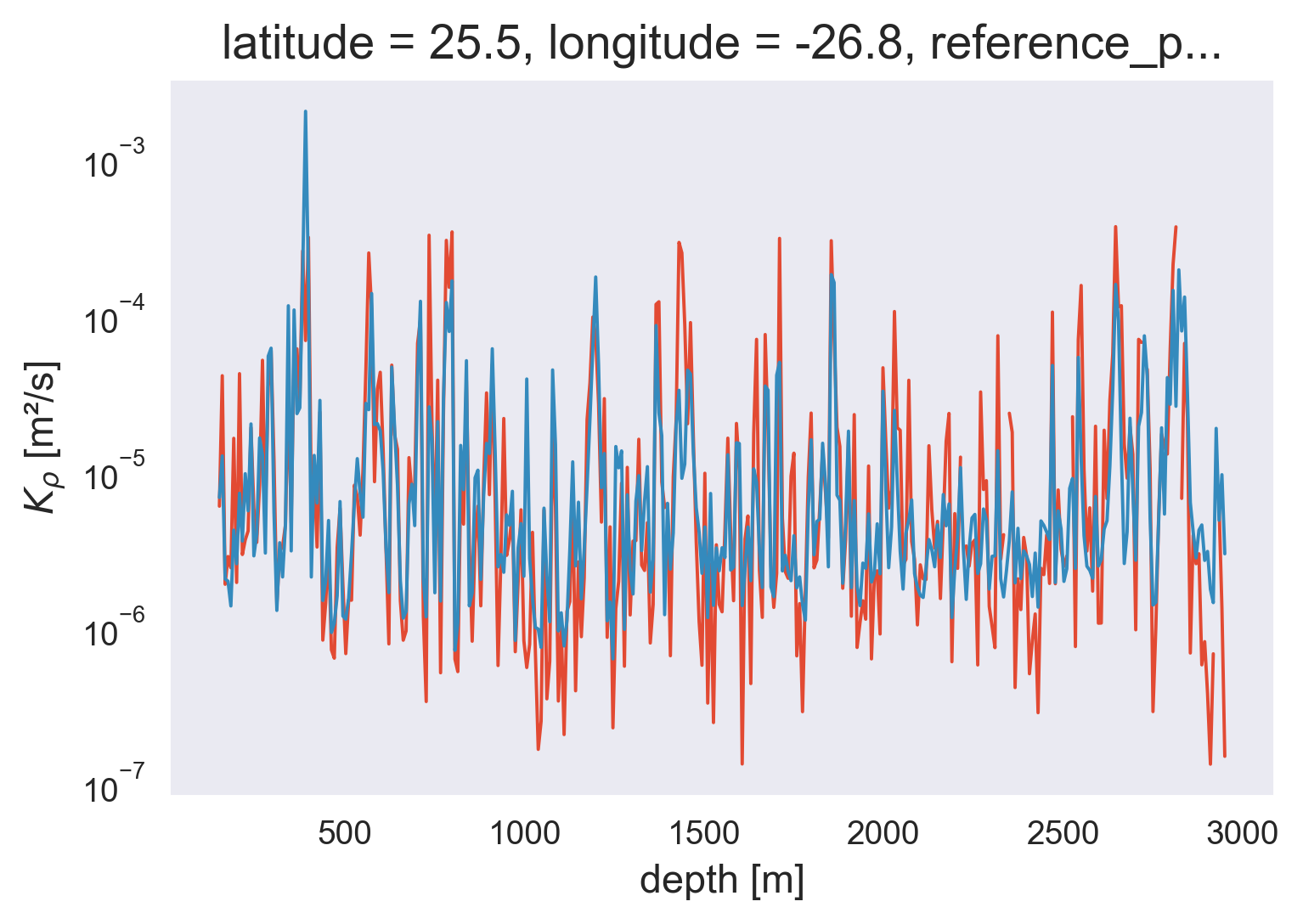
NATRE micro vs finescale#
Here I compare the NATRE microstructure estimates with my Argo finestructure estimates. Key questions:
How well do the two agree?
they don’t! The argo average χ seems to miss any eddy contribution; it agrees really well with the “mean stirring” estimate of Ferrari & Polzin.
If I average the microstructure to 200m bins can I recover the decomposition or not?
TODO; skipped for now since it looks like the Argo estimate misses eddies
%load_ext watermark
%matplotlib inline
import glob
import cf_xarray as cfxr
import dask
import dcpy
import distributed
import matplotlib as mpl
import matplotlib.pyplot as plt
import numpy as np
import tqdm
from IPython.display import Image
import eddydiff as ed
import xarray as xr
xr.set_options(keep_attrs=True)
plt.rcParams["figure.dpi"] = 140
plt.rcParams["savefig.dpi"] = 200
plt.style.use("ggplot")
%watermark -iv
eddydiff : 0.1
cf_xarray : 0.5.3.dev29+g3660810.d20210729
numpy : 1.21.1
dask : 2021.7.2
tqdm : 4.62.0
dcpy : 0.1
xarray : 0.17.1.dev3+g48378c4b1
matplotlib : 3.4.2
distributed: 2021.7.2
Get datasets#
There are 3
NATRE microstructure
NATRE finescale
Argo finescale
Show code cell source
natre = ed.natre.read_natre().load()
bins = ed.sections.choose_bins(natre.gamma_n, depth_range=np.arange(150, 2001, 100))
natre_micro_dens = ed.sections.bin_average_vertical(
natre.reset_coords("pres"), "neutral_density", bins
).rename({"chi": "χ", "eps": "ε", "Krho": "Kρ"})
Show code cell source
natre_micro_depth = (
natre.groupby_bins("pres", bins=np.arange(250, 2050, 200))
.mean(...)
.rename({"chi": "χ", "eps": "ε", "Krho": "Kρ"})
)
natre_micro_depth["pres_bins"].attrs = {
"standard_name": "sea_water_pressure",
"axis": "Z",
"positive": "down",
}
Show code cell source
natre_fine = xr.load_dataset("../datasets/natre-finescale.nc")
natre_fine_depth = (
natre_fine.groupby_bins("pressure", bins=np.arange(250, 2001, 200))
.mean()
.cf.guess_coord_axis()
)
natre_fine_depth.pressure_bins.attrs = {
"positive": "down",
"standard_name": "sea_water_pressure",
}
Show code cell source
argo_fine_profiles = xr.load_dataset(
"../datasets/argomix_natre.zarr", engine="zarr"
).drop("DIRECTION")
argo_fine_profiles["Kt"] = argo_fine_profiles["Kρ"]
argo_fine_profiles["Kt"].attrs["long_name"] = "$K_T$"
# argo_fine_profiles = argo_fine_profiles.where(argo_fine_profiles.ε < 0.1)
bins = cfxr.bounds_to_vertices(natre_micro_dens.gamma_n_bounds, "bounds").data
argo_fine_dens = (
argo_fine_profiles.rename({"γmean": "gamma_n"})
.reset_coords("pressure")
.groupby_bins("gamma_n", bins=bins)
.mean()
)
argo_fine_dens["gamma_n_bins"].attrs = {
"standard_name": "neutral_density",
"axis": "Z",
"positive": "down",
}
argo_fine_dens["npts"] = (
argo_fine_profiles.χ.rename({"γmean": "gamma_n"})
.groupby_bins("gamma_n", bins=bins)
.count()
)
argo_fine_dens = argo_fine_dens.set_coords("pressure")
argo_fine_depth = argo_fine_profiles.groupby_bins(
"pressure", np.arange(250, 2050, 200)
).mean()
argo_fine_depth["pressure_bins"].attrs = {"standard_name": "sea_water_pressure"}
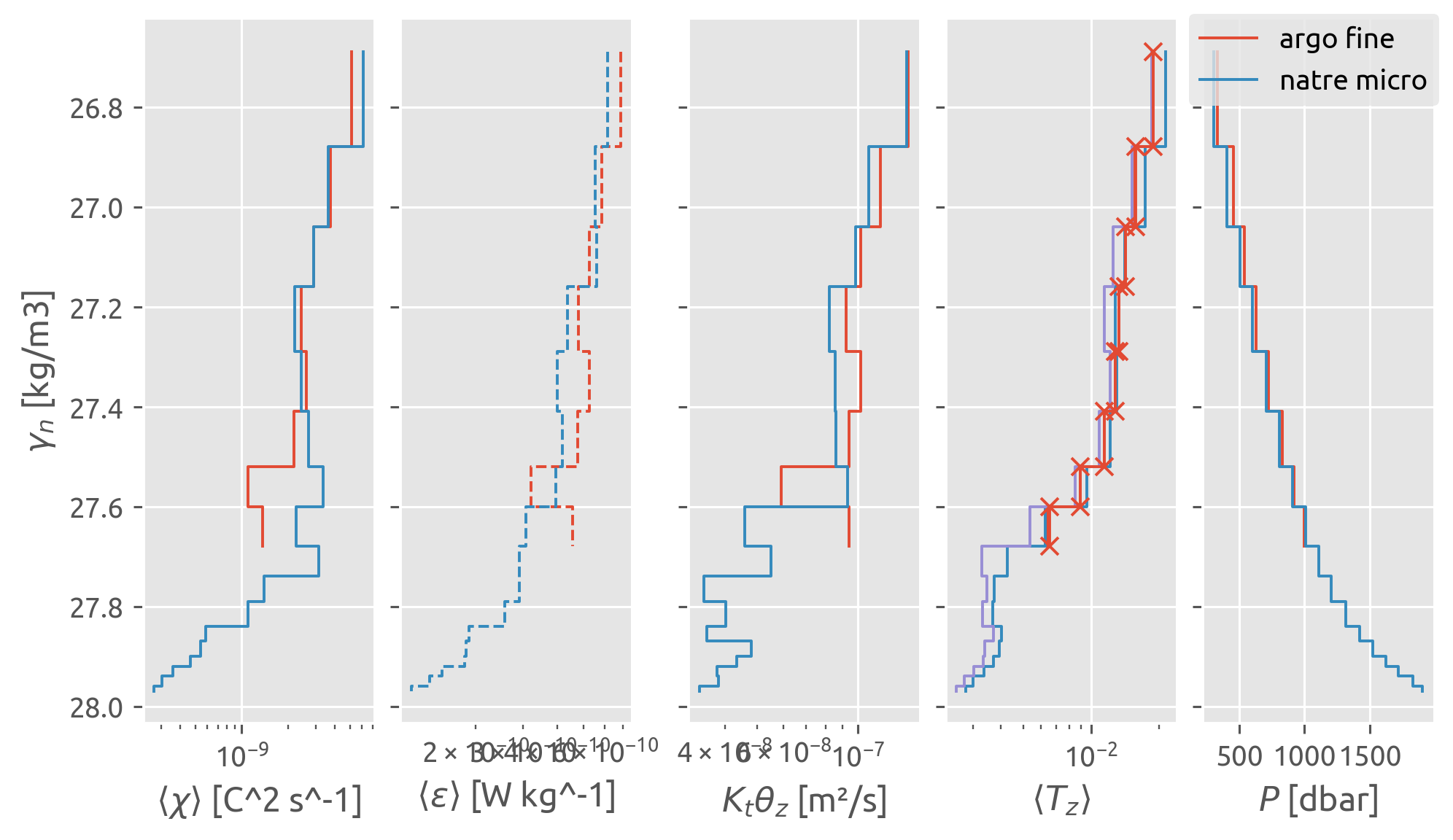
Finescale vs microscale#
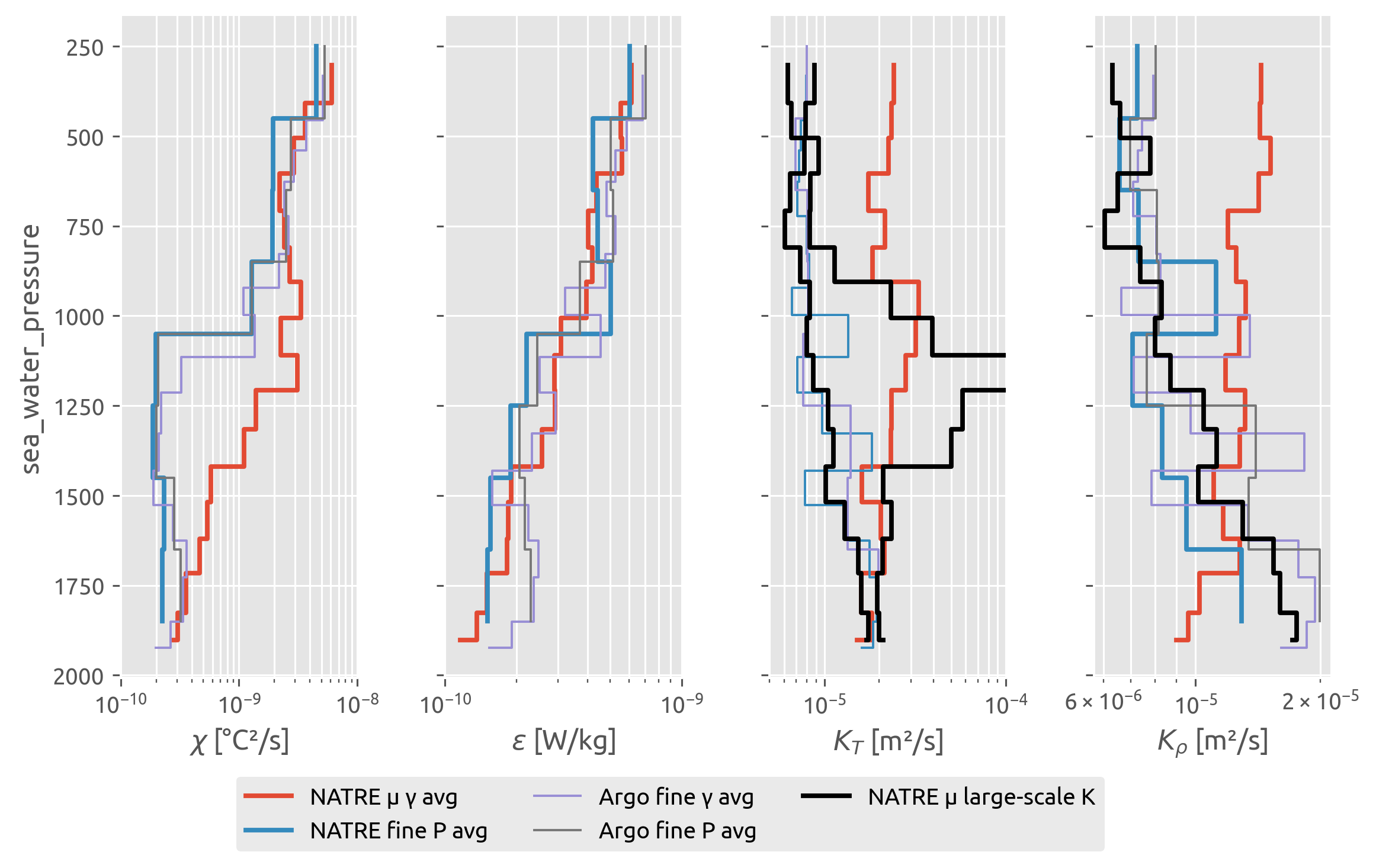
The finescale \(⟨ε^{200}⟩\) estimates agree well with microscale \(⟨ε^{0.5}⟩\)
