at_f_location
- long_name :
- longitude
- units :
- degrees_east
- coordinate :
- YG XG
- axis :
- X
- c_grid_axis_shift :
- -0.5
array([0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0.], dtype=float32) - YG(YG)>f40.0 0.0 0.0 ... 67.56 9.97 -57.01
- standard_name :
- latitude_at_f_location
- long_name :
- latitude
- units :
- degrees_north
- coordinate :
- YG XG
- axis :
- Y
- c_grid_axis_shift :
- -0.5
array([ 0. , 0. , 0. , ..., 67.560646, 9.96973 , -57.005695], dtype=float32) - Z(Z)>f4-5.0 -15.0 ... -5.906e+03
- standard_name :
- depth
- long_name :
- vertical coordinate of cell center
- units :
- m
- positive :
- down
- axis :
- Z
array([-5.000000e+00, -1.500000e+01, -2.500000e+01, -3.500000e+01, -4.500000e+01, -5.500000e+01, -6.500000e+01, -7.500500e+01, -8.502500e+01, -9.509500e+01, -1.053100e+02, -1.158700e+02, -1.271500e+02, -1.397400e+02, -1.544700e+02, -1.724000e+02, -1.947350e+02, -2.227100e+02, -2.574700e+02, -2.999300e+02, -3.506800e+02, -4.099300e+02, -4.774700e+02, -5.527100e+02, -6.347350e+02, -7.224000e+02, -8.144700e+02, -9.097400e+02, -1.007155e+03, -1.105905e+03, -1.205535e+03, -1.306205e+03, -1.409150e+03, -1.517095e+03, -1.634175e+03, -1.765135e+03, -1.914150e+03, -2.084035e+03, -2.276225e+03, -2.491250e+03, -2.729250e+03, -2.990250e+03, -3.274250e+03, -3.581250e+03, -3.911250e+03, -4.264250e+03, -4.640250e+03, -5.039250e+03, -5.461250e+03, -5.906250e+03], dtype=float32) - Zp1(Zp1)>f40.0 -10.0 ... -5.678e+03 -6.134e+03
- standard_name :
- depth_at_w_location
- long_name :
- vertical coordinate of cell interface
- units :
- m
- positive :
- down
- axis :
- Z
- c_grid_axis_shift :
- (-0.5, 0.5)
array([ 0. , -10. , -20. , -30. , -40. , -50. , -60. , -70. , -80.01, -90.04, -100.15, -110.47, -121.27, -133.03, -146.45, -162.49, -182.31, -207.16, -238.26, -276.68, -323.18, -378.18, -441.68, -513.26, -592.16, -677.31, -767.49, -861.45, -958.03, -1056.28, -1155.53, -1255.54, -1356.87, -1461.43, -1572.76, -1695.59, -1834.68, -1993.62, -2174.45, -2378. , -2604.5 , -2854. , -3126.5 , -3422. , -3740.5 , -4082. , -4446.5 , -4834. , -5244.5 , -5678. , -6134.5 ], dtype=float32) - Zu(Zu)>f4-10.0 -20.0 ... -6.134e+03
- standard_name :
- depth_at_upper_w_location
- long_name :
- vertical coordinate of upper cell interface
- units :
- m
- positive :
- down
- axis :
- Z
- c_grid_axis_shift :
- 0.5
array([ -10. , -20. , -30. , -40. , -50. , -60. , -70. , -80.01, -90.04, -100.15, -110.47, -121.27, -133.03, -146.45, -162.49, -182.31, -207.16, -238.26, -276.68, -323.18, -378.18, -441.68, -513.26, -592.16, -677.31, -767.49, -861.45, -958.03, -1056.28, -1155.53, -1255.54, -1356.87, -1461.43, -1572.76, -1695.59, -1834.68, -1993.62, -2174.45, -2378. , -2604.5 , -2854. , -3126.5 , -3422. , -3740.5 , -4082. , -4446.5 , -4834. , -5244.5 , -5678. , -6134.5 ], dtype=float32) - Zl(Zl)>f40.0 -10.0 ... -5.244e+03 -5.678e+03
- standard_name :
- depth_at_lower_w_location
- long_name :
- vertical coordinate of lower cell interface
- units :
- m
- positive :
- down
- axis :
- Z
- c_grid_axis_shift :
- -0.5
array([ 0. , -10. , -20. , -30. , -40. , -50. , -60. , -70. , -80.01, -90.04, -100.15, -110.47, -121.27, -133.03, -146.45, -162.49, -182.31, -207.16, -238.26, -276.68, -323.18, -378.18, -441.68, -513.26, -592.16, -677.31, -767.49, -861.45, -958.03, -1056.28, -1155.53, -1255.54, -1356.87, -1461.43, -1572.76, -1695.59, -1834.68, -1993.62, -2174.45, -2378. , -2604.5 , -2854. , -3126.5 , -3422. , -3740.5 , -4082. , -4446.5 , -4834. , -5244.5 , -5678. ], dtype=float32) - rA(YC, XC)>f4dask.array<chunksize=(1170, 90), meta=np.ndarray>
- standard_name :
- cell_area
- long_name :
- cell area
- units :
- m2
- coordinate :
- YC XC
Array Chunk Bytes 411.33 kiB 411.33 kiB Shape (1170, 90) (1170, 90) Dask graph 1 chunks in 3 graph layers Data type >f4 numpy.ndarray - dxG(YG, XC)>f4dask.array<chunksize=(1170, 90), meta=np.ndarray>
- standard_name :
- cell_x_size_at_v_location
- long_name :
- cell x size
- units :
- m
- coordinate :
- YG XC
Array Chunk Bytes 411.33 kiB 411.33 kiB Shape (1170, 90) (1170, 90) Dask graph 1 chunks in 3 graph layers Data type >f4 numpy.ndarray - dyG(YC, XG)>f4dask.array<chunksize=(1170, 90), meta=np.ndarray>
- standard_name :
- cell_y_size_at_u_location
- long_name :
- cell y size
- units :
- m
- coordinate :
- YC XG
Array Chunk Bytes 411.33 kiB 411.33 kiB Shape (1170, 90) (1170, 90) Dask graph 1 chunks in 3 graph layers Data type >f4 numpy.ndarray - Depth(YC, XC)>f4dask.array<chunksize=(1170, 90), meta=np.ndarray>
- standard_name :
- ocean_depth
- long_name :
- ocean depth
- units :
- m
- coordinate :
- XC YC
Array Chunk Bytes 411.33 kiB 411.33 kiB Shape (1170, 90) (1170, 90) Dask graph 1 chunks in 3 graph layers Data type >f4 numpy.ndarray - rAz(YG, XG)>f4dask.array<chunksize=(1170, 90), meta=np.ndarray>
- standard_name :
- cell_area_at_f_location
- long_name :
- cell area
- units :
- m
- coordinate :
- YG XG
Array Chunk Bytes 411.33 kiB 411.33 kiB Shape (1170, 90) (1170, 90) Dask graph 1 chunks in 3 graph layers Data type >f4 numpy.ndarray - dxC(YC, XG)>f4dask.array<chunksize=(1170, 90), meta=np.ndarray>
- standard_name :
- cell_x_size_at_u_location
- long_name :
- cell x size
- units :
- m
- coordinate :
- YC XG
Array Chunk Bytes 411.33 kiB 411.33 kiB Shape (1170, 90) (1170, 90) Dask graph 1 chunks in 3 graph layers Data type >f4 numpy.ndarray - dyC(YG, XC)>f4dask.array<chunksize=(1170, 90), meta=np.ndarray>
- standard_name :
- cell_y_size_at_v_location
- long_name :
- cell y size
- units :
- m
- coordinate :
- YG XC
Array Chunk Bytes 411.33 kiB 411.33 kiB Shape (1170, 90) (1170, 90) Dask graph 1 chunks in 3 graph layers Data type >f4 numpy.ndarray - rAw(YC, XG)>f4dask.array<chunksize=(1170, 90), meta=np.ndarray>
- standard_name :
- cell_area_at_u_location
- long_name :
- cell area
- units :
- m2
- coordinate :
- YG XC
Array Chunk Bytes 411.33 kiB 411.33 kiB Shape (1170, 90) (1170, 90) Dask graph 1 chunks in 3 graph layers Data type >f4 numpy.ndarray - rAs(YG, XC)>f4dask.array<chunksize=(1170, 90), meta=np.ndarray>
- standard_name :
- cell_area_at_v_location
- long_name :
- cell area
- units :
- m2
- coordinate :
- YG XC
Array Chunk Bytes 411.33 kiB 411.33 kiB Shape (1170, 90) (1170, 90) Dask graph 1 chunks in 3 graph layers Data type >f4 numpy.ndarray - drC(Zp1)>f4dask.array<chunksize=(51,), meta=np.ndarray>
- standard_name :
- cell_z_size_at_w_location
- long_name :
- cell z size
- units :
- m
Array Chunk Bytes 204 B 204 B Shape (51,) (51,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - drF(Z)>f4dask.array<chunksize=(50,), meta=np.ndarray>
- standard_name :
- cell_z_size
- long_name :
- cell z size
- units :
- m
Array Chunk Bytes 200 B 200 B Shape (50,) (50,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - PHrefC(Z)>f4dask.array<chunksize=(50,), meta=np.ndarray>
- standard_name :
- cell_reference_pressure
- long_name :
- Reference Hydrostatic Pressure
- units :
- m2 s-2
Array Chunk Bytes 200 B 200 B Shape (50,) (50,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - PHrefF(Zp1)>f4dask.array<chunksize=(51,), meta=np.ndarray>
- standard_name :
- cell_reference_pressure
- long_name :
- Reference Hydrostatic Pressure
- units :
- m2 s-2
Array Chunk Bytes 204 B 204 B Shape (51,) (51,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - hFacC(Z, YC, XC)>f4dask.array<chunksize=(50, 1170, 90), meta=np.ndarray>
- standard_name :
- cell_vertical_fraction
- long_name :
- vertical fraction of open cell
Array Chunk Bytes 20.08 MiB 20.08 MiB Shape (50, 1170, 90) (50, 1170, 90) Dask graph 1 chunks in 2 graph layers Data type >f4 numpy.ndarray - hFacW(Z, YC, XG)>f4dask.array<chunksize=(50, 1170, 90), meta=np.ndarray>
- standard_name :
- cell_vertical_fraction_at_u_location
- long_name :
- vertical fraction of open cell
Array Chunk Bytes 20.08 MiB 20.08 MiB Shape (50, 1170, 90) (50, 1170, 90) Dask graph 1 chunks in 2 graph layers Data type >f4 numpy.ndarray - hFacS(Z, YG, XC)>f4dask.array<chunksize=(50, 1170, 90), meta=np.ndarray>
- standard_name :
- cell_vertical_fraction_at_v_location
- long_name :
- vertical fraction of open cell
Array Chunk Bytes 20.08 MiB 20.08 MiB Shape (50, 1170, 90) (50, 1170, 90) Dask graph 1 chunks in 2 graph layers Data type >f4 numpy.ndarray - maskC(Z, YC, XC)booldask.array<chunksize=(50, 1170, 90), meta=np.ndarray>
- standard_name :
- sea_binary_mask_at_t_location
- long_name :
- mask denoting wet point at center
Array Chunk Bytes 5.02 MiB 5.02 MiB Shape (50, 1170, 90) (50, 1170, 90) Dask graph 1 chunks in 3 graph layers Data type bool numpy.ndarray - maskW(Z, YC, XG)booldask.array<chunksize=(50, 1170, 90), meta=np.ndarray>
- standard_name :
- cell_vertical_fraction_at_u_location
- long_name :
- mask denoting wet point at interface
Array Chunk Bytes 5.02 MiB 5.02 MiB Shape (50, 1170, 90) (50, 1170, 90) Dask graph 1 chunks in 3 graph layers Data type bool numpy.ndarray - maskS(Z, YG, XC)booldask.array<chunksize=(50, 1170, 90), meta=np.ndarray>
- standard_name :
- cell_vertical_fraction_at_v_location
- long_name :
- mask denoting wet point at interface
Array Chunk Bytes 5.02 MiB 5.02 MiB Shape (50, 1170, 90) (50, 1170, 90) Dask graph 1 chunks in 3 graph layers Data type bool numpy.ndarray - maskCtrlC(Z, YC, XC)booldask.array<chunksize=(50, 1170, 90), meta=np.ndarray>
- standard_name :
- ctrl_vector_3d_mask
- long_name :
- CTRL 3D mask where ctrl vector is active at tracer location
- units :
Array Chunk Bytes 5.02 MiB 5.02 MiB Shape (50, 1170, 90) (50, 1170, 90) Dask graph 1 chunks in 3 graph layers Data type bool numpy.ndarray - maskCtrlS(Z, YG, XC)booldask.array<chunksize=(50, 1170, 90), meta=np.ndarray>
- standard_name :
- ctrl_vector_3d_mask_at_v_location
- long_name :
- CTRL 3D mask where ctrl vector is active at v location
- units :
Array Chunk Bytes 5.02 MiB 5.02 MiB Shape (50, 1170, 90) (50, 1170, 90) Dask graph 1 chunks in 3 graph layers Data type bool numpy.ndarray - rhoRef(Z)>f4dask.array<chunksize=(50,), meta=np.ndarray>
- standard_name :
- reference_density_profile
- long_name :
- 1D, vertical reference density profile
- coordinate :
- Z
- units :
- kg m-3
Array Chunk Bytes 200 B 200 B Shape (50,) (50,) Dask graph 1 chunks in 1 graph layer Data type >f4 numpy.ndarray - maskCtrlW(Z, YC, XG)booldask.array<chunksize=(50, 1170, 90), meta=np.ndarray>
- standard_name :
- ctrl_vector_3d_mask_at_u_location
- long_name :
- CTRL 3D mask where ctrl vector is active at u location
- units :
Array Chunk Bytes 5.02 MiB 5.02 MiB Shape (50, 1170, 90) (50, 1170, 90) Dask graph 1 chunks in 3 graph layers Data type bool numpy.ndarray - iter(time)int64dask.array<chunksize=(1,), meta=np.ndarray>
- standard_name :
- timestep
- long_name :
- model timestep number
Array Chunk Bytes 64 B 8 B Shape (8,) (1,) Dask graph 8 chunks in 9 graph layers Data type int64 numpy.ndarray - time(time)timedelta64[ns]00:00:01 00:00:02 ... 00:00:08
- standard_name :
- time
- long_name :
- Time
- axis :
- T
- calendar :
- gregorian
array([1000000000, 2000000000, 3000000000, 4000000000, 5000000000, 6000000000, 7000000000, 8000000000], dtype='timedelta64[ns]')
- GM_CHIZ(time, Z, YC, XC)float32dask.array<chunksize=(1, 50, 1170, 90), meta=np.ndarray>
- standard_name :
- GM_CHIZ
- long_name :
- Redi Temp variance dissipation rate: Z component
- units :
- degC^2/s
Array Chunk Bytes 160.68 MiB 20.08 MiB Shape (8, 50, 1170, 90) (1, 50, 1170, 90) Dask graph 8 chunks in 33 graph layers Data type float32 numpy.ndarray
- XCPandasIndex
PandasIndex(Float64Index([0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0], dtype='float64', name='XC')) - YCPandasIndex
PandasIndex(Float64Index([ 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, ... -57.27140808105469, 67.6231689453125, 9.48239803314209, -57.27140808105469, 67.53386688232422, 9.48239803314209, -57.27140808105469, 67.47210693359375, 9.48239803314209, -57.27140808105469], dtype='float64', name='YC', length=1170)) - XGPandasIndex
PandasIndex(Float64Index([0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0], dtype='float64', name='XG')) - YGPandasIndex
PandasIndex(Float64Index([ 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, ... -57.00569534301758, 67.76258850097656, 9.969730377197266, -57.00569534301758, 67.6531982421875, 9.969730377197266, -57.00569534301758, 67.5606460571289, 9.969730377197266, -57.00569534301758], dtype='float64', name='YG', length=1170)) - ZPandasIndex
PandasIndex(Float64Index([ -5.0, -15.0, -25.0, -35.0, -45.0, -55.0, -65.0, -75.00499725341797, -85.0250015258789, -95.09500122070312, -105.30999755859375, -115.87000274658203, -127.1500015258789, -139.74000549316406, -154.47000122070312, -172.39999389648438, -194.73500061035156, -222.7100067138672, -257.4700012207031, -299.92999267578125, -350.67999267578125, -409.92999267578125, -477.4700012207031, -552.7100219726562, -634.7349853515625, -722.4000244140625, -814.469970703125, -909.739990234375, -1007.155029296875, -1105.905029296875, -1205.5350341796875, -1306.2049560546875, -1409.1500244140625, -1517.094970703125, -1634.175048828125, -1765.135009765625, -1914.1500244140625, -2084.034912109375, -2276.22509765625, -2491.25, -2729.25, -2990.25, -3274.25, -3581.25, -3911.25, -4264.25, -4640.25, -5039.25, -5461.25, -5906.25], dtype='float64', name='Z')) - Zp1PandasIndex
PandasIndex(Float64Index([ 0.0, -10.0, -20.0, -30.0, -40.0, -50.0, -60.0, -70.0, -80.01000213623047, -90.04000091552734, -100.1500015258789, -110.47000122070312, -121.2699966430664, -133.02999877929688, -146.4499969482422, -162.49000549316406, -182.30999755859375, -207.16000366210938, -238.25999450683594, -276.67999267578125, -323.17999267578125, -378.17999267578125, -441.67999267578125, -513.260009765625, -592.1599731445312, -677.3099975585938, -767.489990234375, -861.4500122070312, -958.030029296875, -1056.280029296875, -1155.530029296875, -1255.5400390625, -1356.8699951171875, -1461.4300537109375, -1572.760009765625, -1695.5899658203125, -1834.6800537109375, -1993.6199951171875, -2174.449951171875, -2378.0, -2604.5, -2854.0, -3126.5, -3422.0, -3740.5, -4082.0, -4446.5, -4834.0, -5244.5, -5678.0, -6134.5], dtype='float64', name='Zp1')) - ZuPandasIndex