NATRE finescale \(⟨K_ρ^{200}⟩\) agrees with Argo finescale \(⟨K_ρ^{200}⟩\) which agrees with \(K_ρ^m\) \(= Γ⟨ε^{0.5}⟩/N_m^2\) i.e. a diapycnal diffusivity for the stirring of the large scale vertical gradient by the microscale. Somehow calculating a 200-m diffusivity using the finestructure \(ε^{200}\) and 200-m scale \(N²\) seems to average us out to the “large scale”
Argo finescale \(⟨χ^{200}⟩ = ⟨Γε^{200}/N\_{200}^2 ; ∂_zT\_{200}^2⟩\) seems to miss the whole eddy contribution to variance production! This is robust to averaging in γ or P, so it is not a question of the bins being too fine for the 200m mean χ estimate in each profile. I think the issue is scaling the 200-m \(ε^{200}\) with 200-m scale gradients to get \(χ^{200}\)
NATRE microscale \(⟨K_ρ⟩\) and \(⟨K_T⟩\) is quite different near the top.
The large-scale mean \(K_T^m\) is the (meaningless) diffusivity required to explain \(⟨χ⟩\) by microscale stirring on the mean vertical gradient. that’s why there’s a peak where variance is actually being generated by mesoscale stirring of the lateral gradient.
TODO:
The disagreement between microscale \(⟨K_ρ⟩\) and the \(K_ρ^m\) or finescale \(K_ρ\) is the key I think. I think the 200-m averaging inherent in the finescale estimate is filtering out the mesoscale. BUT then the 200m averaged microscale \(⟨K_ρ⟩\) should also reflect this? But maybe I should calculate the 200m scale diffusivity instead of averaging the 0.5m scale diffusivity in 200m bins.
Is there any way I can estimate the microscale \(K_ρ\) that gets me back to the “large-scale” \(K_ρ^m\)
Look at individual profiles of \(χ, ε, χ^{200}, ε^{200}\); is the 200-m average taking out the eddy stirring contribution to \(T_z\)? Since they are compensated it makes sense that \(N² ≪ gαT_z^2\) at scales smaller than horizontal stirring vertical scale
Show code cell source
f, axx = plt.subplots(1, 4, sharey=True)
ax = dict(zip(["χ", "ε", "Kt", "Kρ"], axx))
hdl = []
for var, axis in ax.items():
for ds, lw in zip(
[
natre_micro_dens,
natre_fine_depth.sel(criteria="kunze"),
# natre_micro_depth,
argo_fine_dens,
argo_fine_depth,
],
[2, 2, 1, 1],
):
if var in ds:
h = ds[var].cf.plot.step(
y="sea_water_pressure", xscale="log", ax=axis, lw=lw
)
if var == "Kρ":
hdl.append(h[0])
natre_micro_dens["Kt_m"].cf.plot.step(
y="sea_water_pressure", xscale="log", ax=ax["Kt"], lw=2, color="k", _labels=False
)
h = natre_micro_dens["Krho_m"].cf.plot.step(
y="sea_water_pressure", xscale="log", ax=ax["Kt"], lw=2, color="k", _labels=False
)
h = natre_micro_dens["Krho_m"].cf.plot.step(
y="sea_water_pressure", xscale="log", ax=ax["Kρ"], lw=2, color="k", _labels=False
)
hdl.append(h[0])
f.legend(
hdl,
[
"NATRE μ γ avg",
# "NATRE μ P avg",
"NATRE fine P avg",
"Argo fine γ avg",
"Argo fine P avg",
"NATRE μ large-scale K",
],
ncol=3,
loc="center",
bbox_to_anchor=(0.5, 0),
)
ax["χ"].set_xlim([1e-10, 1e-8])
ax["ε"].set_xlim([1e-10, 1e-9])
ax["Kt"].set_xlim([5e-6, 1e-4])
[aa.grid(True, which="minor", axis="x") for aa in axx]
ax["Kt"].set_title("")
dcpy.plots.clean_axes(axx)
plt.tight_layout(h_pad=0.1, w_pad=0.1)
plt.gcf().set_size_inches((9, 5))
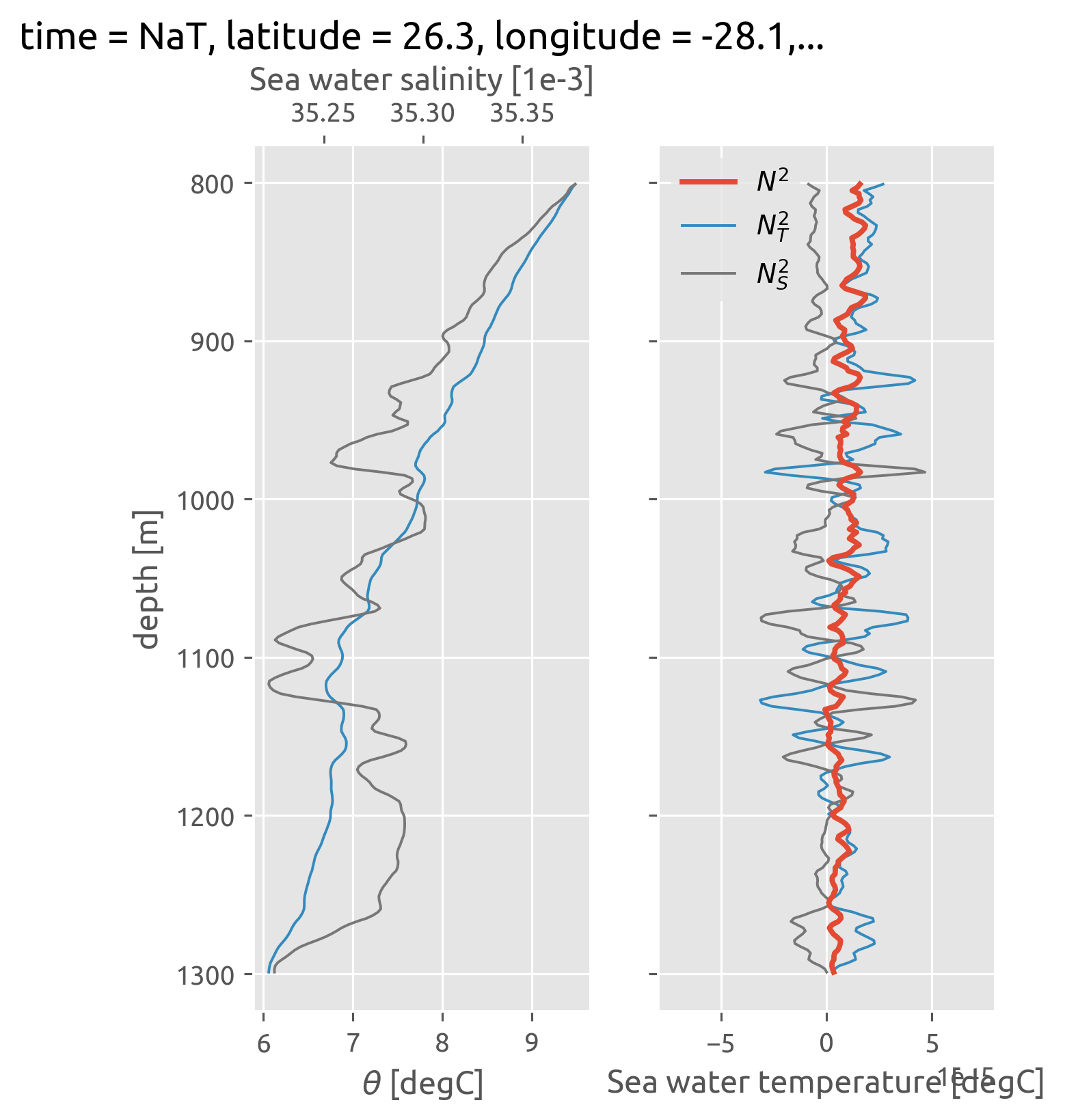
Studying N² vs \(T_z, S_z\)#
Well this is pretty clear!
There are compensated vertical gradients of \(T_z\) and \(S_z\) of ≈ 10-30m scale. So a 200-m diffusivity is acting on the much larger scale gradient \(∂_z θ_m\) not \(∂_z(θ_m + θ_e)\), the latter being the mean for the microscale diffusivity.
Another interesting thing is that \(⟨K_ρ^μ⟩ = K_ρ^m\) i.e. “large-scale” average \(K_ρ\) is the diffusivity for the “large-scale” because there is no stirring of density!
The other way to think of it is that \(K_T ≠ K_ρ\) at scales of spiciness i.e. compensated T-S variability.
Show code cell source
natre["NT2"] = 9.81 * dcpy.eos.alpha(natre.salt, natre.temp, natre.pres) * natre.Tz
natre["NS2"] = (
9.81
* dcpy.eos.beta(natre.salt, natre.temp, natre.pres)
* natre.salt.interpolate_na("depth").differentiate("depth")
)
profile = natre.isel(latitude=3, longitude=6).temp.interpolate_na("depth")
dcpy.ts.PlotSpectrum(profile)
dcpy.ts.PlotSpectrum(profile.rolling(depth=7, center=True).mean(), ax=plt.gca())
dcpy.ts.PlotSpectrum(profile.rolling(depth=31, center=True).mean(), ax=plt.gca())
0.4000000000000341
0.4000000000000341
0.4000000000000341
([<matplotlib.lines.Line2D at 0x7fc910df1910>],
<AxesSubplot:title={'center':' Sea water temperature\n[degrees_Celsius]'}, xlabel='Wavelength/2π', ylabel='PSD'>)
%matplotlib inline
plt.rcParams["figure.dpi"] = 140
profile = (
natre.isel(latitude=3, longitude=6)
.rolling(depth=21, center=True, min_periods=1)
.mean()
.coarsen(depth=5, boundary="trim")
.mean()
.sel(depth=slice(800, 1300))
)
profile["NT2"] = (
-9.81
* dcpy.eos.alpha(profile.salt, profile.temp, profile.pres)
* profile.theta.interpolate_na("depth").differentiate("depth")
)
profile["NS2"] = (
9.81
* dcpy.eos.beta(profile.salt, profile.temp, profile.pres)
* profile.salt.interpolate_na("depth").differentiate("depth")
)
f, ax = plt.subplots(1, 2, sharey=True)
profile.theta.cf.plot(ax=ax[0], color="C1")
axS = ax[0].twiny()
axS.grid(False)
profile.salt.cf.plot(color="C3", ax=axS)
# ax2 = ax[0].twiny()
# profile.gamma_n.cf.plot(color="k", ax=ax2)
# ax2.grid(False)
profile.N2.cf.plot(lw=2, zorder=5, ax=ax[1])
profile.NT2.cf.plot(ax=ax[1])
profile.NS2.cf.plot(color="C3", ax=ax[1])
ax[1].set_xlim([-8e-5, 8e-5])
ax[1].legend(["$N^2$", "$N_T^2$", "$N_S^2$"])
dcpy.plots.clean_axes(ax)
[aa.set_title("") for aa in ax]
f.set_size_inches((5, 6))
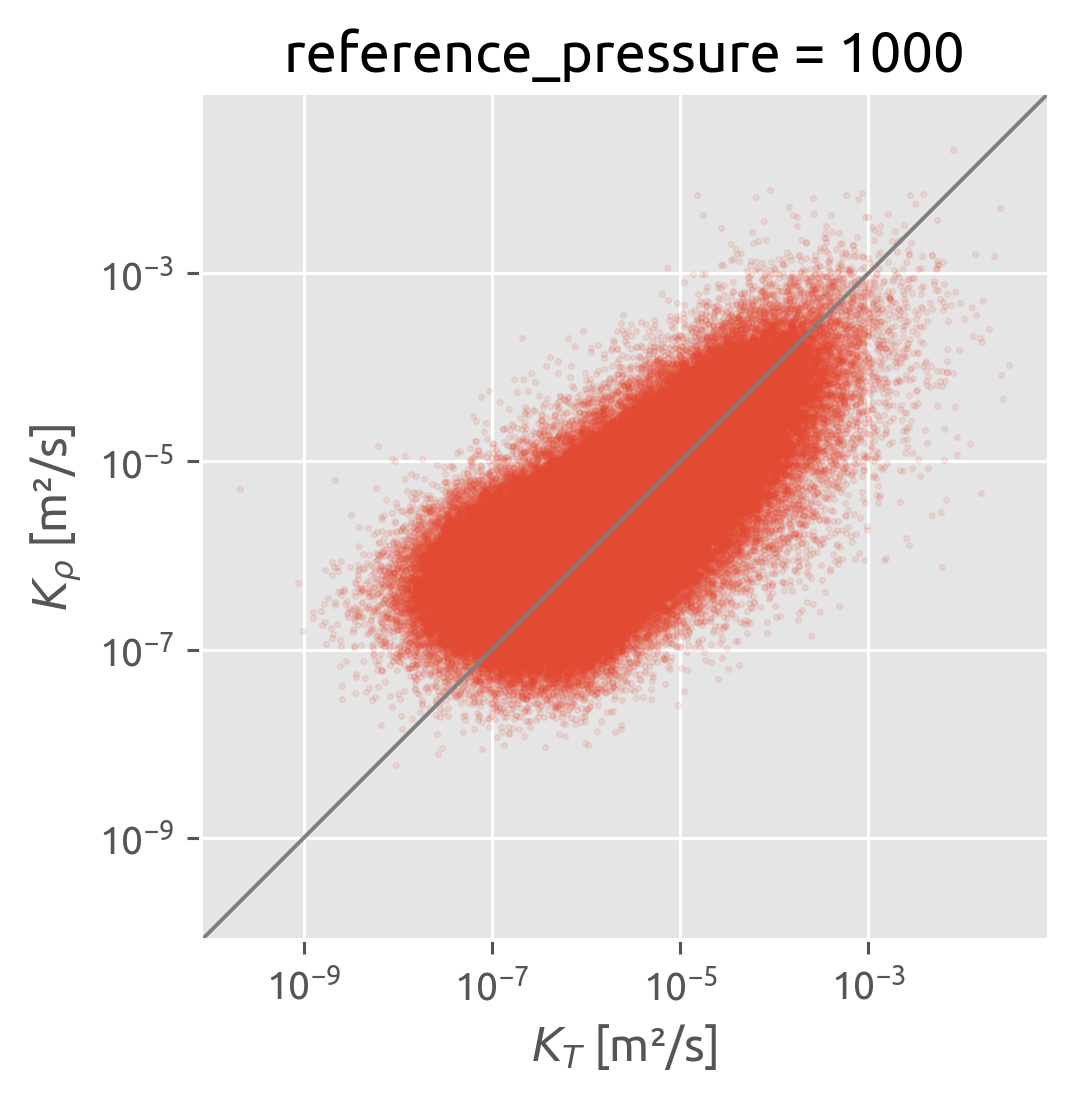
Studying \(K_T\) vs \(K_ρ\)#
I’m wondering if the \(K_T = K_ρ\) assumption really breaks down at the 200-m scale?
calculate \(K_T, K_ρ\) from the microstructure data at various averaging scales.
natre.plot.scatter(x="Kt", y="Krho", s=2, alpha=0.1)
plt.gca().set_xscale("log")
plt.gca().set_yscale("log")
dcpy.plots.line45()
Old stuff#
def decode_bounds_to_intervals(dataset, bounds_dim="bounds"):
dataset = dataset.copy(deep=True)
all_bounds = dataset.cf.bounds
for name in set(dataset.variables) & set(all_bounds):
bounds_name = all_bounds[name][0]
vertices = cfxr.bounds_to_vertices(dataset[bounds_name], bounds_dim)
intervals = ed.intervals_from_vertex(vertices)
dataset[name] = dataset[name].copy(data=intervals)
del dataset[name].attrs["bounds"]
dataset = dataset.drop(bounds_name)
return dataset
micro = decode_bounds_to_intervals(micro)
argo_fine.npts.plot()
[<matplotlib.lines.Line2D at 0x7ff00197a6a0>]
f, ax = plt.subplots(1, 5, sharey=True, constrained_layout=True)
kwargs = dict(y="neutral_density", xscale="log")
argo_fine.gamma_n_bins.attrs["standard_name"] = "neutral_density"
masked_argo_fine = argo_fine.where(argo_fine.npts > 300)
for var in argo_fine.variables:
masked_argo_fine[var].attrs = argo_fine[var].attrs
masked_argo_fine.χ.cf.plot.step(ax=ax[0], **kwargs)
micro.chi.cf.plot.step(ax=ax[0], **kwargs)
masked_argo_fine.ε.cf.plot.step(ax=ax[1], color="C0", ls="--", **kwargs)
micro.eps.cf.plot.step(ax=ax[1], color="C1", ls="--", **kwargs)
masked_argo_fine.KtTz.cf.plot.step(ax=ax[2], **kwargs)
micro.KtTz.cf.plot.step(ax=ax[2], **kwargs)
# micro.KrhoTz.cf.plot.step(ax=ax[2], **kwargs)
masked_argo_fine.Tzmean.cf.plot.step(ax=ax[3], **kwargs)
micro.Tz.cf.plot.step(ax=ax[3], **kwargs)
micro.dTdz_m.cf.plot.step(ax=ax[3], **kwargs)
masked_argo_fine.mean_dTdz_seg.cf.plot.step(ax=ax[3], color="C0", marker="x", **kwargs)
masked_argo_fine.pressure.cf.plot.step(ax=ax[4], y="neutral_density")
micro.pres.cf.plot.step(ax=ax[4], y="neutral_density")
ax[-1].set_xlabel("$P$ [dbar]")
f.legend(["argo fine", "natre micro"])
dcpy.plots.clean_axes(ax)
[aa.set_title("") for aa in ax]
f.set_size_inches((7, 4))
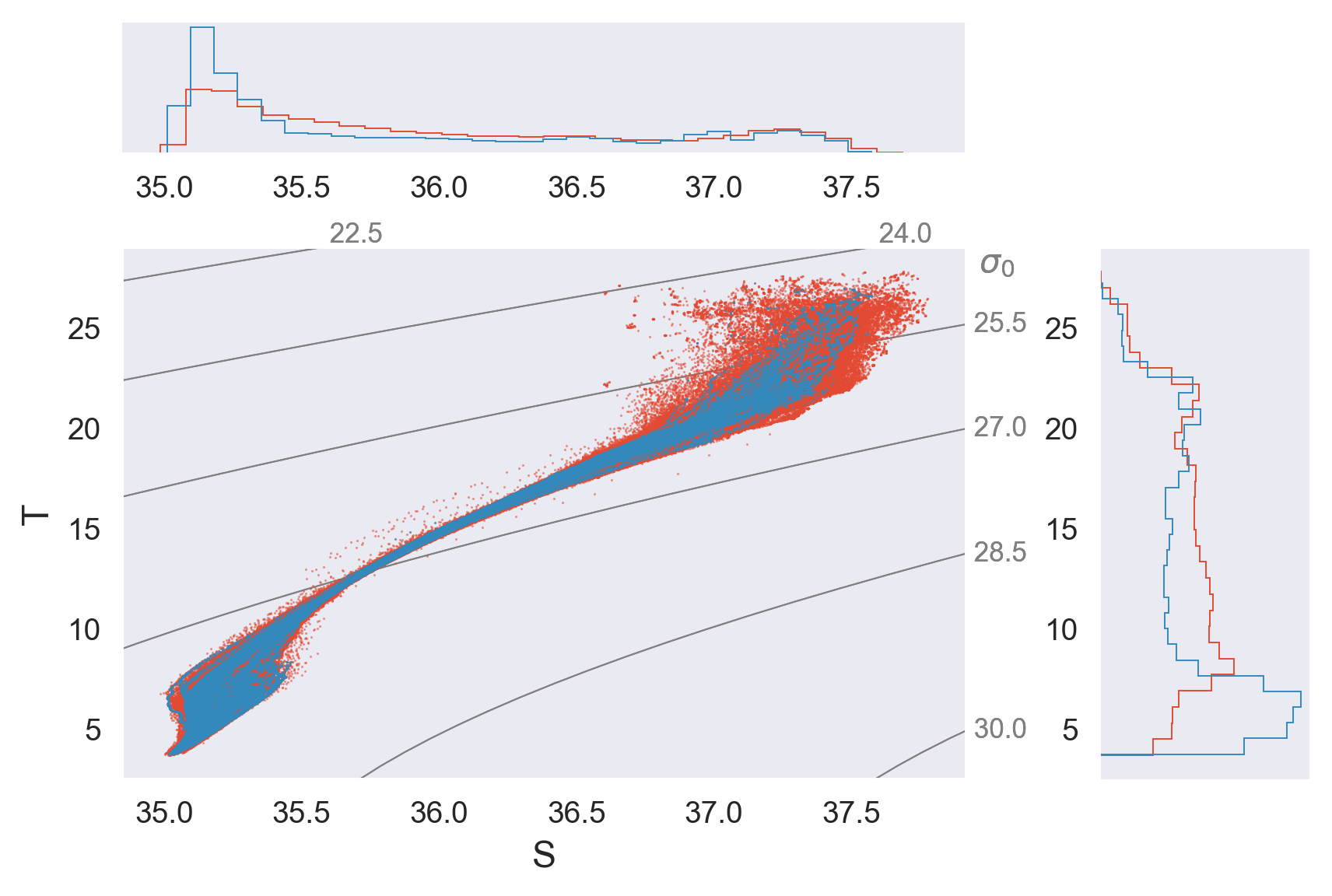
T-S#
Hmm.. the T-S diagram for Argo profiles with which we can calculate finescale estimates should tell us if it samples any eddy stirring at all.