PandasIndex(Float64Index([ -10.0, -20.0, -30.0, -40.0, -50.0, -60.0, -70.0, -80.01000213623047, -90.04000091552734, -100.1500015258789, -110.47000122070312, -121.2699966430664, -133.02999877929688, -146.4499969482422, -162.49000549316406, -182.30999755859375, -207.16000366210938, -238.25999450683594, -276.67999267578125, -323.17999267578125, -378.17999267578125, -441.67999267578125, -513.260009765625, -592.1599731445312, -677.3099975585938, -767.489990234375, -861.4500122070312, -958.030029296875, -1056.280029296875, -1155.530029296875, -1255.5400390625, -1356.8699951171875, -1461.4300537109375, -1572.760009765625, -1695.5899658203125, -1834.6800537109375, -1993.6199951171875, -2174.449951171875, -2378.0, -2604.5, -2854.0, -3126.5, -3422.0, -3740.5, -4082.0, -4446.5, -4834.0, -5244.5, -5678.0, -6134.5], dtype='float64', name='Zu')) - ZlPandasIndex
PandasIndex(Float64Index([ 0.0, -10.0, -20.0, -30.0, -40.0, -50.0, -60.0, -70.0, -80.01000213623047, -90.04000091552734, -100.1500015258789, -110.47000122070312, -121.2699966430664, -133.02999877929688, -146.4499969482422, -162.49000549316406, -182.30999755859375, -207.16000366210938, -238.25999450683594, -276.67999267578125, -323.17999267578125, -378.17999267578125, -441.67999267578125, -513.260009765625, -592.1599731445312, -677.3099975585938, -767.489990234375, -861.4500122070312, -958.030029296875, -1056.280029296875, -1155.530029296875, -1255.5400390625, -1356.8699951171875, -1461.4300537109375, -1572.760009765625, -1695.5899658203125, -1834.6800537109375, -1993.6199951171875, -2174.449951171875, -2378.0, -2604.5, -2854.0, -3126.5, -3422.0, -3740.5, -4082.0, -4446.5, -4834.0, -5244.5, -5678.0], dtype='float64', name='Zl')) - timePandasIndex
PandasIndex(TimedeltaIndex(['0 days 00:00:01', '0 days 00:00:02', '0 days 00:00:03', '0 days 00:00:04', '0 days 00:00:05', '0 days 00:00:06', '0 days 00:00:07', '0 days 00:00:08'], dtype='timedelta64[ns]', name='time', freq=None))
- Conventions :
- CF-1.6
- title :
- netCDF wrapper of MITgcm MDS binary data
- source :
- MITgcm
- history :
- Created by calling `open_mdsdataset(grid_dir='/glade/work/dcherian/mitgcm/ECCOV4/release4/run/diags/../', iters=None, prefix=None, read_grid=True, delta_t=1, ref_date=None, calendar='gregorian', geometry='sphericalpolar', grid_vars_to_coords=False, swap_dims=False, endian='>', chunks={}, ignore_unknown_vars=False, default_dtype=None, nx=None, ny=None, nz=None, llc_method='smallchunks', extra_metadata=None, extra_variables=None, data_dir='/glade/work/dcherian/mitgcm/ECCOV4/release4/run/diags/GM_CHIZ_INST/', levels=None)`
chiz = ecco.llc_tiles_to_xda(
ecco.read_llc_to_tiles(
dirname + "GM_CHIZ_INST/",
"GM_CHIZ.0000000001.data",
nk=50,
nl=1,
use_xmitgcm=True,
),
var_type="c",
dim4="depth",
dim5="time",
)
chiz
read_llc_to_tiles: full_filename: /glade/work/dcherian/mitgcm/ECCOV4/release4/run/diags/GM_CHIZ_INST//GM_CHIZ.0000000001.data
<xarray.DataArray 'llc-2d8997a5c3544544991ec6632d79eee9' (time: 1, k: 50,
tile: 13, j: 90, i: 90)>
dask.array<llc, shape=(1, 50, 13, 90, 90), dtype=>f4, chunksize=(1, 1, 1, 90, 90), chunktype=numpy.ndarray>
Coordinates:
* k (k) int64 0 1 2 3 4 5 6 7 8 9 10 ... 40 41 42 43 44 45 46 47 48 49
* time (time) int64 0
* tile (tile) int64 0 1 2 3 4 5 6 7 8 9 10 11 12
* j (j) int64 0 1 2 3 4 5 6 7 8 9 10 ... 80 81 82 83 84 85 86 87 88 89
* i (i) int64 0 1 2 3 4 5 6 7 8 9 10 ... 80 81 82 83 84 85 86 87 88 89xarray.DataArray
'llc-2d8997a5c3544544991ec6632d79eee9'
- time: 1
- k: 50
- tile: 13
- j: 90
- i: 90
- dask.array<chunksize=(1, 1, 1, 90, 90), meta=np.ndarray>
Array Chunk Bytes 20.08 MiB 31.64 kiB Shape (1, 50, 13, 90, 90) (1, 1, 1, 90, 90) Dask graph 650 chunks in 1 graph layer Data type >f4 numpy.ndarray - k(k)int640 1 2 3 4 5 6 ... 44 45 46 47 48 49
- dims :
- ['k']
- attrs :
- {'standard_name': 'z_grid_index', 'axis': 'Z', 'long_name': 'z-dimension of the t grid', 'swap_dim': 'Z'}
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49]) - time(time)int640
array([0])
- tile(tile)int640 1 2 3 4 5 6 7 8 9 10 11 12
- standard_name :
- tile_index
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12])
- j(j)int640 1 2 3 4 5 6 ... 84 85 86 87 88 89
- standard_name :
- y_grid_index
- axis :
- Y
- long_name :
- y-dimension of the t grid
- swap_dim :
- YC
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89]) - i(i)int640 1 2 3 4 5 6 ... 84 85 86 87 88 89
- standard_name :
- x_grid_index
- axis :
- X
- long_name :
- x-dimension of the t grid
- swap_dim :
- XC
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89])
- kPandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49], dtype='int64', name='k')) - timePandasIndex
PandasIndex(Int64Index([0], dtype='int64', name='time'))
- tilePandasIndex
PandasIndex(Int64Index([0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12], dtype='int64', name='tile'))
- jPandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89], dtype='int64', name='j')) - iPandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89], dtype='int64', name='i'))
chix.sel(tile=5).plot()
<matplotlib.collections.QuadMesh at 0x2b9eaf65a830>
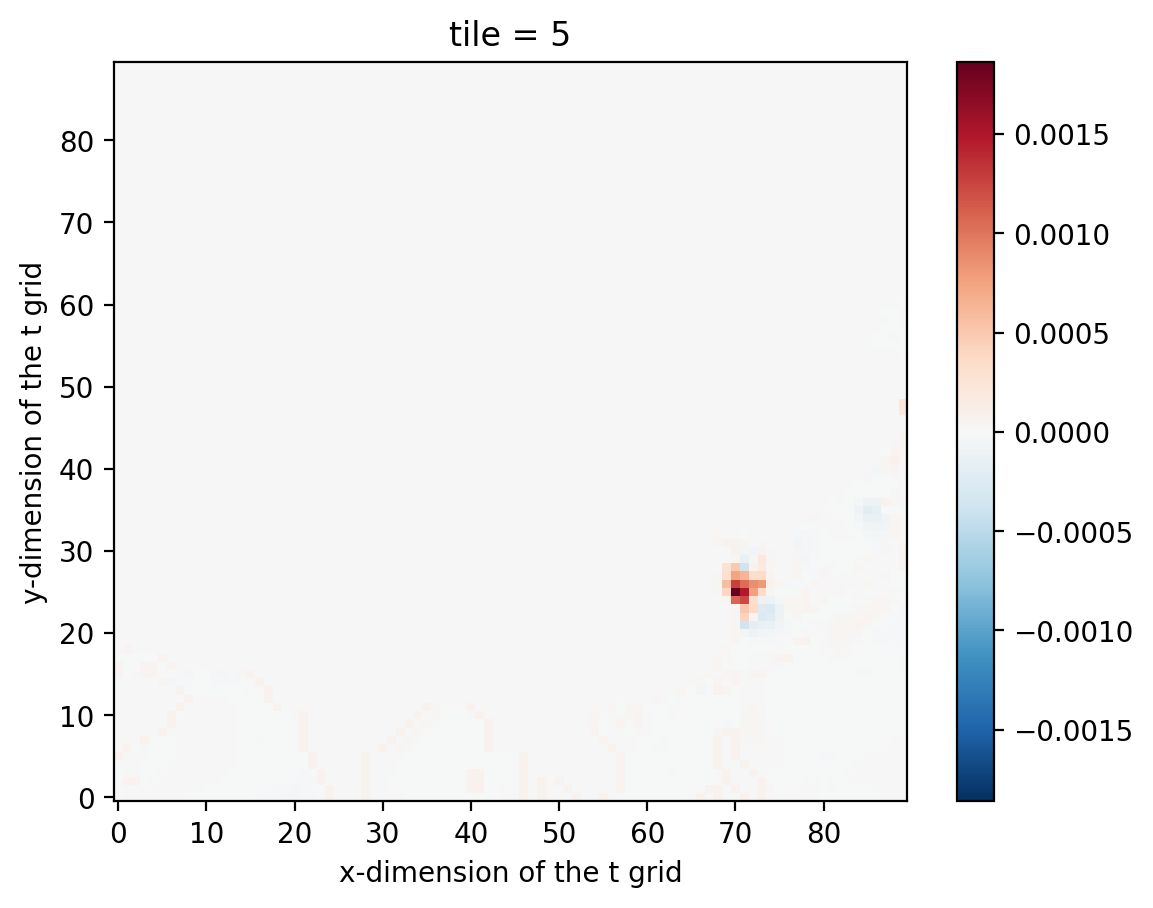
Tracking down gradient calculation in gmredi_ytransport.F#
Here, I saved the dTdz in gmredi_ytransport.F as GM_CHIYO
dTdz_v replicates the calculation.
This helped tracked down a bug in ds.drC_kl (we want drC.isel(kp1=slice(-1)) instead of drC.isel(kp1=slice(1, None))
ΔT_z = -1 * grid.diff(ds.THETA, "Z")
ΔT_z_y = grid.interp(ΔT_z, "Y")
dTdz_v = grid.interp(ΔT_z_y / ds.drC_kl, "Z")
(ds.GM_CHIYO).isel(time=5, j_g=21, i=20).plot()
(dTdz_v).isel(time=5, j_g=21, i=20).plot()
(grid.interp(ds.Tz, axis=["Z", "Y"])).isel(time=5, j_g=21, i=20).plot()
[<matplotlib.lines.Line2D at 0x2ab5ea93fd00>]
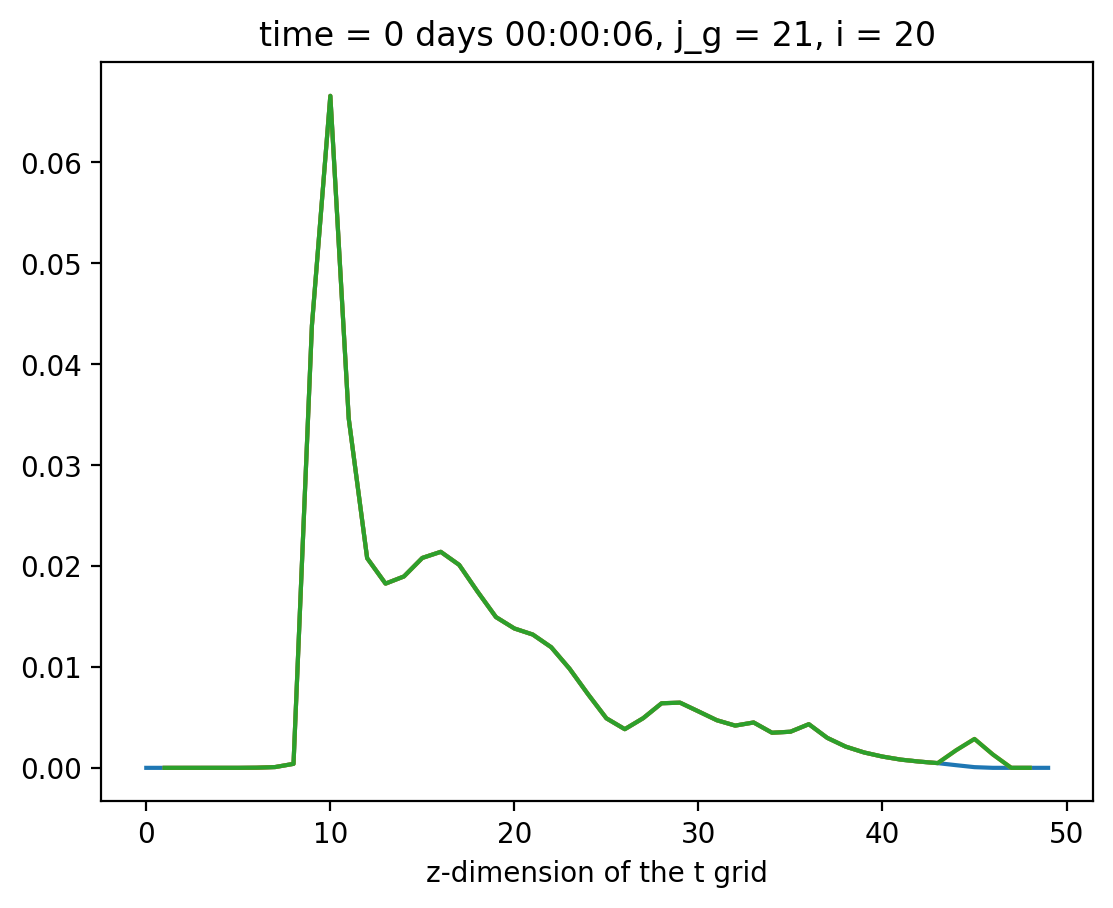
Test ECCO Kredi application#
In input_init/README(link) we see
total_diffkr_r009bit11.bin vert. diff. of release 1 (this field plus xx is the total)
total_kapgm_r009bit11.bin Kappa GM of release 1 (this field plus xx is the total)
total_kapredi_r009bit11.bin Kappa Redi of release 1 (this field plus xx is the total)
xx_*.* control adjustments
I suspect by commenting out the file name, I set a constant and the control adjustments were applied
And it turns out the control adjustments need to be rescaled so that makes it a pain to check.
It does approximately match so that seems to be what’s going on.
path = "/glade/u/home/dcherian/work/mitgcm/ECCOV4/release4/input_init/"
import xarray as xr
natre_mean = xr.open_dataset("../datasets/ecco-chi.nc").isel(time=1)
import numpy as np
import xmitgcm
kapredi_adj = xmitgcm.open_mdsdataset(
path,
grid_dir=path + "../run/",
prefix="xx_kapredi",
geometry="llc",
extra_variables=dict(xx_kapredi=dict(dims=["k", "j", "i"], attrs={})),
).load()
kapredi_adj
<xarray.Dataset>
Dimensions: (i: 90, i_g: 90, j: 90, j_g: 90, k: 50, k_u: 50, k_l: 50,
k_p1: 51, face: 13, time: 1)
Coordinates: (12/44)
* i (i) int64 0 1 2 3 4 5 6 7 8 9 ... 80 81 82 83 84 85 86 87 88 89
* i_g (i_g) int64 0 1 2 3 4 5 6 7 8 9 ... 81 82 83 84 85 86 87 88 89
* j (j) int64 0 1 2 3 4 5 6 7 8 9 ... 80 81 82 83 84 85 86 87 88 89
* j_g (j_g) int64 0 1 2 3 4 5 6 7 8 9 ... 81 82 83 84 85 86 87 88 89
* k (k) int64 0 1 2 3 4 5 6 7 8 9 ... 40 41 42 43 44 45 46 47 48 49
* k_u (k_u) int64 0 1 2 3 4 5 6 7 8 9 ... 41 42 43 44 45 46 47 48 49
... ...