argo_fine_full_profiles = xr.load_dataset("argo_profiles_used_for_finestructure.nc")
mask = argo_fine_full_profiles.fine_deepest_bin > 1200
kwargs = dict(hexbin=False, size=0.1)
_, ax = dcpy.oceans.TSplot(
argo_fine_full_profiles.PSAL,
argo_fine_full_profiles.THETA,
**kwargs,
)
dcpy.oceans.TSplot(
argo_fine_full_profiles.where(mask, drop=True).PSAL,
argo_fine_full_profiles.where(mask, drop=True).THETA,
**kwargs,
ax=ax,
);
Extract Argo profiles used for finescale estimates#
import argopy
argo_profiles = argo_natre.argo.point2profile()
argo_profiles
<xarray.Dataset>
Dimensions: (N_PROF: 4386, N_LEVELS: 1012)
Coordinates:
* N_PROF (N_PROF) int64 26 46 2 27 47 ... 3983 471 3677 3975
* N_LEVELS (N_LEVELS) int64 0 1 2 3 4 ... 1008 1009 1010 1011
LATITUDE (N_PROF) float64 23.08 27.41 23.79 ... 23.93 25.86
LONGITUDE (N_PROF) float64 -34.05 -32.88 ... -34.35 -30.77
TIME (N_PROF) datetime64[ns] 2005-01-03T05:11:07 ... 20...
Data variables: (12/14)
CONFIG_MISSION_NUMBER (N_PROF) int64 1 1 1 1 1 1 1 1 ... 3 -1 21 -1 3 -1 23
CYCLE_NUMBER (N_PROF) int64 103 102 104 104 103 ... 79 85 299 160
DATA_MODE (N_PROF) <U1 'D' 'D' 'D' 'D' 'D' ... 'A' 'A' 'A' 'A'
DIRECTION (N_PROF) <U1 'A' 'A' 'A' 'A' 'A' ... 'A' 'A' 'A' 'A'
PLATFORM_NUMBER (N_PROF) int64 1900073 1900075 ... 6901146 6901206
POSITION_QC (N_PROF) int64 1 1 1 1 1 1 1 1 1 ... 1 1 1 1 1 1 1 1
... ...
PSAL (N_PROF, N_LEVELS) float64 37.46 37.46 ... nan nan
PSAL_QC (N_PROF) int64 1 2 1 1 2 1 1 2 1 ... 1 1 1 1 1 1 1 1
TEMP (N_PROF, N_LEVELS) float64 24.08 24.08 ... nan nan
TEMP_QC (N_PROF) int64 1 1 1 1 1 1 1 1 1 ... 1 1 1 1 1 1 1 1
THETA (N_PROF, N_LEVELS) float64 24.3 24.3 24.3 ... nan nan
TIME_QC (N_PROF) int64 1 1 1 1 1 1 1 1 1 ... 1 1 1 1 1 1 1 1
Attributes:
DATA_ID: ARGO
DOI: http://doi.org/10.17882/42182
Fetched_from: https://www.ifremer.fr/erddap
Fetched_by: deepak
Fetched_date: 2021/05/10
Fetched_constraints: [x=-35.00/-25.00; y=23.00/28.00; z=0.0/2000.0; t=20...
Fetched_uri: https://www.ifremer.fr/erddap/tabledap/ArgoFloats.n...
history: Variables filtered according to DATA_MODE; Variable...- N_PROF: 4386
- N_LEVELS: 1012
- N_PROF(N_PROF)int6426 46 2 27 ... 3983 471 3677 3975
array([ 26, 46, 2, ..., 471, 3677, 3975])
- N_LEVELS(N_LEVELS)int640 1 2 3 4 ... 1008 1009 1010 1011
array([ 0, 1, 2, ..., 1009, 1010, 1011])
- LATITUDE(N_PROF)float6423.08 27.41 23.79 ... 23.93 25.86
- _CoordinateAxisType :
- Lat
- actual_range :
- [6.98806449e-320 6.31920604e+268]
- axis :
- Y
- colorBarMaximum :
- 90.0
- colorBarMinimum :
- -90.0
- ioos_category :
- Location
- long_name :
- Latitude of the station, best estimate
- standard_name :
- latitude
- units :
- degrees_north
- valid_max :
- 90.0
- valid_min :
- -90.0
array([23.079 , 27.407 , 23.785 , ..., 27.62774667, 23.9312 , 25.865 ]) - LONGITUDE(N_PROF)float64-34.05 -32.88 ... -34.35 -30.77
- _CoordinateAxisType :
- Lon
- actual_range :
- [1.02468406e-030 1.09650976e+227]
- axis :
- X
- colorBarMaximum :
- 180.0
- colorBarMinimum :
- -180.0
- ioos_category :
- Location
- long_name :
- Longitude of the station, best estimate
- standard_name :
- longitude
- units :
- degrees_east
- valid_max :
- 180.0
- valid_min :
- -180.0
array([-34.046 , -32.875 , -25.578 , ..., -29.39144333, -34.3531 , -30.765 ]) - TIME(N_PROF)datetime64[ns]2005-01-03T05:11:07 ... 2020-12-...