maskW (k, face, j, i_g) bool False False False ... False False False
maskS (k, face, j_g, i) bool False False False ... False False False
maskCtrlC (k, face, j, i) bool False False False ... False False False
maskCtrlS (k, face, j_g, i) bool False False False ... False False False
rhoRef (k) >f4 1.024e+03 1.024e+03 1.024e+03 ... 1.052e+03 1.054e+03
maskCtrlW (k, face, j, i_g) bool False False False ... False False False
Data variables:
xx_kapredi (time, k, face, j, i) >f4 0.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0
Attributes:
Conventions: CF-1.6
title: netCDF wrapper of MITgcm MDS binary data
source: MITgcm
history: Created by calling `open_mdsdataset(grid_dir='/glade/u/home...xarray.Dataset
- i: 90
- i_g: 90
- j: 90
- j_g: 90
- k: 50
- k_u: 50
- k_l: 50
- k_p1: 51
- face: 13
- time: 1
- i(i)int640 1 2 3 4 5 6 ... 84 85 86 87 88 89
- standard_name :
- x_grid_index
- axis :
- X
- long_name :
- x-dimension of the t grid
- swap_dim :
- XC
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89]) - i_g(i_g)int640 1 2 3 4 5 6 ... 84 85 86 87 88 89
- standard_name :
- x_grid_index_at_u_location
- axis :
- X
- long_name :
- x-dimension of the u grid
- c_grid_axis_shift :
- -0.5
- swap_dim :
- XG
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89]) - j(j)int640 1 2 3 4 5 6 ... 84 85 86 87 88 89
- standard_name :
- y_grid_index
- axis :
- Y
- long_name :
- y-dimension of the t grid
- swap_dim :
- YC
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89]) - j_g(j_g)int640 1 2 3 4 5 6 ... 84 85 86 87 88 89
- standard_name :
- y_grid_index_at_v_location
- axis :
- Y
- long_name :
- y-dimension of the v grid
- c_grid_axis_shift :
- -0.5
- swap_dim :
- YG
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89]) - k(k)int640 1 2 3 4 5 6 ... 44 45 46 47 48 49
- standard_name :
- z_grid_index
- axis :
- Z
- long_name :
- z-dimension of the t grid
- swap_dim :
- Z
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49]) - k_u(k_u)int640 1 2 3 4 5 6 ... 44 45 46 47 48 49
- standard_name :
- z_grid_index_at_upper_w_location
- axis :
- Z
- long_name :
- z-dimension of the w grid
- c_grid_axis_shift :
- 0.5
- swap_dim :
- Zu
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49]) - k_l(k_l)int640 1 2 3 4 5 6 ... 44 45 46 47 48 49
- standard_name :
- z_grid_index_at_lower_w_location
- axis :
- Z
- long_name :
- z-dimension of the w grid
- c_grid_axis_shift :
- -0.5
- swap_dim :
- Zl
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49]) - k_p1(k_p1)int640 1 2 3 4 5 6 ... 45 46 47 48 49 50
- standard_name :
- z_grid_index_at_w_location
- axis :
- Z
- long_name :
- z-dimension of the w grid
- c_grid_axis_shift :
- (-0.5, 0.5)
- swap_dim :
- Zp1
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50]) - face(face)int640 1 2 3 4 5 6 7 8 9 10 11 12
- standard_name :
- face_index
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12])
- iter(time)int64129
- standard_name :
- timestep
- long_name :
- model timestep number
array([129])
- time(time)timedelta64[ns]00:02:09
- standard_name :
- time
- long_name :
- Time
- axis :
- T
- calendar :
- gregorian
array([129000000000], dtype='timedelta64[ns]')
- XC(face, j, i)>f40.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0
- standard_name :
- longitude
- long_name :
- longitude
- units :
- degrees_east
- coordinate :
- YC XC
array([[[ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], ..., [ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ], [ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ], [ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ]], [[ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ], [ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ], [ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ], ... [ -40.5 , -40.5 , -40.5 , ..., -40.5 , -40.5 , -40.5 ], [ -39.5 , -39.5 , -39.5 , ..., -39.5 , -39.5 , -39.5 ], [ -38.5 , -38.5 , -38.5 , ..., -38.5 , -38.5 , -38.5 ]], [[-127.5 , -127.5 , -127.5 , ..., 0. , 0. , 0. ], [-126.5 , -126.5 , -126.5 , ..., 0. , 0. , 0. ], [-125.5 , -125.5 , -125.5 , ..., 0. , 0. , 0. ], ..., [ -40.5 , -40.5 , -40.5 , ..., 0. , 0. , 0. ], [ -39.5 , -39.5 , -39.5 , ..., 0. , 0. , 0. ], [ -38.5 , -38.5 , -38.5 , ..., 0. , 0. , 0. ]]], dtype=float32) - YC(face, j, i)>f40.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0
- standard_name :
- latitude
- long_name :
- latitude
- units :
- degrees_north
- coordinate :
- YC XC
array([[[ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], ..., [-58.321964 , -58.321964 , -58.321964 , ..., -58.321964 , -58.321964 , -58.321964 ], [-57.79962 , -57.79962 , -57.79962 , ..., -57.79962 , -57.79962 , -57.79962 ], [-57.271408 , -57.271408 , -57.271408 , ..., -57.271408 , -57.271408 , -57.271408 ]], [[-56.73891 , -56.73891 , -56.73891 , ..., -56.73891 , -56.73891 , -56.73891 ], [-56.2021 , -56.2021 , -56.2021 , ..., -56.2021 , -56.2021 , -56.2021 ], [-55.65936 , -55.65936 , -55.65936 , ..., -55.65936 , -55.65936 , -55.65936 ], ... [ 9.482398 , 8.516253 , 7.5699615, ..., -55.65936 , -56.2021 , -56.73891 ], [ 9.482398 , 8.516253 , 7.5699615, ..., -55.65936 , -56.2021 , -56.73891 ], [ 9.482398 , 8.516253 , 7.5699615, ..., -55.65936 , -56.2021 , -56.73891 ]], [[-57.271408 , -57.79962 , -58.321964 , ..., 0. , 0. , 0. ], [-57.271408 , -57.79962 , -58.321964 , ..., 0. , 0. , 0. ], [-57.271408 , -57.79962 , -58.321964 , ..., 0. , 0. , 0. ], ..., [-57.271408 , -57.79962 , -58.321964 , ..., 0. , 0. , 0. ], [-57.271408 , -57.79962 , -58.321964 , ..., 0. , 0. , 0. ], [-57.271408 , -57.79962 , -58.321964 , ..., 0. , 0. , 0. ]]], dtype=float32) - XG(face, j_g, i_g)>f40.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0
- standard_name :
- longitude_at_f_location
- long_name :
- longitude
- units :
- degrees_east
- coordinate :
- YG XG
array([[[ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], ..., [ -38. , -37. , -36. , ..., 49. , 50. , 51. ], [ -38. , -37. , -36. , ..., 49. , 50. , 51. ], [ -38. , -37. , -36. , ..., 49. , 50. , 51. ]], [[ -38. , -37. , -36. , ..., 49. , 50. , 51. ], [ -38. , -37. , -36. , ..., 49. , 50. , 51. ], [ -38. , -37. , -36. , ..., 49. , 50. , 51. ], ... [ -41. , -41. , -41. , ..., -41. , -41. , -41. ], [ -40. , -40. , -40. , ..., -40. , -40. , -40. ], [ -39. , -39. , -39. , ..., -39. , -39. , -39. ]], [[-128. , -128. , -128. , ..., 0. , 0. , 0. ], [-127. , -127. , -127. , ..., 0. , 0. , 0. ], [-126. , -126. , -126. , ..., 0. , 0. , 0. ], ..., [ -41. , -41. , -41. , ..., 0. , 0. , 0. ], [ -40. , -40. , -40. , ..., 0. , 0. , 0. ], [ -39. , -39. , -39. , ..., 0. , 0. , 0. ]]], dtype=float32) - YG(face, j_g, i_g)>f40.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0
- standard_name :
- latitude_at_f_location
- long_name :
- latitude
- units :
- degrees_north
- coordinate :
- YG XG
array([[[ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], ..., [-58.58002 , -58.58002 , -58.58002 , ..., -58.58002 , -58.58002 , -58.58002 ], [-58.06183 , -58.06183 , -58.06183 , ..., -58.06183 , -58.06183 , -58.06183 ], [-57.53605 , -57.53605 , -57.53605 , ..., -57.53605 , -57.53605 , -57.53605 ]], [[-57.005695, -57.005695, -57.005695, ..., -57.005695, -57.005695, -57.005695], [-56.471046, -56.471046, -56.471046, ..., -56.471046, -56.471046, -56.471046], [-55.931786, -55.931786, -55.931786, ..., -55.931786, -55.931786, -55.931786], ... [ 9.96973 , 8.997536, 8.039881, ..., -55.384766, -55.931786, -56.471046], [ 9.96973 , 8.997536, 8.039881, ..., -55.384766, -55.931786, -56.471046], [ 9.96973 , 8.997536, 8.039881, ..., -55.384766, -55.931786, -56.471046]], [[-57.005695, -57.53605 , -58.06183 , ..., 0. , 0. , 0. ], [-57.005695, -57.53605 , -58.06183 , ..., 0. , 0. , 0. ], [-57.005695, -57.53605 , -58.06183 , ..., 0. , 0. , 0. ], ..., [-57.005695, -57.53605 , -58.06183 , ..., 0. , 0. , 0. ], [-57.005695, -57.53605 , -58.06183 , ..., 0. , 0. , 0. ], [-57.005695, -57.53605 , -58.06183 , ..., 0. , 0. , 0. ]]], dtype=float32) - CS(face, j, i)>f40.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0
- standard_name :
- Cos of grid orientation angle
- long_name :
- AngleCS
- units :
- coordinate :
- YC XC
array([[[ 0.0000000e+00, 0.0000000e+00, 0.0000000e+00, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], [ 0.0000000e+00, 0.0000000e+00, 0.0000000e+00, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], [ 0.0000000e+00, 0.0000000e+00, 0.0000000e+00, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], ..., [ 1.0000000e+00, 1.0000000e+00, 1.0000000e+00, ..., 1.0000000e+00, 1.0000000e+00, 1.0000000e+00], [ 1.0000000e+00, 1.0000000e+00, 1.0000000e+00, ..., 1.0000000e+00, 1.0000000e+00, 1.0000000e+00], [ 1.0000000e+00, 1.0000000e+00, 1.0000000e+00, ..., 1.0000000e+00, 1.0000000e+00, 1.0000000e+00]], [[ 1.0000000e+00, 1.0000000e+00, 1.0000000e+00, ..., 1.0000000e+00, 1.0000000e+00, 1.0000000e+00], [ 1.0000000e+00, 1.0000000e+00, 1.0000000e+00, ..., 1.0000000e+00, 1.0000000e+00, 1.0000000e+00], [ 1.0000000e+00, 1.0000000e+00, 1.0000000e+00, ..., 1.0000000e+00, 1.0000000e+00, 1.0000000e+00], ... [-1.8010396e-15, -1.7962386e-15, -8.9699497e-16, ..., -6.3420986e-15, -6.3420986e-15, 6.5240599e-15], [ 8.9924353e-16, 1.7962386e-15, 1.3445239e-15, ..., -0.0000000e+00, -0.0000000e+00, -6.5240599e-15], [ 2.5525710e-18, -8.9924353e-16, -4.4752880e-16, ..., 1.2508176e-14, -0.0000000e+00, -0.0000000e+00]], [[-0.0000000e+00, 1.3431713e-14, 1.3431713e-14, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], [ 6.5240599e-15, -1.3431713e-14, -6.6167042e-15, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], [-6.5240599e-15, -0.0000000e+00, -6.8150090e-15, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], ..., [ 6.5240599e-15, -0.0000000e+00, 6.8150090e-15, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], [-6.5240599e-15, 1.3431713e-14, 6.6167042e-15, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], [-0.0000000e+00, -1.3431713e-14, -1.3431713e-14, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00]]], dtype=float32) - SN(face, j, i)>f40.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0
- standard_name :
- Sin of grid orientation angle
- long_name :
- AngleSN
- units :
- coordinate :
- YC XC
array([[[ 0.0000000e+00, 0.0000000e+00, 0.0000000e+00, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], [ 0.0000000e+00, 0.0000000e+00, 0.0000000e+00, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], [ 0.0000000e+00, 0.0000000e+00, 0.0000000e+00, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], ..., [ 1.3431713e-14, -6.6167042e-15, -6.8150090e-15, ..., 6.8150090e-15, 6.6167042e-15, -1.3431713e-14], [ 1.3431713e-14, -1.3431713e-14, -0.0000000e+00, ..., -0.0000000e+00, 1.3431713e-14, -1.3431713e-14], [-0.0000000e+00, 6.5240599e-15, -6.5240599e-15, ..., 6.5240599e-15, -6.5240599e-15, -0.0000000e+00]], [[-0.0000000e+00, 6.5240599e-15, -6.5240599e-15, ..., 6.5240599e-15, -6.5240599e-15, -0.0000000e+00], [-0.0000000e+00, -0.0000000e+00, 6.3420986e-15, ..., -6.3420986e-15, -0.0000000e+00, -0.0000000e+00], [-1.2508176e-14, -0.0000000e+00, 6.3420986e-15, ..., -6.3420986e-15, -0.0000000e+00, 1.2508176e-14], ... [-1.0000000e+00, -1.0000000e+00, -1.0000000e+00, ..., -1.0000000e+00, -1.0000000e+00, -1.0000000e+00], [-1.0000000e+00, -1.0000000e+00, -1.0000000e+00, ..., -1.0000000e+00, -1.0000000e+00, -1.0000000e+00], [-1.0000000e+00, -1.0000000e+00, -1.0000000e+00, ..., -1.0000000e+00, -1.0000000e+00, -1.0000000e+00]], [[-1.0000000e+00, -1.0000000e+00, -1.0000000e+00, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], [-1.0000000e+00, -1.0000000e+00, -1.0000000e+00, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], [-1.0000000e+00, -1.0000000e+00, -1.0000000e+00, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], ..., [-1.0000000e+00, -1.0000000e+00, -1.0000000e+00, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], [-1.0000000e+00, -1.0000000e+00, -1.0000000e+00, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], [-1.0000000e+00, -1.0000000e+00, -1.0000000e+00, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00]]], dtype=float32) - Z(k)>f4-5.0 -15.0 ... -5.906e+03
- standard_name :
- depth
- long_name :
- vertical coordinate of cell center
- units :
- m
- positive :
- down
array([-5.000000e+00, -1.500000e+01, -2.500000e+01, -3.500000e+01, -4.500000e+01, -5.500000e+01, -6.500000e+01, -7.500500e+01, -8.502500e+01, -9.509500e+01, -1.053100e+02, -1.158700e+02, -1.271500e+02, -1.397400e+02, -1.544700e+02, -1.724000e+02, -1.947350e+02, -2.227100e+02, -2.574700e+02, -2.999300e+02, -3.506800e+02, -4.099300e+02, -4.774700e+02, -5.527100e+02, -6.347350e+02, -7.224000e+02, -8.144700e+02, -9.097400e+02, -1.007155e+03, -1.105905e+03, -1.205535e+03, -1.306205e+03, -1.409150e+03, -1.517095e+03, -1.634175e+03, -1.765135e+03, -1.914150e+03, -2.084035e+03, -2.276225e+03, -2.491250e+03, -2.729250e+03, -2.990250e+03, -3.274250e+03, -3.581250e+03, -3.911250e+03, -4.264250e+03, -4.640250e+03, -5.039250e+03, -5.461250e+03, -5.906250e+03], dtype=float32) - Zp1(k_p1)>f40.0 -10.0 ... -5.678e+03 -6.134e+03
- standard_name :
- depth_at_w_location
- long_name :
- vertical coordinate of cell interface
- units :
- m
- positive :
- down
array([ 0. , -10. , -20. , -30. , -40. , -50. , -60. , -70. , -80.01, -90.04, -100.15, -110.47, -121.27, -133.03, -146.45, -162.49, -182.31, -207.16, -238.26, -276.68, -323.18, -378.18, -441.68, -513.26, -592.16, -677.31, -767.49, -861.45, -958.03, -1056.28, -1155.53, -1255.54, -1356.87, -1461.43, -1572.76, -1695.59, -1834.68, -1993.62, -2174.45, -2378. , -2604.5 , -2854. , -3126.5 , -3422. , -3740.5 , -4082. , -4446.5 , -4834. , -5244.5 , -5678. , -6134.5 ], dtype=float32) - Zu(k_u)>f4-10.0 -20.0 ... -6.134e+03
- standard_name :
- depth_at_upper_w_location
- long_name :
- vertical coordinate of upper cell interface
- units :
- m
- positive :
- down
array([ -10. , -20. , -30. , -40. , -50. , -60. , -70. , -80.01, -90.04, -100.15, -110.47, -121.27, -133.03, -146.45, -162.49, -182.31, -207.16, -238.26, -276.68, -323.18, -378.18, -441.68, -513.26, -592.16, -677.31, -767.49, -861.45, -958.03, -1056.28, -1155.53, -1255.54, -1356.87, -1461.43, -1572.76, -1695.59, -1834.68, -1993.62, -2174.45, -2378. , -2604.5 , -2854. , -3126.5 , -3422. , -3740.5 , -4082. , -4446.5 , -4834. , -5244.5 , -5678. , -6134.5 ], dtype=float32) - Zl(k_l)>f40.0 -10.0 ... -5.244e+03 -5.678e+03
- standard_name :
- depth_at_lower_w_location
- long_name :
- vertical coordinate of lower cell interface
- units :
- m
- positive :
- down
array([ 0. , -10. , -20. , -30. , -40. , -50. , -60. , -70. , -80.01, -90.04, -100.15, -110.47, -121.27, -133.03, -146.45, -162.49, -182.31, -207.16, -238.26, -276.68, -323.18, -378.18, -441.68, -513.26, -592.16, -677.31, -767.49, -861.45, -958.03, -1056.28, -1155.53, -1255.54, -1356.87, -1461.43, -1572.76, -1695.59, -1834.68, -1993.62, -2174.45, -2378. , -2604.5 , -2854. , -3126.5 , -3422. , -3740.5 , -4082. , -4446.5 , -4834. , -5244.5 , -5678. ], dtype=float32) - rA(face, j, i)>f40.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0
- standard_name :
- cell_area
- long_name :
- cell area
- units :
- m2
- coordinate :
- YC XC
array([[[0.0000000e+00, 0.0000000e+00, 0.0000000e+00, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], [0.0000000e+00, 0.0000000e+00, 0.0000000e+00, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], [0.0000000e+00, 0.0000000e+00, 0.0000000e+00, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], ..., [3.3636692e+09, 3.3636692e+09, 3.3636692e+09, ..., 3.3636692e+09, 3.3636692e+09, 3.3636692e+09], [3.4631982e+09, 3.4631982e+09, 3.4631982e+09, ..., 3.4631982e+09, 3.4631982e+09, 3.4631982e+09], [3.5442668e+09, 3.5442668e+09, 3.5442668e+09, ..., 3.5442668e+09, 3.5442668e+09, 3.5442668e+09]], [[3.6245120e+09, 3.6245120e+09, 3.6245120e+09, ..., 3.6245120e+09, 3.6245120e+09, 3.6245120e+09], [3.7078205e+09, 3.7078205e+09, 3.7078205e+09, ..., 3.7078205e+09, 3.7078205e+09, 3.7078205e+09], [3.8142915e+09, 3.8142915e+09, 3.8142915e+09, ..., 3.8142915e+09, 3.8142915e+09, 3.8142915e+09], ... [1.1852359e+10, 1.1706299e+10, 1.1415347e+10, ..., 3.8142915e+09, 3.7078205e+09, 3.6245120e+09], [1.1852359e+10, 1.1706299e+10, 1.1415347e+10, ..., 3.8142915e+09, 3.7078205e+09, 3.6245120e+09], [1.1852359e+10, 1.1706299e+10, 1.1415347e+10, ..., 3.8142915e+09, 3.7078205e+09, 3.6245120e+09]], [[3.5442668e+09, 3.4631982e+09, 3.3636692e+09, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], [3.5442668e+09, 3.4631982e+09, 3.3636692e+09, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], [3.5442668e+09, 3.4631982e+09, 3.3636692e+09, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], ..., [3.5442668e+09, 3.4631982e+09, 3.3636692e+09, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], [3.5442668e+09, 3.4631982e+09, 3.3636692e+09, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], [3.5442668e+09, 3.4631982e+09, 3.3636692e+09, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00]]], dtype=float32) - dxG(face, j_g, i)>f40.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0
- standard_name :
- cell_x_size_at_v_location
- long_name :
- cell x size
- units :
- m
- coordinate :
- YG XC
array([[[ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], ..., [ 57957.625, 57957.625, 57957.625, ..., 57957.625, 57957.625, 57957.625], [ 58813.305, 58813.305, 58813.305, ..., 58813.305, 58813.305, 58813.305], [ 59676.61 , 59676.61 , 59676.61 , ..., 59676.61 , 59676.61 , 59676.61 ]], [[ 60542.324, 60542.324, 60542.324, ..., 60542.324, 60542.324, 60542.324], [ 61409.805, 61409.805, 61409.805, ..., 61409.805, 61409.805, 61409.805], [ 62279.344, 62279.344, 62279.344, ..., 62279.344, 62279.344, 62279.344], ... [108086.08 , 106469.66 , 103581.64 , ..., 60816.504, 59953.41 , 59441.125], [108086.08 , 106469.66 , 103581.64 , ..., 60816.504, 59953.41 , 59441.125], [108086.08 , 106469.66 , 103581.64 , ..., 60816.504, 59953.41 , 59441.125]], [[ 58963.117, 58455.164, 57611.023, ..., 0. , 0. , 0. ], [ 58963.117, 58455.164, 57611.023, ..., 0. , 0. , 0. ], [ 58963.117, 58455.164, 57611.023, ..., 0. , 0. , 0. ], ..., [ 58963.117, 58455.164, 57611.023, ..., 0. , 0. , 0. ], [ 58963.117, 58455.164, 57611.023, ..., 0. , 0. , 0. ], [ 58963.117, 58455.164, 57611.023, ..., 0. , 0. , 0. ]]], dtype=float32) - dyG(face, j, i_g)>f40.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0
- standard_name :
- cell_y_size_at_u_location
- long_name :
- cell y size
- units :
- m
- coordinate :
- YC XG
array([[[ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], ..., [ 57611.023, 57611.023, 57611.023, ..., 57611.023, 57611.023, 57611.023], [ 58455.164, 58455.164, 58455.164, ..., 58455.164, 58455.164, 58455.164], [ 58963.117, 58963.117, 58963.117, ..., 58963.117, 58963.117, 58963.117]], [[ 59441.125, 59441.125, 59441.125, ..., 59441.125, 59441.125, 59441.125], [ 59953.41 , 59953.41 , 59953.41 , ..., 59953.41 , 59953.41 , 59953.41 ], [ 60816.504, 60816.504, 60816.504, ..., 60816.504, 60816.504, 60816.504], ... [109498.625, 109809.445, 110084.7 , ..., 63155.766, 62279.344, 61409.805], [109498.625, 109809.445, 110084.7 , ..., 63155.766, 62279.344, 61409.805], [109498.625, 109809.445, 110084.7 , ..., 63155.766, 62279.344, 61409.805]], [[ 60542.324, 59676.61 , 58813.305, ..., 0. , 0. , 0. ], [ 60542.324, 59676.61 , 58813.305, ..., 0. , 0. , 0. ], [ 60542.324, 59676.61 , 58813.305, ..., 0. , 0. , 0. ], ..., [ 60542.324, 59676.61 , 58813.305, ..., 0. , 0. , 0. ], [ 60542.324, 59676.61 , 58813.305, ..., 0. , 0. , 0. ], [ 60542.324, 59676.61 , 58813.305, ..., 0. , 0. , 0. ]]], dtype=float32) - Depth(face, j, i)>f40.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0
- standard_name :
- ocean_depth
- long_name :
- ocean depth
- units :
- m
- coordinate :
- XC YC