- _CoordinateAxisType :
- Time
- actual_range :
- [1.04831271e-309 1.04662720e-309]
- axis :
- T
- ioos_category :
- Time
- long_name :
- Julian day (UTC) of the station relative to REFERENCE_DATE_TIME
- standard_name :
- time
- time_origin :
- 01-JAN-1970 00:00:00
array(['2005-01-03T05:11:07.000000000', '2005-01-04T13:14:52.000000000', '2005-01-08T21:29:31.000000000', ..., '2020-12-28T20:39:20.000000000', '2020-12-29T11:36:29.000000000', '2020-12-29T13:43:29.000000000'], dtype='datetime64[ns]')
- CONFIG_MISSION_NUMBER(N_PROF)int641 1 1 1 1 1 1 ... -1 21 -1 3 -1 23
- actual_range :
- [ -1 33554432]
- colorBarMaximum :
- 100.0
- colorBarMinimum :
- 0.0
- conventions :
- 1...N, 1 : first complete mission
- ioos_category :
- Statistics
- long_name :
- Unique number denoting the missions performed by the float
- casted :
- 1
array([ 1, 1, 1, ..., 3, -1, 23])
- CYCLE_NUMBER(N_PROF)int64103 102 104 104 ... 79 85 299 160
- long_name :
- Float cycle number
- convention :
- 0..N, 0 : launch cycle (if exists), 1 : first complete cycle
- casted :
- 1
array([103, 102, 104, ..., 85, 299, 160])
- DATA_MODE(N_PROF)<U1'D' 'D' 'D' 'D' ... 'A' 'A' 'A' 'A'
- long_name :
- Delayed mode or real time data
- convention :
- R : real time; D : delayed mode; A : real time with adjustment
- casted :
- 1
array(['D', 'D', 'D', ..., 'A', 'A', 'A'], dtype='<U1')
- DIRECTION(N_PROF)<U1'A' 'A' 'A' 'A' ... 'A' 'A' 'A' 'A'
- long_name :
- Direction of the station profiles
- convention :
- A: ascending profiles, D: descending profiles
- casted :
- 1
array(['A', 'A', 'A', ..., 'A', 'A', 'A'], dtype='<U1')
- PLATFORM_NUMBER(N_PROF)int641900073 1900075 ... 6901146 6901206
- long_name :
- Float unique identifier
- convention :
- WMO float identifier : A9IIIII
- casted :
- 1
array([1900073, 1900075, 1900072, ..., 3901989, 6901146, 6901206])
- POSITION_QC(N_PROF)int641 1 1 1 1 1 1 1 ... 1 1 1 1 1 1 1 1
- long_name :
- Global quality flag of POSITION_QC profile
- convention :
- Argo reference table 2a
- casted :
- 1
array([1, 1, 1, ..., 1, 1, 1])
- PRES(N_PROF, N_LEVELS)float644.2 12.8 22.3 32.6 ... nan nan nan
- long_name :
- Sea Pressure
- standard_name :
- sea_water_pressure
- units :
- decibar
- valid_min :
- 0.0
- valid_max :
- 12000.0
- resolution :
- 0.1
- axis :
- Z
- casted :
- 1
array([[ 4.19999981, 12.80000019, 22.29999924, ..., nan, nan, nan], [ 4.5 , 13.5 , 23.79999924, ..., nan, nan, nan], [ 4.0999999 , 5.19999981, 15.19999981, ..., nan, nan, nan], ..., [ 3.70000005, 4.5999999 , 5.5999999 , ..., nan, nan, nan], [24.39999962, 30.5 , 35.29999924, ..., nan, nan, nan], [ 3.97000003, 5.86999989, 7.86999989, ..., nan, nan, nan]]) - PRES_QC(N_PROF)int641 1 1 1 1 1 1 1 ... 1 1 1 1 1 1 1 1
- long_name :
- Global quality flag of PRES_QC profile
- convention :
- Argo reference table 2a
- casted :
- 1
array([1, 1, 1, ..., 1, 1, 1])
- PSAL(N_PROF, N_LEVELS)float6437.46 37.46 37.46 ... nan nan nan
- long_name :
- PRACTICAL SALINITY
- standard_name :
- sea_water_salinity
- units :
- psu
- valid_min :
- 0.0
- valid_max :
- 43.0
- resolution :
- 0.001
- casted :
- 1
array([[37.46099854, 37.46099091, 37.46099091, ..., nan, nan, nan], [37.20439148, 37.20436096, 37.20436096, ..., nan, nan, nan], [37.22338104, 37.22238922, 37.22238922, ..., nan, nan, nan], ..., [37.36199951, 37.36299896, 37.36500168, ..., nan, nan, nan], [37.44200134, 37.44200134, 37.44200134, ..., nan, nan, nan], [37.4108696 , 37.41289902, 37.41300964, ..., nan, nan, nan]]) - PSAL_QC(N_PROF)int641 2 1 1 2 1 1 2 ... 1 1 1 1 1 1 1 1
- long_name :
- Global quality flag of PSAL_QC profile
- convention :
- Argo reference table 2a
- casted :
- 1
array([1, 2, 1, ..., 1, 1, 1])
- TEMP(N_PROF, N_LEVELS)float6424.08 24.08 24.09 ... nan nan nan
- long_name :
- SEA TEMPERATURE IN SITU ITS-90 SCALE
- standard_name :
- sea_water_temperature
- units :
- degree_Celsius
- valid_min :
- -2.0
- valid_max :
- 40.0
- resolution :
- 0.001
- casted :
- 1
array([[24.08300018, 24.08499908, 24.08600044, ..., nan, nan, nan], [22.21299934, 22.21299934, 22.21500015, ..., nan, nan, nan], [22.14299965, 22.14699936, 22.15099907, ..., nan, nan, nan], ..., [22.13199997, 22.12999916, 22.12999916, ..., nan, nan, nan], [23.52300072, 23.52400017, 23.52199936, ..., nan, nan, nan], [22.60199928, 22.60700035, 22.60899925, ..., nan, nan, nan]]) - TEMP_QC(N_PROF)int641 1 1 1 1 1 1 1 ... 1 1 1 1 1 1 1 1
- long_name :
- Global quality flag of TEMP_QC profile
- convention :
- Argo reference table 2a
- casted :
- 1
array([1, 1, 1, ..., 1, 1, 1])
- THETA(N_PROF, N_LEVELS)float6424.3 24.3 24.3 24.3 ... nan nan nan
- long_name :
- SEA TEMPERATURE IN SITU ITS-90 SCALE
- standard_name :
- sea_water_potential_temperature
- units :
- degC
- valid_min :
- -2.0
- valid_max :
- 40.0
- resolution :
- 0.001
- casted :
- 1
array([[24.29887065, 24.29903939, 24.29801114, ..., nan, nan, nan], [22.41709966, 22.41528073, 22.41521143, ..., nan, nan, nan], [22.34676527, 22.35056697, 22.35257413, ..., nan, nan, nan], ..., [22.3359062 , 22.33371242, 22.33351257, ..., nan, nan, nan], [23.7311773 , 23.73089706, 23.72787256, ..., nan, nan, nan], [22.80879642, 22.81344103, 22.81504269, ..., nan, nan, nan]]) - TIME_QC(N_PROF)int641 1 1 1 1 1 1 1 ... 1 1 1 1 1 1 1 1
- long_name :
- Global quality flag of TIME_QC profile
- convention :
- Argo reference table 2a
- casted :
- 1
array([1, 1, 1, ..., 1, 1, 1])
- DATA_ID :
- ARGO
- DOI :
- http://doi.org/10.17882/42182
- Fetched_from :
- https://www.ifremer.fr/erddap
- Fetched_by :
- deepak
- Fetched_date :
- 2021/05/10
- Fetched_constraints :
- [x=-35.00/-25.00; y=23.00/28.00; z=0.0/2000.0; t=2005-01-01/2010-12-31]
- Fetched_uri :
- https://www.ifremer.fr/erddap/tabledap/ArgoFloats.nc?data_mode,latitude,longitude,position_qc,time,time_qc,direction,platform_number,cycle_number,config_mission_number,vertical_sampling_scheme,pres,temp,psal,pres_qc,temp_qc,psal_qc,pres_adjusted,temp_adjusted,psal_adjusted,pres_adjusted_qc,temp_adjusted_qc,psal_adjusted_qc,pres_adjusted_error,temp_adjusted_error,psal_adjusted_error&longitude>=-35&longitude<=-25&latitude>=23&latitude<=28&pres>=0&pres<=2000&time>=1104537600.0&time<=1293753600.0&distinct()&orderBy("time,pres")
- history :
- Variables filtered according to DATA_MODE; Variables selected according to QC; Transformed with point2profile
argo_fine_profiles
<xarray.Dataset>
Dimensions: (profile: 3583, pressure_: 20, nbnds: 2)
Coordinates:
CONFIG_MISSION_NUMBER (profile) int64 1 1 1 1 1 1 1 1 1 ... 5 4 2 5 2 2 5 2
CYCLE_NUMBER (profile) int64 103 104 104 105 ... 180 181 229 182
PLATFORM_NUMBER (profile) int64 1900073 1900072 ... 4901585 6902572
flag (profile, pressure_) float64 -1.0 -1.0 ... -1.0 nan
latitude (profile) float64 23.08 23.79 23.11 ... 26.0 27.96
longitude (profile) float64 -34.05 -25.58 ... -32.41 -27.37
npts (profile, pressure_) float64 0.0 0.0 0.0 ... 0.0 nan
p_bounds (profile, pressure_, nbnds) float64 nan nan ... nan
pressure (profile, pressure_) float64 nan nan nan ... nan nan
* pressure_ (pressure_) int64 0 1 2 3 4 5 6 ... 14 15 16 17 18 19
γ_bounds (profile, pressure_, nbnds) float64 nan nan ... nan
γmean (profile, pressure_) float64 nan nan nan ... nan nan
Dimensions without coordinates: profile, nbnds
Data variables: (12/14)
KtTz (profile, pressure_) float64 nan nan nan ... nan nan
Kρ (profile, pressure_) float64 nan nan nan ... nan nan
N2mean (profile, pressure_) float64 nan nan nan ... nan nan
Tmld (profile) float64 112.4 95.1 112.3 ... 77.88 89.6
Tmode (profile) float64 132.5 135.3 132.2 ... 79.96 99.7
Tzlin (profile, pressure_) float64 nan nan nan ... nan nan
... ...
ε (profile, pressure_) float64 nan nan nan ... nan nan
ξvar (profile, pressure_) float64 nan nan nan ... nan nan
ξvargm (profile, pressure_) float64 nan nan nan ... nan nan
σmld (profile) float64 92.4 95.1 112.3 ... 80.2 77.88 89.6
σmode (profile) float64 132.5 115.1 132.2 ... 79.96 99.7
χ (profile, pressure_) float64 nan nan nan ... nan nan- profile: 3583
- pressure_: 20
- nbnds: 2
- CONFIG_MISSION_NUMBER(profile)int641 1 1 1 1 1 1 1 ... 5 4 2 5 2 2 5 2
array([1, 1, 1, ..., 2, 5, 2])
- CYCLE_NUMBER(profile)int64103 104 104 105 ... 180 181 229 182
array([103, 104, 104, ..., 181, 229, 182])
- PLATFORM_NUMBER(profile)int641900073 1900072 ... 4901585 6902572
array([1900073, 1900072, 1900073, ..., 6902572, 4901585, 6902572])
- flag(profile, pressure_)float64-1.0 -1.0 -1.0 ... -1.0 -1.0 nan
- flag_meanings :
- too_coarse too_short N²_variance_too_high too_unstratified too_little_bandwidth no_internal_waves good_data
- flag_values :
- [-2, -1, 1, 2, 3, 4, 5]
array([[-1., -1., -1., ..., -1., nan, nan], [-1., -1., -1., ..., -1., nan, nan], [-1., -1., -1., ..., -1., nan, nan], ..., [ 5., 5., -1., ..., -1., -1., -1.], [ 5., 5., 5., ..., nan, nan, nan], [ 5., 5., -1., ..., -1., -1., nan]]) - latitude(profile)float6423.08 23.79 23.11 ... 26.0 27.96