array([[[ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], ..., [2977.8345 , 2503.9146 , 2562.9644 , ..., 5431.1567 , 5356.7544 , 5331.2 ], [3125.5176 , 2908.5 , 3046.7676 , ..., 5440.6733 , 5383.5386 , 5332.443 ], [3203.0798 , 3283.116 , 3312.7979 , ..., 5455.109 , 5433.769 , 5367.7656 ]], [[3284.1084 , 3485.7 , 3485.7 , ..., 5455.109 , 5444.462 , 5387.942 ], [3350.4321 , 3403.6208 , 3485.7 , ..., 5367.847 , 5379.0474 , 5359.3574 ], [3324.7493 , 3126.5 , 2243.637 , ..., 5244.5 , 5244.5 , 5196.211 ], ... [3384.1433 , 3728.414 , 4524. , ..., 3185.6 , 3107.3855 , 3246.3552 ], [3638.9377 , 3570.711 , 4352.1484 , ..., 3031.5972 , 2908.5 , 2908.5 ], [3965.616 , 3676.4983 , 4082. , ..., 3285.413 , 3066.3777 , 3055.8303 ]], [[3920.2725 , 3809.5889 , 3877.8499 , ..., 0. , 0. , 0. ], [3984.9038 , 3906.704 , 3950.57 , ..., 0. , 0. , 0. ], [3982.3193 , 3961.0894 , 4053.4758 , ..., 0. , 0. , 0. ], ..., [3374.1394 , 3362.2715 , 3243.364 , ..., 0. , 0. , 0. ], [2949.6958 , 3126.5 , 3213.2534 , ..., 0. , 0. , 0. ], [3076.3447 , 3119.1152 , 3126.5 , ..., 0. , 0. , 0. ]]], dtype=float32) - rAz(face, j_g, i_g)>f40.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0
- standard_name :
- cell_area_at_f_location
- long_name :
- cell area
- units :
- m
- coordinate :
- YG XG
array([[[0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], ..., [3.31241549e+09, 3.31241549e+09, 3.31241549e+09, ..., 3.31241549e+09, 3.31241549e+09, 3.31241549e+09], [3.41554739e+09, 3.41554739e+09, 3.41554739e+09, ..., 3.41554739e+09, 3.41554739e+09, 3.41554739e+09], [3.50457702e+09, 3.50457702e+09, 3.50457702e+09, ..., 3.50457702e+09, 3.50457702e+09, 3.50457702e+09]], [[3.58424499e+09, 3.58424499e+09, 3.58424499e+09, ..., 3.58424499e+09, 3.58424499e+09, 3.58424499e+09], [3.66506675e+09, 3.66506675e+09, 3.66506675e+09, ..., 3.66506675e+09, 3.66506675e+09, 3.66506675e+09], [3.75804186e+09, 3.75804186e+09, 3.75804186e+09, ..., 3.75804186e+09, 3.75804186e+09, 3.75804186e+09], ... [1.18844058e+10, 1.17948221e+10, 1.15813806e+10, ..., 3.87148928e+09, 3.75804186e+09, 3.66506675e+09], [1.18844058e+10, 1.17948221e+10, 1.15813806e+10, ..., 3.87148928e+09, 3.75804186e+09, 3.66506675e+09], [1.18844058e+10, 1.17948221e+10, 1.15813806e+10, ..., 3.87148928e+09, 3.75804186e+09, 3.66506675e+09]], [[3.58424499e+09, 3.50457702e+09, 3.41554739e+09, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [3.58424499e+09, 3.50457702e+09, 3.41554739e+09, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [3.58424499e+09, 3.50457702e+09, 3.41554739e+09, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], ..., [3.58424499e+09, 3.50457702e+09, 3.41554739e+09, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [3.58424499e+09, 3.50457702e+09, 3.41554739e+09, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [3.58424499e+09, 3.50457702e+09, 3.41554739e+09, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00]]], dtype=float32) - dxC(face, j, i_g)>f40.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0
- standard_name :
- cell_x_size_at_u_location
- long_name :
- cell x size
- units :
- m
- coordinate :
- YC XG
array([[[ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], ..., [ 58384.344, 58384.344, 58384.344, ..., 58384.344, 58384.344, 58384.344], [ 59244.46 , 59244.46 , 59244.46 , ..., 59244.46 , 59244.46 , 59244.46 ], [ 60109.234, 60109.234, 60109.234, ..., 60109.234, 60109.234, 60109.234]], [[ 60975.85 , 60975.85 , 60975.85 , ..., 60975.85 , 60975.85 , 60975.85 ], [ 61844.16 , 61844.16 , 61844.16 , ..., 61844.16 , 61844.16 , 61844.16 ], [ 62716.535, 62716.535, 62716.535, ..., 62716.535, 62716.535, 62716.535], ... [108536.33 , 107413.53 , 105206.3 , ..., 61299.195, 60340.285, 59681.477], [108536.33 , 107413.53 , 105206.3 , ..., 61299.195, 60340.285, 59681.477], [108536.33 , 107413.53 , 105206.3 , ..., 61299.195, 60340.285, 59681.477]], [[ 59201.66 , 58725.49 , 58072.91 , ..., 0. , 0. , 0. ], [ 59201.66 , 58725.49 , 58072.91 , ..., 0. , 0. , 0. ], [ 59201.66 , 58725.49 , 58072.91 , ..., 0. , 0. , 0. ], ..., [ 59201.66 , 58725.49 , 58072.91 , ..., 0. , 0. , 0. ], [ 59201.66 , 58725.49 , 58072.91 , ..., 0. , 0. , 0. ], [ 59201.66 , 58725.49 , 58072.91 , ..., 0. , 0. , 0. ]]], dtype=float32) - dyC(face, j_g, i)>f40.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0
- standard_name :
- cell_y_size_at_v_location
- long_name :
- cell y size
- units :
- m
- coordinate :
- YG XC
array([[[ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], ..., [ 57150.88 , 57150.88 , 57150.88 , ..., 57150.88 , 57150.88 , 57150.88 ], [ 58072.91 , 58072.91 , 58072.91 , ..., 58072.91 , 58072.91 , 58072.91 ], [ 58725.49 , 58725.49 , 58725.49 , ..., 58725.49 , 58725.49 , 58725.49 ]], [[ 59201.66 , 59201.66 , 59201.66 , ..., 59201.66 , 59201.66 , 59201.66 ], [ 59681.477, 59681.477, 59681.477, ..., 59681.477, 59681.477, 59681.477], [ 60340.285, 60340.285, 60340.285, ..., 60340.285, 60340.285, 60340.285], ... [109658.375, 109951.62 , 110208.53 , ..., 62716.535, 61844.16 , 60975.85 ], [109658.375, 109951.62 , 110208.53 , ..., 62716.535, 61844.16 , 60975.85 ], [109658.375, 109951.62 , 110208.53 , ..., 62716.535, 61844.16 , 60975.85 ]], [[ 60109.234, 59244.46 , 58384.344, ..., 0. , 0. , 0. ], [ 60109.234, 59244.46 , 58384.344, ..., 0. , 0. , 0. ], [ 60109.234, 59244.46 , 58384.344, ..., 0. , 0. , 0. ], ..., [ 60109.234, 59244.46 , 58384.344, ..., 0. , 0. , 0. ], [ 60109.234, 59244.46 , 58384.344, ..., 0. , 0. , 0. ], [ 60109.234, 59244.46 , 58384.344, ..., 0. , 0. , 0. ]]], dtype=float32) - rAw(face, j, i_g)>f40.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0
- standard_name :
- cell_area_at_u_location
- long_name :
- cell area
- units :
- m2
- coordinate :
- YG XC
array([[[0.0000000e+00, 0.0000000e+00, 0.0000000e+00, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], [0.0000000e+00, 0.0000000e+00, 0.0000000e+00, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], [0.0000000e+00, 0.0000000e+00, 0.0000000e+00, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], ..., [3.3636692e+09, 3.3636692e+09, 3.3636692e+09, ..., 3.3636692e+09, 3.3636692e+09, 3.3636692e+09], [3.4631982e+09, 3.4631982e+09, 3.4631982e+09, ..., 3.4631982e+09, 3.4631982e+09, 3.4631982e+09], [3.5442668e+09, 3.5442668e+09, 3.5442668e+09, ..., 3.5442668e+09, 3.5442668e+09, 3.5442668e+09]], [[3.6245120e+09, 3.6245120e+09, 3.6245120e+09, ..., 3.6245120e+09, 3.6245120e+09, 3.6245120e+09], [3.7078205e+09, 3.7078205e+09, 3.7078205e+09, ..., 3.7078205e+09, 3.7078205e+09, 3.7078205e+09], [3.8142915e+09, 3.8142915e+09, 3.8142915e+09, ..., 3.8142915e+09, 3.8142915e+09, 3.8142915e+09], ... [1.1884406e+10, 1.1794822e+10, 1.1581381e+10, ..., 3.8714893e+09, 3.7580419e+09, 3.6650668e+09], [1.1884406e+10, 1.1794822e+10, 1.1581381e+10, ..., 3.8714893e+09, 3.7580419e+09, 3.6650668e+09], [1.1884406e+10, 1.1794822e+10, 1.1581381e+10, ..., 3.8714893e+09, 3.7580419e+09, 3.6650668e+09]], [[3.5842450e+09, 3.5045770e+09, 3.4155474e+09, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], [3.5842450e+09, 3.5045770e+09, 3.4155474e+09, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], [3.5842450e+09, 3.5045770e+09, 3.4155474e+09, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], ..., [3.5842450e+09, 3.5045770e+09, 3.4155474e+09, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], [3.5842450e+09, 3.5045770e+09, 3.4155474e+09, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], [3.5842450e+09, 3.5045770e+09, 3.4155474e+09, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00]]], dtype=float32) - rAs(face, j_g, i)>f40.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0
- standard_name :
- cell_area_at_v_location
- long_name :
- cell area
- units :
- m2
- coordinate :
- YG XC
array([[[0.0000000e+00, 0.0000000e+00, 0.0000000e+00, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], [0.0000000e+00, 0.0000000e+00, 0.0000000e+00, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], [0.0000000e+00, 0.0000000e+00, 0.0000000e+00, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], ..., [3.3124155e+09, 3.3124155e+09, 3.3124155e+09, ..., 3.3124155e+09, 3.3124155e+09, 3.3124155e+09], [3.4155474e+09, 3.4155474e+09, 3.4155474e+09, ..., 3.4155474e+09, 3.4155474e+09, 3.4155474e+09], [3.5045770e+09, 3.5045770e+09, 3.5045770e+09, ..., 3.5045770e+09, 3.5045770e+09, 3.5045770e+09]], [[3.5842450e+09, 3.5842450e+09, 3.5842450e+09, ..., 3.5842450e+09, 3.5842450e+09, 3.5842450e+09], [3.6650668e+09, 3.6650668e+09, 3.6650668e+09, ..., 3.6650668e+09, 3.6650668e+09, 3.6650668e+09], [3.7580419e+09, 3.7580419e+09, 3.7580419e+09, ..., 3.7580419e+09, 3.7580419e+09, 3.7580419e+09], ... [1.1852359e+10, 1.1706299e+10, 1.1415347e+10, ..., 3.8142915e+09, 3.7078205e+09, 3.6245120e+09], [1.1852359e+10, 1.1706299e+10, 1.1415347e+10, ..., 3.8142915e+09, 3.7078205e+09, 3.6245120e+09], [1.1852359e+10, 1.1706299e+10, 1.1415347e+10, ..., 3.8142915e+09, 3.7078205e+09, 3.6245120e+09]], [[3.5442668e+09, 3.4631982e+09, 3.3636692e+09, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], [3.5442668e+09, 3.4631982e+09, 3.3636692e+09, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], [3.5442668e+09, 3.4631982e+09, 3.3636692e+09, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], ..., [3.5442668e+09, 3.4631982e+09, 3.3636692e+09, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], [3.5442668e+09, 3.4631982e+09, 3.3636692e+09, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00], [3.5442668e+09, 3.4631982e+09, 3.3636692e+09, ..., 0.0000000e+00, 0.0000000e+00, 0.0000000e+00]]], dtype=float32) - drC(k_p1)>f45.0 10.0 10.0 ... 422.0 445.0 228.2
- standard_name :
- cell_z_size_at_w_location
- long_name :
- cell z size
- units :
- m
array([ 5. , 10. , 10. , 10. , 10. , 10. , 10. , 10.005, 10.02 , 10.07 , 10.215, 10.56 , 11.28 , 12.59 , 14.73 , 17.93 , 22.335, 27.975, 34.76 , 42.46 , 50.75 , 59.25 , 67.54 , 75.24 , 82.025, 87.665, 92.07 , 95.27 , 97.415, 98.75 , 99.63 , 100.67 , 102.945, 107.945, 117.08 , 130.96 , 149.015, 169.885, 192.19 , 215.025, 238. , 261. , 284. , 307. , 330. , 353. , 376. , 399. , 422. , 445. , 228.25 ], dtype=float32) - drF(k)>f410.0 10.0 10.0 ... 433.5 456.5
- standard_name :
- cell_z_size
- long_name :
- cell z size
- units :
- m
array([ 10. , 10. , 10. , 10. , 10. , 10. , 10. , 10.01, 10.03, 10.11, 10.32, 10.8 , 11.76, 13.42, 16.04, 19.82, 24.85, 31.1 , 38.42, 46.5 , 55. , 63.5 , 71.58, 78.9 , 85.15, 90.18, 93.96, 96.58, 98.25, 99.25, 100.01, 101.33, 104.56, 111.33, 122.83, 139.09, 158.94, 180.83, 203.55, 226.5 , 249.5 , 272.5 , 295.5 , 318.5 , 341.5 , 364.5 , 387.5 , 410.5 , 433.5 , 456.5 ], dtype=float32) - PHrefC(k)>f449.05 147.1 ... 5.357e+04 5.794e+04
- standard_name :
- cell_reference_pressure
- long_name :
- Reference Hydrostatic Pressure
- units :
- m2 s-2
array([4.9049999e+01, 1.4714999e+02, 2.4525000e+02, 3.4335001e+02, 4.4145001e+02, 5.3954999e+02, 6.3765002e+02, 7.3579907e+02, 8.3409528e+02, 9.3288196e+02, 1.0330911e+03, 1.1366847e+03, 1.2473416e+03, 1.3708494e+03, 1.5153507e+03, 1.6912440e+03, 1.9103503e+03, 2.1847852e+03, 2.5257808e+03, 2.9423132e+03, 3.4401709e+03, 4.0214133e+03, 4.6839805e+03, 5.4220850e+03, 6.2267505e+03, 7.0867441e+03, 7.9899507e+03, 8.9245498e+03, 9.8801904e+03, 1.0848928e+04, 1.1826299e+04, 1.2813871e+04, 1.3823762e+04, 1.4882702e+04, 1.6031257e+04, 1.7315975e+04, 1.8777811e+04, 2.0444383e+04, 2.2329768e+04, 2.4439162e+04, 2.6773943e+04, 2.9334352e+04, 3.2120393e+04, 3.5132062e+04, 3.8369363e+04, 4.1832293e+04, 4.5520852e+04, 4.9435043e+04, 5.3574863e+04, 5.7940312e+04], dtype=float32) - PHrefF(k_p1)>f40.0 98.1 ... 5.57e+04 6.018e+04
- standard_name :
- cell_reference_pressure
- long_name :
- Reference Hydrostatic Pressure
- units :
- m2 s-2
array([ 0. , 98.1 , 196.2 , 294.3 , 392.4 , 490.5 , 588.6 , 686.7 , 784.8981, 883.2924, 982.4715, 1083.7107, 1189.6587, 1305.0243, 1436.6746, 1594.0269, 1788.461 , 2032.2396, 2337.3306, 2714.2307, 3170.3958, 3709.9458, 4332.881 , 5035.0806, 5809.09 , 6644.411 , 7529.0767, 8450.824 , 9398.274 , 10362.106 , 11335.749 , 12316.848 , 13310.895 , 14336.628 , 15428.775 , 16633.738 , 17998.21 , 19557.412 , 21331.355 , 23328.18 , 25550.145 , 27997.74 , 30670.965 , 33569.82 , 36694.305 , 40044.42 , 43620.164 , 47421.54 , 51448.547 , 55701.18 , 60179.445 ], dtype=float32) - hFacC(k, face, j, i)>f40.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0
- standard_name :
- cell_vertical_fraction
- long_name :
- vertical fraction of open cell
array([[[[0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], ..., [1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ]], [[1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ], ... [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ]], [[0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], ..., [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ]]]], dtype=float32) - hFacW(k, face, j, i_g)>f40.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0
- standard_name :
- cell_vertical_fraction_at_u_location
- long_name :
- vertical fraction of open cell
array([[[[0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], ..., [1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ]], [[1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ], ... [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ]], [[0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], ..., [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ]]]], dtype=float32) - hFacS(k, face, j_g, i)>f40.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0
- standard_name :
- cell_vertical_fraction_at_v_location
- long_name :
- vertical fraction of open cell
array([[[[0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], ..., [1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ]], [[1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ], ... [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ]], [[0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], ..., [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ]]]], dtype=float32) - maskC(k, face, j, i)boolFalse False False ... False False
- standard_name :
- sea_binary_mask_at_t_location
- long_name :
- mask denoting wet point at center
array([[[[False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], ..., [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True]], [[ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], ..., [ True, True, True, ..., False, True, True], [ True, True, True, ..., False, True, True], [ True, True, True, ..., False, False, True]], [[ True, True, True, ..., False, False, True], [ True, True, True, ..., True, False, True], [ True, True, True, ..., True, True, True], ..., ... ..., [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False]], [[False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], ..., [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False]], [[False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], ..., [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False]]]]) - maskW(k, face, j, i_g)boolFalse False False ... False False
- standard_name :
- cell_vertical_fraction_at_u_location
- long_name :
- mask denoting wet point at interface
array([[[[False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], ..., [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True]], [[ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], ..., [ True, True, True, ..., False, False, True], [ True, True, True, ..., False, False, True], [ True, True, True, ..., False, False, False]], [[ True, True, True, ..., False, False, False], [ True, True, True, ..., True, False, False], [ True, True, True, ..., True, True, True], ..., ... ..., [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False]], [[False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], ..., [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False]], [[False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], ..., [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False]]]]) - maskS(k, face, j_g, i)boolFalse False False ... False False
- standard_name :
- cell_vertical_fraction_at_v_location
- long_name :
- mask denoting wet point at interface
array([[[[False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], ..., [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True]], [[ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], ..., [ True, True, True, ..., False, True, True], [ True, True, True, ..., False, True, True], [ True, True, True, ..., False, False, True]], [[ True, True, True, ..., False, False, True], [ True, True, True, ..., False, False, True], [ True, True, True, ..., True, False, True], ..., ... ..., [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False]], [[False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], ..., [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False]], [[False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], ..., [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False]]]]) - maskCtrlC(k, face, j, i)boolFalse False False ... False False
- standard_name :
- ctrl_vector_3d_mask
- long_name :
- CTRL 3D mask where ctrl vector is active at tracer location
- units :
array([[[[False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], ..., [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True]], [[ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], ..., [ True, True, True, ..., False, True, True], [ True, True, True, ..., False, True, True], [ True, True, True, ..., False, False, True]], [[ True, True, True, ..., False, False, True], [ True, True, True, ..., True, False, True], [ True, True, True, ..., True, True, True], ..., ... ..., [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False]], [[False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], ..., [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False]], [[False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], ..., [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False]]]]) - maskCtrlS(k, face, j_g, i)boolFalse False False ... False False
- standard_name :
- ctrl_vector_3d_mask_at_v_location
- long_name :
- CTRL 3D mask where ctrl vector is active at v location
- units :
array([[[[False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], ..., [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True]], [[ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], ..., [ True, True, True, ..., False, True, True], [ True, True, True, ..., False, True, True], [ True, True, True, ..., False, False, True]], [[ True, True, True, ..., False, False, True], [ True, True, True, ..., False, False, True], [ True, True, True, ..., True, False, True], ..., ... ..., [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False]], [[False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], ..., [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False]], [[False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], ..., [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False]]]]) - rhoRef(k)>f41.024e+03 1.024e+03 ... 1.054e+03
- standard_name :
- reference_density_profile
- long_name :
- 1D, vertical reference density profile
- coordinate :
- Z
- units :
- kg m-3
array([1023.5776 , 1023.6208 , 1023.66394, 1023.991 , 1024.0343 , 1024.0776 , 1024.3971 , 1024.7086 , 1024.7523 , 1025.0563 , 1025.3527 , 1025.3989 , 1025.6919 , 1025.9822 , 1026.0472 , 1026.3529 , 1026.6698 , 1027.0033 , 1027.3589 , 1027.7407 , 1028.3246 , 1028.5933 , 1029.0645 , 1029.5631 , 1030.0851 , 1030.4861 , 1031.0481 , 1031.4844 , 1032.0652 , 1032.518 , 1032.9736 , 1033.5654 , 1034.0369 , 1034.53 , 1035.1935 , 1035.7921 , 1036.471 , 1037.2421 , 1038.2471 , 1039.2203 , 1040.2919 , 1041.4601 , 1042.7234 , 1044.0795 , 1045.5265 , 1047.0621 , 1048.6838 , 1050.3894 , 1052.1759 , 1054.0406 ], dtype=float32) - maskCtrlW(k, face, j, i_g)boolFalse False False ... False False
- standard_name :
- ctrl_vector_3d_mask_at_u_location
- long_name :
- CTRL 3D mask where ctrl vector is active at u location
- units :
array([[[[False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], ..., [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True]], [[ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], ..., [ True, True, True, ..., False, False, True], [ True, True, True, ..., False, False, True], [ True, True, True, ..., False, False, False]], [[ True, True, True, ..., False, False, False], [ True, True, True, ..., True, False, False], [ True, True, True, ..., True, True, True], ..., ... ..., [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False]], [[False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], ..., [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False]], [[False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], ..., [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False]]]])