array([23.079 , 23.785 , 23.115 , ..., 27.692 , 26.00449, 27.961 ])
- longitude(profile)float64-34.05 -25.58 ... -32.41 -27.37
array([-34.046 , -25.578 , -34.184 , ..., -27.586 , -32.41227, -27.371 ]) - npts(profile, pressure_)float640.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 nan
- description :
- number of points in segment
array([[ 0., 0., 0., ..., 0., nan, nan], [ 0., 0., 0., ..., 0., nan, nan], [ 0., 0., 0., ..., 0., nan, nan], ..., [ 19., 14., 4., ..., 0., 0., 0.], [100., 99., 99., ..., nan, nan, nan], [ 19., 15., 4., ..., 0., 0., nan]]) - p_bounds(profile, pressure_, nbnds)float64nan nan nan nan ... nan nan nan nan
array([[[ nan, nan], [ nan, nan], [ nan, nan], ..., [ nan, nan], [ nan, nan], [ nan, nan]], [[ nan, nan], [ nan, nan], [ nan, nan], ..., [ nan, nan], [ nan, nan], [ nan, nan]], [[ nan, nan], [ nan, nan], [ nan, nan], ..., ... ..., [ nan, nan], [ nan, nan], [ nan, nan]], [[101.95999908, 300. ], [204. , 400. ], [302. , 498. ], ..., [ nan, nan], [ nan, nan], [ nan, nan]], [[119.69999695, 299.8999939 ], [209.69999695, 350.3999939 ], [ nan, nan], ..., [ nan, nan], [ nan, nan], [ nan, nan]]]) - pressure(profile, pressure_)float64nan nan nan nan ... nan nan nan nan
- axis :
- Z
- bounds :
- p_bounds
- positive :
- down
- standard_name :
- sea_water_pressure
array([[ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], ..., [209.99999619, 285.05000305, nan, ..., nan, nan, nan], [200.97999954, 302. , 400. , ..., nan, nan, nan], [209.79999542, 280.04999542, nan, ..., nan, nan, nan]]) - pressure_(pressure_)int640 1 2 3 4 5 6 ... 14 15 16 17 18 19
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19]) - γ_bounds(profile, pressure_, nbnds)float64nan nan nan nan ... nan nan nan nan
array([[[ nan, nan], [ nan, nan], [ nan, nan], ..., [ nan, nan], [ nan, nan], [ nan, nan]], [[ nan, nan], [ nan, nan], [ nan, nan], ..., [ nan, nan], [ nan, nan], [ nan, nan]], [[ nan, nan], [ nan, nan], [ nan, nan], ..., ... ..., [ nan, nan], [ nan, nan], [ nan, nan]], [[26.01802761, 26.67128058], [26.37578484, 26.86517511], [26.67407304, 27.01725056], ..., [ nan, nan], [ nan, nan], [ nan, nan]], [[26.18965179, 26.68132286], [26.48551074, 26.79334891], [ nan, nan], ..., [ nan, nan], [ nan, nan], [ nan, nan]]]) - γmean(profile, pressure_)float64nan nan nan nan ... nan nan nan nan
- bounds :
- γ_bounds
- casted :
- 1
- long_name :
- $γ_n$
- resolution :
- 0.001
- standard_name :
- neutral_density
- units :
- kg/m3
- valid_max :
- 43.0
- valid_min :
- 0.0
array([[ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], ..., [26.48515563, 26.65522995, nan, ..., nan, nan, nan], [26.36340187, 26.65571826, 26.86758021, ..., nan, nan, nan], [26.46223804, 26.63708881, nan, ..., nan, nan, nan]])
- KtTz(profile, pressure_)float64nan nan nan nan ... nan nan nan nan
- long_name :
- $K_ρθ_z$
- units :
- °Cm/s
array([[ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], ..., [7.29478138e-08, 2.69641156e-08, nan, ..., nan, nan, nan], [3.17735223e-07, 4.86244183e-08, 5.21133852e-08, ..., nan, nan, nan], [1.66432455e-07, 8.03898726e-08, nan, ..., nan, nan, nan]]) - Kρ(profile, pressure_)float64nan nan nan nan ... nan nan nan nan
- long_name :
- $K_ρ$
- units :
- m²/s
array([[ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], ..., [3.59037654e-06, 1.43011102e-06, nan, ..., nan, nan, nan], [1.28061120e-05, 2.24881020e-06, 3.34742500e-06, ..., nan, nan, nan], [7.85027337e-06, 4.29463023e-06, nan, ..., nan, nan, nan]]) - N2mean(profile, pressure_)float64nan nan nan nan ... nan nan nan nan
- description :
- mean of quadratic fit of N² with pressure
- long_name :
- $N²$
array([[ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], ..., [2.20933756e-05, 2.15535867e-05, nan, ..., nan, nan, nan], [3.14779142e-05, 2.43592211e-05, 1.76240853e-05, ..., nan, nan, nan], [2.60349795e-05, 2.11982850e-05, nan, ..., nan, nan, nan]]) - Tmld(profile)float64112.4 95.1 112.3 ... 77.88 89.6
- description :
- ΔT criterion applied first time
- units :
- m
array([112.40000153, 95.09999847, 112.30000305, ..., 80.19999695, 77.87999725, 89.59999847]) - Tmode(profile)float64132.5 135.3 132.2 ... 79.96 99.7
- description :
- ΔT criterion applied second time
- units :
- m
array([132.5 , 135.30000305, 132.19999695, ..., 90.19999695, 79.95999908, 99.69999695]) - Tzlin(profile, pressure_)float64nan nan nan nan ... nan nan nan nan
- description :
- linear fit of T vs pressure
- long_name :
- $T_z^{lin}$
array([[ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], ..., [0.02004671, 0.01860232, nan, ..., nan, nan, nan], [0.02470548, 0.02164925, 0.01549148, ..., nan, nan, nan], [0.02100139, 0.01878803, nan, ..., nan, nan, nan]]) - Tzmean(profile, pressure_)float64nan nan nan nan ... nan nan nan nan
- description :
- mean of quadratic fit of Tz with pressure; like N² fitting for strain
- long_name :
- $T_z^{quad}$
array([[ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], ..., [0.02031759, 0.01885456, nan, ..., nan, nan, nan], [0.02481122, 0.02162229, 0.0155682 , ..., nan, nan, nan], [0.02120085, 0.01871869, nan, ..., nan, nan, nan]]) - mean_dTdz_seg(profile, pressure_)float64nan nan nan nan ... nan nan nan nan
- description :
- mean of dTdz values in segment
- long_name :
- $⟨T_z⟩$
array([[ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], ..., [0.02099382, 0.01903718, nan, ..., nan, nan, nan], [0.02480703, 0.02181404, 0.0155757 , ..., nan, nan, nan], [0.02209709, 0.01862588, nan, ..., nan, nan, nan]]) - ε(profile, pressure_)float64nan nan nan nan ... nan nan nan nan
- long_name :
- $ε$
- units :
- W/kg
array([[ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], ..., [3.88852199e-10, 1.51102548e-10, nan, ..., nan, nan, nan], [1.97608548e-09, 2.68533632e-10, 2.89201090e-10, ..., nan, nan, nan], [1.00190030e-09, 4.46281609e-10, nan, ..., nan, nan, nan]]) - ξvar(profile, pressure_)float64nan nan nan nan ... nan nan nan nan
- description :
- strain variance
- long_name :
- $ξ$
array([[ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], ..., [0.09462641, 0.05380143, nan, ..., nan, nan, nan], [0.21496643, 0.21644211, 0.21566597, ..., nan, nan, nan], [0.13632284, 0.10018238, nan, ..., nan, nan, nan]]) - ξvargm(profile, pressure_)float64nan nan nan nan ... nan nan nan nan
- description :
- GM strain variance
- long_name :
- $ξ_{gm}$
array([[ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], ..., [0.10710723, 0.09636968, nan, ..., nan, nan, nan], [0.12808864, 0.30392183, 0.24419597, ..., nan, nan, nan], [0.10559203, 0.10383195, nan, ..., nan, nan, nan]]) - σmld(profile)float6492.4 95.1 112.3 ... 80.2 77.88 89.6
- description :
- Δσ criterion applied first time
- units :
- m
array([ 92.40000153, 95.09999847, 112.30000305, ..., 80.19999695, 77.87999725, 89.59999847]) - σmode(profile)float64132.5 115.1 132.2 ... 79.96 99.7
- description :
- Δσ criterion applied second time
- units :
- m
array([132.5 , 115.09999847, 132.19999695, ..., 90.19999695, 79.95999908, 99.69999695]) - χ(profile, pressure_)float64nan nan nan nan ... nan nan nan nan
- long_name :
- $χ$
- units :
- °C²/s
array([[ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], ..., [2.96424817e-09, 1.01679314e-09, nan, ..., nan, nan, nan], [1.57667951e-08, 2.10274220e-09, 1.62262331e-09, ..., nan, nan, nan], [7.05701851e-09, 3.00958698e-09, nan, ..., nan, nan, nan]])
profiles = []
for nprof in tqdm.notebook.tqdm(range(argo_fine_profiles.sizes["profile"])):
prof = argo_fine_profiles.isel(profile=nprof)
selected = argo_profiles.query(
{
"N_PROF": f"CYCLE_NUMBER == {prof.CYCLE_NUMBER.item()} & PLATFORM_NUMBER == {prof.PLATFORM_NUMBER.item()} & DIRECTION == 'A'"
}
)
assert selected.sizes["N_PROF"] == 1
selected["fine_deepest_bin"] = (
"N_PROF",
[prof.pressure.max("pressure_").data],
)
profiles.append(selected)
selected
<xarray.Dataset>
Dimensions: (N_PROF: 1, N_LEVELS: 1012)
Coordinates:
* N_PROF (N_PROF) int64 4238
* N_LEVELS (N_LEVELS) int64 0 1 2 3 4 ... 1008 1009 1010 1011
LATITUDE (N_PROF) float64 27.96
LONGITUDE (N_PROF) float64 -27.37
TIME (N_PROF) datetime64[ns] 2019-11-23T14:25:04
Data variables: (12/15)
CONFIG_MISSION_NUMBER (N_PROF) int64 2
CYCLE_NUMBER (N_PROF) int64 182
DATA_MODE (N_PROF) <U1 'D'
DIRECTION (N_PROF) <U1 'A'
PLATFORM_NUMBER (N_PROF) int64 6902572
POSITION_QC (N_PROF) int64 1
... ...
PSAL_QC (N_PROF) int64 1
TEMP (N_PROF, N_LEVELS) float64 23.61 23.59 ... nan nan
TEMP_QC (N_PROF) int64 1
THETA (N_PROF, N_LEVELS) float64 23.83 23.81 ... nan nan
TIME_QC (N_PROF) int64 1
fine_deepest_bin (N_PROF) float64 280.0
Attributes:
DATA_ID: ARGO
DOI: http://doi.org/10.17882/42182
Fetched_from: https://www.ifremer.fr/erddap
Fetched_by: deepak
Fetched_date: 2021/05/10
Fetched_constraints: [x=-35.00/-25.00; y=23.00/28.00; z=0.0/2000.0; t=20...
Fetched_uri: https://www.ifremer.fr/erddap/tabledap/ArgoFloats.n...
history: Variables filtered according to DATA_MODE; Variable...- N_PROF: 1
- N_LEVELS: 1012
- N_PROF(N_PROF)int644238
array([4238])
- N_LEVELS(N_LEVELS)int640 1 2 3 4 ... 1008 1009 1010 1011
array([ 0, 1, 2, ..., 1009, 1010, 1011])
- LATITUDE(N_PROF)float6427.96
- _CoordinateAxisType :
- Lat
- actual_range :
- [6.98806449e-320 6.31920604e+268]
- axis :
- Y
- colorBarMaximum :
- 90.0
- colorBarMinimum :
- -90.0
- ioos_category :
- Location
- long_name :
- Latitude of the station, best estimate
- standard_name :
- latitude
- units :
- degrees_north
- valid_max :
- 90.0
- valid_min :
- -90.0
array([27.961])
- LONGITUDE(N_PROF)float64-27.37
- _CoordinateAxisType :
- Lon
- actual_range :
- [1.02468406e-030 1.09650976e+227]
- axis :
- X
- colorBarMaximum :
- 180.0
- colorBarMinimum :
- -180.0
- ioos_category :
- Location
- long_name :
- Longitude of the station, best estimate
- standard_name :
- longitude
- units :
- degrees_east
- valid_max :
- 180.0
- valid_min :
- -180.0
array([-27.371])
- TIME(N_PROF)datetime64[ns]2019-11-23T14:25:04
- _CoordinateAxisType :
- Time
- actual_range :
- [1.04831271e-309 1.04662720e-309]
- axis :
- T
- ioos_category :
- Time
- long_name :
- Julian day (UTC) of the station relative to REFERENCE_DATE_TIME
- standard_name :
- time
- time_origin :
- 01-JAN-1970 00:00:00
array(['2019-11-23T14:25:04.000000000'], dtype='datetime64[ns]')
- CONFIG_MISSION_NUMBER(N_PROF)int642
- actual_range :
- [ -1 33554432]
- colorBarMaximum :
- 100.0
- colorBarMinimum :
- 0.0
- conventions :
- 1...N, 1 : first complete mission
- ioos_category :
- Statistics
- long_name :
- Unique number denoting the missions performed by the float
- casted :
- 1
array([2])
- CYCLE_NUMBER(N_PROF)int64182
- long_name :
- Float cycle number
- convention :
- 0..N, 0 : launch cycle (if exists), 1 : first complete cycle
- casted :
- 1
array([182])
- DATA_MODE(N_PROF)<U1'D'
- long_name :
- Delayed mode or real time data
- convention :
- R : real time; D : delayed mode; A : real time with adjustment
- casted :
- 1
array(['D'], dtype='<U1')
- DIRECTION(N_PROF)<U1'A'
- long_name :
- Direction of the station profiles
- convention :
- A: ascending profiles, D: descending profiles
- casted :
- 1
array(['A'], dtype='<U1')
- PLATFORM_NUMBER(N_PROF)int646902572
- long_name :
- Float unique identifier
- convention :
- WMO float identifier : A9IIIII
- casted :
- 1
array([6902572])
- POSITION_QC(N_PROF)int641
- long_name :
- Global quality flag of POSITION_QC profile
- convention :
- Argo reference table 2a
- casted :
- 1
array([1])
- PRES(N_PROF, N_LEVELS)float645.5 10.0 14.7 19.4 ... nan nan nan
- long_name :
- Sea Pressure
- standard_name :
- sea_water_pressure
- units :
- decibar
- valid_min :
- 0.0
- valid_max :
- 12000.0
- resolution :
- 0.1
- axis :
- Z
- casted :
- 1
array([[ 5.5 , 10. , 14.69999981, ..., nan, nan, nan]]) - PRES_QC(N_PROF)int641
- long_name :
- Global quality flag of PRES_QC profile
- convention :
- Argo reference table 2a
- casted :
- 1
array([1])
- PSAL(N_PROF, N_LEVELS)float6437.34 37.34 37.34 ... nan nan nan
- long_name :
- PRACTICAL SALINITY
- standard_name :
- sea_water_salinity
- units :
- psu
- valid_min :
- 0.0
- valid_max :
- 43.0
- resolution :
- 0.001
- casted :
- 1
array([[37.34207153, 37.34006882, 37.34106827, ..., nan, nan, nan]]) - PSAL_QC(N_PROF)int641
- long_name :
- Global quality flag of PSAL_QC profile
- convention :
- Argo reference table 2a
- casted :
- 1
array([1])
- TEMP(N_PROF, N_LEVELS)float6423.61 23.59 23.57 ... nan nan nan
- long_name :
- SEA TEMPERATURE IN SITU ITS-90 SCALE
- standard_name :
- sea_water_temperature
- units :
- degree_Celsius
- valid_min :
- -2.0
- valid_max :
- 40.0
- resolution :
- 0.001
- casted :
- 1
array([[23.61499977, 23.59399986, 23.56599998, ..., nan, nan, nan]]) - TEMP_QC(N_PROF)int641
- long_name :
- Global quality flag of TEMP_QC profile
- convention :
- Argo reference table 2a
- casted :
- 1
array([1])
- THETA(N_PROF, N_LEVELS)float6423.83 23.81 23.78 ... nan nan nan
- long_name :
- SEA TEMPERATURE IN SITU ITS-90 SCALE
- standard_name :
- sea_water_potential_temperature
- units :
- degC
- valid_min :
- -2.0
- valid_max :
- 40.0
- resolution :
- 0.001
- casted :
- 1
array([[23.82763461, 23.80555503, 23.77639455, ..., nan, nan, nan]]) - TIME_QC(N_PROF)int641
- long_name :
- Global quality flag of TIME_QC profile
- convention :
- Argo reference table 2a
- casted :
- 1
array([1])
- fine_deepest_bin(N_PROF)float64280.0
array([280.04999542])
- DATA_ID :
- ARGO
- DOI :
- http://doi.org/10.17882/42182
- Fetched_from :
- https://www.ifremer.fr/erddap
- Fetched_by :
- deepak
- Fetched_date :
- 2021/05/10
- Fetched_constraints :
- [x=-35.00/-25.00; y=23.00/28.00; z=0.0/2000.0; t=2005-01-01/2010-12-31]
- Fetched_uri :
- https://www.ifremer.fr/erddap/tabledap/ArgoFloats.nc?data_mode,latitude,longitude,position_qc,time,time_qc,direction,platform_number,cycle_number,config_mission_number,vertical_sampling_scheme,pres,temp,psal,pres_qc,temp_qc,psal_qc,pres_adjusted,temp_adjusted,psal_adjusted,pres_adjusted_qc,temp_adjusted_qc,psal_adjusted_qc,pres_adjusted_error,temp_adjusted_error,psal_adjusted_error&longitude>=-35&longitude<=-25&latitude>=23&latitude<=28&pres>=0&pres<=2000&time>=1104537600.0&time<=1293753600.0&distinct()&orderBy("time,pres")
- history :