- xx_kapredi(time, k, face, j, i)>f40.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0
array([[[[[ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], ..., [ 9.91381705e-03, 7.36427447e-03, 5.49139781e-03, ..., -6.09950488e-03, -5.78788714e-03, -5.75005310e-03], [ 1.40299965e-02, 9.83888283e-03, 7.29267346e-03, ..., -3.25040868e-03, -4.09374759e-03, -5.58908843e-03], [ 2.11328287e-02, 1.51925171e-02, 1.14498697e-02, ..., 1.41750311e-03, -4.90736391e-04, -3.33952089e-03]], [[ 2.89764702e-02, 2.15260740e-02, 1.64005980e-02, ..., 5.87992975e-03, 3.27166729e-03, -3.28127178e-04], [ 3.61302309e-02, 2.75439024e-02, 2.11683363e-02, ..., 8.92235618e-03, 6.08210359e-03, 2.42907042e-03], [ 4.21850421e-02, 3.27062681e-02, 2.54961718e-02, ..., 1.04417633e-02, 7.82990921e-03, 4.67536738e-03], ... 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00]], [[ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], ..., [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00], [ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00, ..., 0.00000000e+00, 0.00000000e+00, 0.00000000e+00]]]]], dtype=float32)
- iPandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89], dtype='int64', name='i')) - i_gPandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89], dtype='int64', name='i_g')) - jPandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89], dtype='int64', name='j')) - j_gPandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89], dtype='int64', name='j_g')) - kPandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49], dtype='int64', name='k')) - k_uPandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49], dtype='int64', name='k_u')) - k_lPandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49], dtype='int64', name='k_l')) - k_p1PandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50], dtype='int64', name='k_p1')) - facePandasIndex
PandasIndex(Int64Index([0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12], dtype='int64', name='face'))
- timePandasIndex
PandasIndex(TimedeltaIndex(['0 days 00:02:09'], dtype='timedelta64[ns]', name='time', freq=None))
- Conventions :
- CF-1.6
- title :
- netCDF wrapper of MITgcm MDS binary data
- source :
- MITgcm
- history :
- Created by calling `open_mdsdataset(grid_dir='/glade/u/home/dcherian/work/mitgcm/ECCOV4/release4/input_init/../run/', iters='all', prefix='xx_kapredi', read_grid=True, delta_t=1, ref_date=None, calendar='gregorian', geometry='llc', grid_vars_to_coords=True, swap_dims=None, endian='>', chunks=None, ignore_unknown_vars=False, default_dtype=None, nx=None, ny=None, nz=None, llc_method='smallchunks', extra_metadata=None, extra_variables={'xx_kapredi': {'dims': ['k', 'j', 'i'], 'attrs': {}}}, data_dir='/glade/u/home/dcherian/work/mitgcm/ECCOV4/release4/input_init/', levels=None)`
natre_mean.GM_Kux.load()
<xarray.DataArray 'GM_Kux' (node: 2, k: 50)>
array([[1653.3713 , 1756.2273 , 1774.7654 , 1698.0707 , 1606.0331 ,
1502.6423 , 1411.8212 , 1353.8236 , 1326.3242 , 1335.0173 ,
1382.8555 , 1460.9812 , 1550.7936 , 1637.4607 , 1704.886 ,
1736.8871 , 1719.9307 , 1662.3121 , 1585.8231 , 1500.4946 ,
1408.286 , 1299.3516 , 1193.2825 , 1113.6604 , 1086.0511 ,
1114.4277 , 1220.7032 , 1396.9984 , 1558.5225 , 1537.3969 ,
1229.34 , 750.17395, 366.39853, 213.548 , 228.79117,
287.18646, 343.04477, 408.48822, 487.57626, 579.28723,
710.1041 , 898.3161 , 1090.6859 , 1212.5089 , 1229.1472 ,
1170.0338 , 1088.2152 , 1028.699 , 999.34344, 1000. ],
[1122.2953 , 1138.5017 , 1138.5872 , 1127.2064 , 1121.2599 ,
1119.6086 , 1118.9813 , 1114.1803 , 1100.417 , 1080.9762 ,
1060.82 , 1044.8038 , 1036.1597 , 1035.7036 , 1040.701 ,
1040.6842 , 1025.8724 , 993.8615 , 949.4629 , 908.0531 ,
887.2899 , 903.4036 , 956.21436, 1028.5127 , 1086.1965 ,
1094.7805 , 1055.2051 , 999.22345, 958.0874 , 941.8081 ,
944.84357, 962.67786, 988.8215 , 1001.7471 , 980.7318 ,
939.7799 , 920.5631 , 944.925 , 995.82355, 1041.636 ,
1061.2731 , 1056.8977 , 1041.72 , 1026.6104 , 1016.8036 ,
1011.05994, 1007.35706, 1004.25977, 1000.74286, 1000. ]],
dtype=float32)
Coordinates:
* k (k) int64 0 1 2 3 4 5 6 7 8 9 10 ... 40 41 42 43 44 45 46 47 48 49
face int64 2
time timedelta64[ns] 00:23:48
Z (k) float32 -5.0 -15.0 -25.0 ... -5.039e+03 -5.461e+03 -5.906e+03
drF (k) float32 10.0 10.0 10.0 10.0 10.0 ... 387.5 410.5 433.5 456.5
PHrefC (k) float32 49.05 147.1 245.2 ... 4.944e+04 5.357e+04 5.794e+04
rhoRef (k) float32 1.024e+03 1.024e+03 1.024e+03 ... 1.052e+03 1.054e+03
iter float64 1.428e+03
* node (node) object 'adj' 'const'xarray.DataArray
'GM_Kux'
- node: 2
- k: 50
- 1.653e+03 1.756e+03 1.775e+03 1.698e+03 ... 1.004e+03 1.001e+03 1e+03
array([[1653.3713 , 1756.2273 , 1774.7654 , 1698.0707 , 1606.0331 , 1502.6423 , 1411.8212 , 1353.8236 , 1326.3242 , 1335.0173 , 1382.8555 , 1460.9812 , 1550.7936 , 1637.4607 , 1704.886 , 1736.8871 , 1719.9307 , 1662.3121 , 1585.8231 , 1500.4946 , 1408.286 , 1299.3516 , 1193.2825 , 1113.6604 , 1086.0511 , 1114.4277 , 1220.7032 , 1396.9984 , 1558.5225 , 1537.3969 , 1229.34 , 750.17395, 366.39853, 213.548 , 228.79117, 287.18646, 343.04477, 408.48822, 487.57626, 579.28723, 710.1041 , 898.3161 , 1090.6859 , 1212.5089 , 1229.1472 , 1170.0338 , 1088.2152 , 1028.699 , 999.34344, 1000. ], [1122.2953 , 1138.5017 , 1138.5872 , 1127.2064 , 1121.2599 , 1119.6086 , 1118.9813 , 1114.1803 , 1100.417 , 1080.9762 , 1060.82 , 1044.8038 , 1036.1597 , 1035.7036 , 1040.701 , 1040.6842 , 1025.8724 , 993.8615 , 949.4629 , 908.0531 , 887.2899 , 903.4036 , 956.21436, 1028.5127 , 1086.1965 , 1094.7805 , 1055.2051 , 999.22345, 958.0874 , 941.8081 , 944.84357, 962.67786, 988.8215 , 1001.7471 , 980.7318 , 939.7799 , 920.5631 , 944.925 , 995.82355, 1041.636 , 1061.2731 , 1056.8977 , 1041.72 , 1026.6104 , 1016.8036 , 1011.05994, 1007.35706, 1004.25977, 1000.74286, 1000. ]], dtype=float32) - k(k)int640 1 2 3 4 5 6 ... 44 45 46 47 48 49
- standard_name :
- z_grid_index
- axis :
- Z
- long_name :
- z-dimension of the t grid
- swap_dim :
- Z
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49]) - face()int642
- standard_name :
- face_index
array(2)
- time()timedelta64[ns]00:23:48
- standard_name :
- time
- long_name :
- Time
- axis :
- T
- calendar :
- gregorian
array(1428000000000, dtype='timedelta64[ns]')
- Z(k)float32-5.0 -15.0 ... -5.906e+03
- standard_name :
- depth
- long_name :
- vertical coordinate of cell center
- units :
- m
- positive :
- down
array([-5.000000e+00, -1.500000e+01, -2.500000e+01, -3.500000e+01, -4.500000e+01, -5.500000e+01, -6.500000e+01, -7.500500e+01, -8.502500e+01, -9.509500e+01, -1.053100e+02, -1.158700e+02, -1.271500e+02, -1.397400e+02, -1.544700e+02, -1.724000e+02, -1.947350e+02, -2.227100e+02, -2.574700e+02, -2.999300e+02, -3.506800e+02, -4.099300e+02, -4.774700e+02, -5.527100e+02, -6.347350e+02, -7.224000e+02, -8.144700e+02, -9.097400e+02, -1.007155e+03, -1.105905e+03, -1.205535e+03, -1.306205e+03, -1.409150e+03, -1.517095e+03, -1.634175e+03, -1.765135e+03, -1.914150e+03, -2.084035e+03, -2.276225e+03, -2.491250e+03, -2.729250e+03, -2.990250e+03, -3.274250e+03, -3.581250e+03, -3.911250e+03, -4.264250e+03, -4.640250e+03, -5.039250e+03, -5.461250e+03, -5.906250e+03], dtype=float32) - drF(k)float3210.0 10.0 10.0 ... 433.5 456.5
- standard_name :
- cell_z_size
- long_name :
- cell z size
- units :
- m
array([ 10. , 10. , 10. , 10. , 10. , 10. , 10. , 10.01, 10.03, 10.11, 10.32, 10.8 , 11.76, 13.42, 16.04, 19.82, 24.85, 31.1 , 38.42, 46.5 , 55. , 63.5 , 71.58, 78.9 , 85.15, 90.18, 93.96, 96.58, 98.25, 99.25, 100.01, 101.33, 104.56, 111.33, 122.83, 139.09, 158.94, 180.83, 203.55, 226.5 , 249.5 , 272.5 , 295.5 , 318.5 , 341.5 , 364.5 , 387.5 , 410.5 , 433.5 , 456.5 ], dtype=float32) - PHrefC(k)float3249.05 147.1 ... 5.357e+04 5.794e+04
- standard_name :
- cell_reference_pressure
- long_name :
- Reference Hydrostatic Pressure
- units :
- m2 s-2
array([4.9049999e+01, 1.4714999e+02, 2.4525000e+02, 3.4335001e+02, 4.4145001e+02, 5.3954999e+02, 6.3765002e+02, 7.3579907e+02, 8.3409528e+02, 9.3288196e+02, 1.0330911e+03, 1.1366847e+03, 1.2473416e+03, 1.3708494e+03, 1.5153507e+03, 1.6912440e+03, 1.9103503e+03, 2.1847852e+03, 2.5257808e+03, 2.9423132e+03, 3.4401709e+03, 4.0214133e+03, 4.6839805e+03, 5.4220850e+03, 6.2267505e+03, 7.0867441e+03, 7.9899507e+03, 8.9245498e+03, 9.8801904e+03, 1.0848928e+04, 1.1826299e+04, 1.2813871e+04, 1.3823762e+04, 1.4882702e+04, 1.6031257e+04, 1.7315975e+04, 1.8777811e+04, 2.0444383e+04, 2.2329768e+04, 2.4439162e+04, 2.6773943e+04, 2.9334352e+04, 3.2120393e+04, 3.5132062e+04, 3.8369363e+04, 4.1832293e+04, 4.5520852e+04, 4.9435043e+04, 5.3574863e+04, 5.7940312e+04], dtype=float32) - rhoRef(k)float321.024e+03 1.024e+03 ... 1.054e+03
- standard_name :
- reference_density_profile
- long_name :
- 1D, vertical reference density profile
- coordinate :
- Z
- units :
- kg m-3
array([1023.5776 , 1023.6208 , 1023.66394, 1023.991 , 1024.0343 , 1024.0776 , 1024.3971 , 1024.7086 , 1024.7523 , 1025.0563 , 1025.3527 , 1025.3989 , 1025.6919 , 1025.9822 , 1026.0472 , 1026.3529 , 1026.6698 , 1027.0033 , 1027.3589 , 1027.7407 , 1028.3246 , 1028.5933 , 1029.0645 , 1029.5631 , 1030.0851 , 1030.4861 , 1031.0481 , 1031.4844 , 1032.0652 , 1032.518 , 1032.9736 , 1033.5654 , 1034.0369 , 1034.53 , 1035.1935 , 1035.7921 , 1036.471 , 1037.2421 , 1038.2471 , 1039.2203 , 1040.2919 , 1041.4601 , 1042.7234 , 1044.0795 , 1045.5265 , 1047.0621 , 1048.6838 , 1050.3894 , 1052.1759 , 1054.0406 ], dtype=float32) - iter()float641.428e+03
- standard_name :
- timestep
- long_name :
- model timestep number
array(1428.)
- node(node)object'adj' 'const'
array(['adj', 'const'], dtype=object)
- kPandasIndex
PandasIndex(Int64Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49], dtype='int64', name='k')) - nodePandasIndex
PandasIndex(Index(['adj', 'const'], dtype='object', name='node'))
(kapredi_adj).sel(face=2).isel(time=0, i=slice(10, 18), j=slice(10, 20)).mean(
["i", "j"]
).xx_kapredi.plot(color="k")
(natre_mean.GM_Kux / 2000 - 0.5).sel(node="const").plot.line(hue="node")
[<matplotlib.lines.Line2D at 0x2ba4c4f1bd60>]