- Variables filtered according to DATA_MODE; Variables selected according to QC; Transformed with point2profile
argo_fine_full_profiles = xr.concat(profiles, dim="N_PROF")
argo_fine_full_profiles["fine_deepest_bin"].attrs[
"description"
] = "Deepest bin with a finestructure estimate"
# sanity check
fields = ["CYCLE_NUMBER", "PLATFORM_NUMBER"]
for data in fields:
np.testing.assert_equal(
argo_fine_full_profiles[data].data, argo_fine_profiles[data].data
)
argo_fine_full_profiles["THETA"] = dcpy.eos.ptmp(
argo_fine_full_profiles.PSAL,
argo_fine_full_profiles.TEMP,
argo_fine_full_profiles.PRES,
natre.reference_pressure.item(),
)
argo_fine_full_profiles.to_netcdf("argo_profiles_used_for_finestructure.nc")
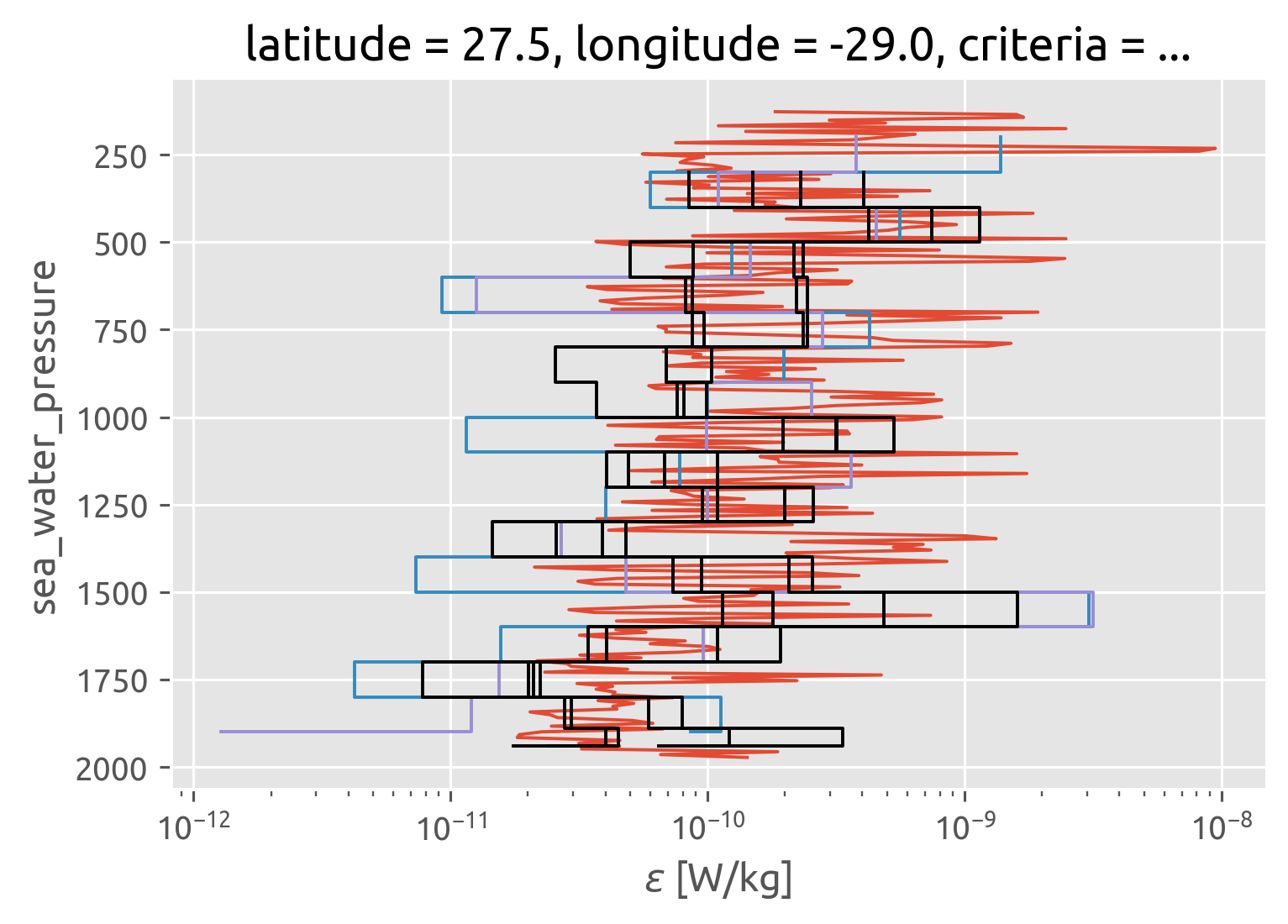
Finescale estimate with NATRE data#
def fix_profile(profile):
fixed = (
profile.unstack()
.squeeze()
.cf.guess_coord_axis()
.cf.dropna("Z", how="any")
.set_coords("pres")
.drop("depth")
.swap_dims({"depth": "pres"})
)
assert fixed.temp.isnull().sum() == 0
return fixed
stacked = (
natre.load()
.reset_coords("pres")
.cf.interpolate_na("Z")
.stack({"latlon": ["longitude", "latitude"]})
)
results = [
dask.delayed(dcpy.finestructure.process_profile)(
fix_profile(stacked.isel(latlon=[idx]))
)
for idx in range(1, stacked.sizes["latlon"])
]
profile = fix_profile(stacked.isel(latlon=[40]))
ms = dcpy.finestructure.do_mixsea_shearstrain(profile, dz_segment=200)
dc = dcpy.finestructure.process_profile(profile, dz_segment=200, criteria=None)
profile.eps.coarsen(pres=20, boundary="trim").mean().plot(y="pres")
ms.eps.isel(kind=0).plot.step(y="depth_bin", xscale="log", yincrease=False)
ms.eps.isel(kind=1).plot.step(y="depth_bin", xscale="log", yincrease=False)
for crit in dc.criteria:
dc.sel(criteria=crit).ε.cf.plot.step(color="k")
from dask.distributed import Client
client = Client("tcp://127.0.0.1:41667")
client
Client
Client-a53a850d-e9a0-11eb-8526-0028f8270ea7
| Connection method: Direct | |
| Dashboard: http://127.0.0.1:8787/status |
Scheduler Info
Scheduler
Scheduler-d6f7fb0f-8f7b-4ee3-8176-31ebb7885697
| Comm: tcp://127.0.0.1:41667 | Workers: 4 |
| Dashboard: http://127.0.0.1:8787/status | Total threads: 8 |
| Started: 1 minute ago | Total memory: 31.10 GiB |
Workers
Worker: 0
| Comm: tcp://127.0.0.1:38589 | Total threads: 2 |
| Dashboard: http://127.0.0.1:38143/status | Memory: 7.77 GiB |
| Nanny: tcp://127.0.0.1:45089 | |
| Local directory: /tmp/dask-worker-space/worker-j3tr7azt | |
| Tasks executing: 0 | Tasks in memory: 0 |
| Tasks ready: 0 | Tasks in flight: 0 |
| CPU usage: 2.0% | Last seen: Just now |
| Memory usage: 307.97 MiB | Spilled bytes: 0 B |
| Read bytes: 84.62 kiB | Write bytes: 5.47 MiB |
Worker: 1
| Comm: tcp://127.0.0.1:34279 | Total threads: 2 |
| Dashboard: http://127.0.0.1:36725/status | Memory: 7.77 GiB |
| Nanny: tcp://127.0.0.1:45517 | |
| Local directory: /tmp/dask-worker-space/worker-3ble6b2n | |
| Tasks executing: 0 | Tasks in memory: 0 |
| Tasks ready: 0 | Tasks in flight: 0 |
| CPU usage: 2.0% | Last seen: Just now |
| Memory usage: 305.44 MiB | Spilled bytes: 0 B |
| Read bytes: 102.43 kiB | Write bytes: 12.61 MiB |
Worker: 2
| Comm: tcp://127.0.0.1:36211 | Total threads: 2 |
| Dashboard: http://127.0.0.1:45765/status | Memory: 7.77 GiB |
| Nanny: tcp://127.0.0.1:33209 | |
| Local directory: /tmp/dask-worker-space/worker-oa6g1apu | |
| Tasks executing: 0 | Tasks in memory: 0 |
| Tasks ready: 0 | Tasks in flight: 0 |
| CPU usage: 2.0% | Last seen: Just now |
| Memory usage: 304.01 MiB | Spilled bytes: 0 B |
| Read bytes: 86.19 kiB | Write bytes: 5.47 MiB |
Worker: 3
| Comm: tcp://127.0.0.1:43585 | Total threads: 2 |
| Dashboard: http://127.0.0.1:45443/status | Memory: 7.77 GiB |
| Nanny: tcp://127.0.0.1:37171 | |
| Local directory: /tmp/dask-worker-space/worker-hnwyf6y6 | |
| Tasks executing: 0 | Tasks in memory: 0 |
| Tasks ready: 0 | Tasks in flight: 0 |
| CPU usage: 4.0% | Last seen: Just now |
| Memory usage: 306.35 MiB | Spilled bytes: 0 B |
| Read bytes: 62.84 kiB | Write bytes: 8.10 MiB |
computed = dask.compute(results, scheduler="threads")
/home/deepak/work/python/dcpy/dcpy/finestructure.py:311: RuntimeWarning: invalid value encountered in true_divide
scale = (ξvar / ξgmvar) ** 2 * h_Rω * L_Nf
/home/deepak/work/python/dcpy/dcpy/finestructure.py:311: RuntimeWarning: invalid value encountered in true_divide
scale = (ξvar / ξgmvar) ** 2 * h_Rω * L_Nf
def fix_result(profile):
return (
profile.rename_dims({"pressure": "pressure_"})
.reset_coords()
.set_coords(["latitude", "longitude"])
.expand_dims(["latitude", "longitude"])
.assign_coords({"pressure_": np.arange(profile.sizes["pressure"])})
)
combined = xr.combine_by_coords([fix_result(ds) for ds in computed[0]])
combined = combined.cf.guess_coord_axis()
combined.to_netcdf("../datasets/natre-finescale.nc")
avg = combined.groupby_bins("pressure", np.arange(150, 2060, 100)).mean()
avg.N2mean.plot()
micro.N2.plot(x="pres")
micro.N2_m.plot(x="pres")
[<matplotlib.lines.Line2D at 0x7fefa3e34760>]
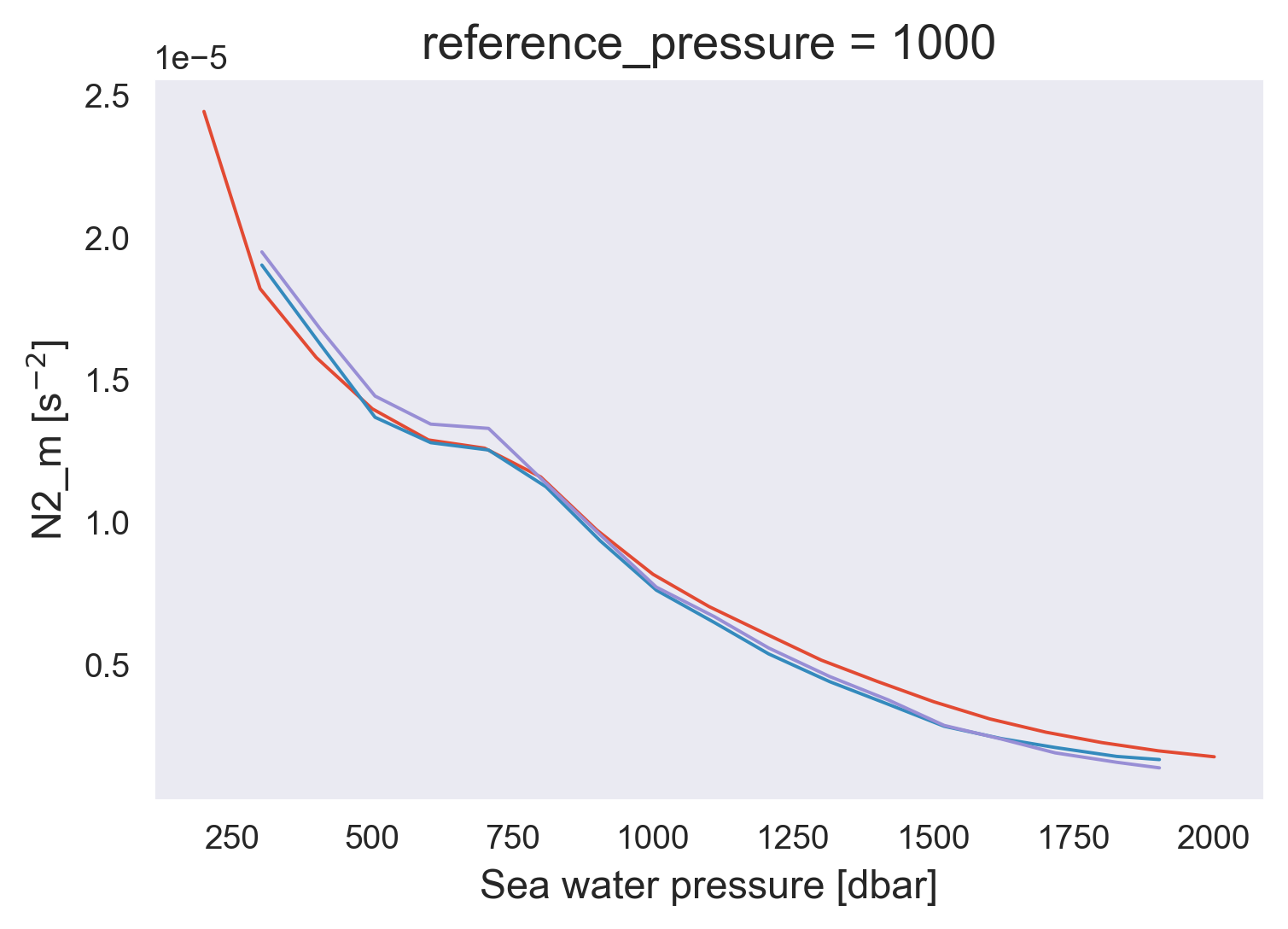
avg.Tzmean.plot()
micro.Tz.plot(x="pres")
micro.dTdz_m.plot(x="pres")
[<matplotlib.lines.Line2D at 0x7feff10f9400>]
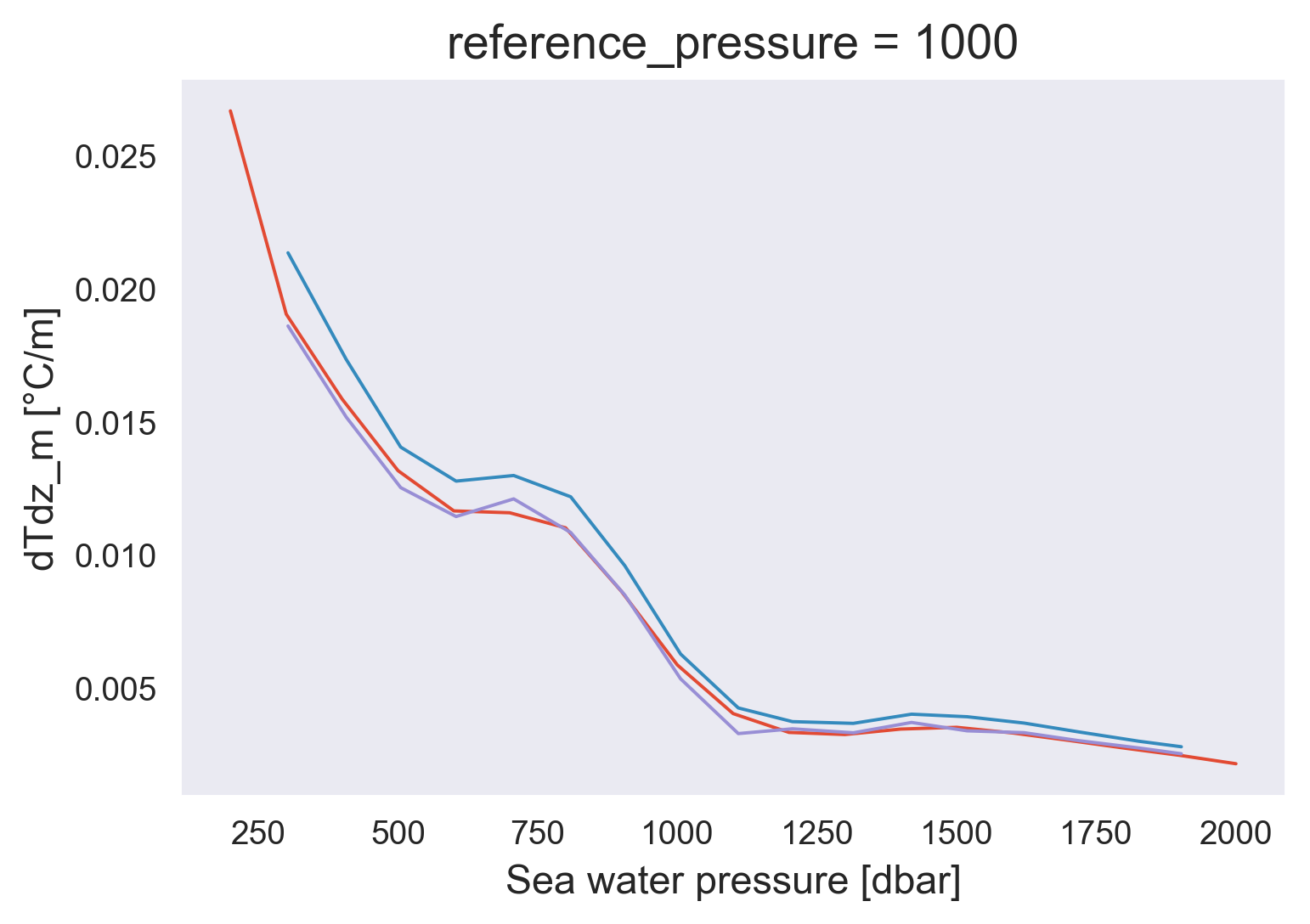
avg.ε.plot(y="pressure_bins", hue="criteria", xscale="log")
natre.eps.mean(["latitude", "longitude"]).coarsen(
depth=50, boundary="trim"
).mean().plot(y="depth")
[<matplotlib.lines.Line2D at 0x7fefdd4f9eb0>]
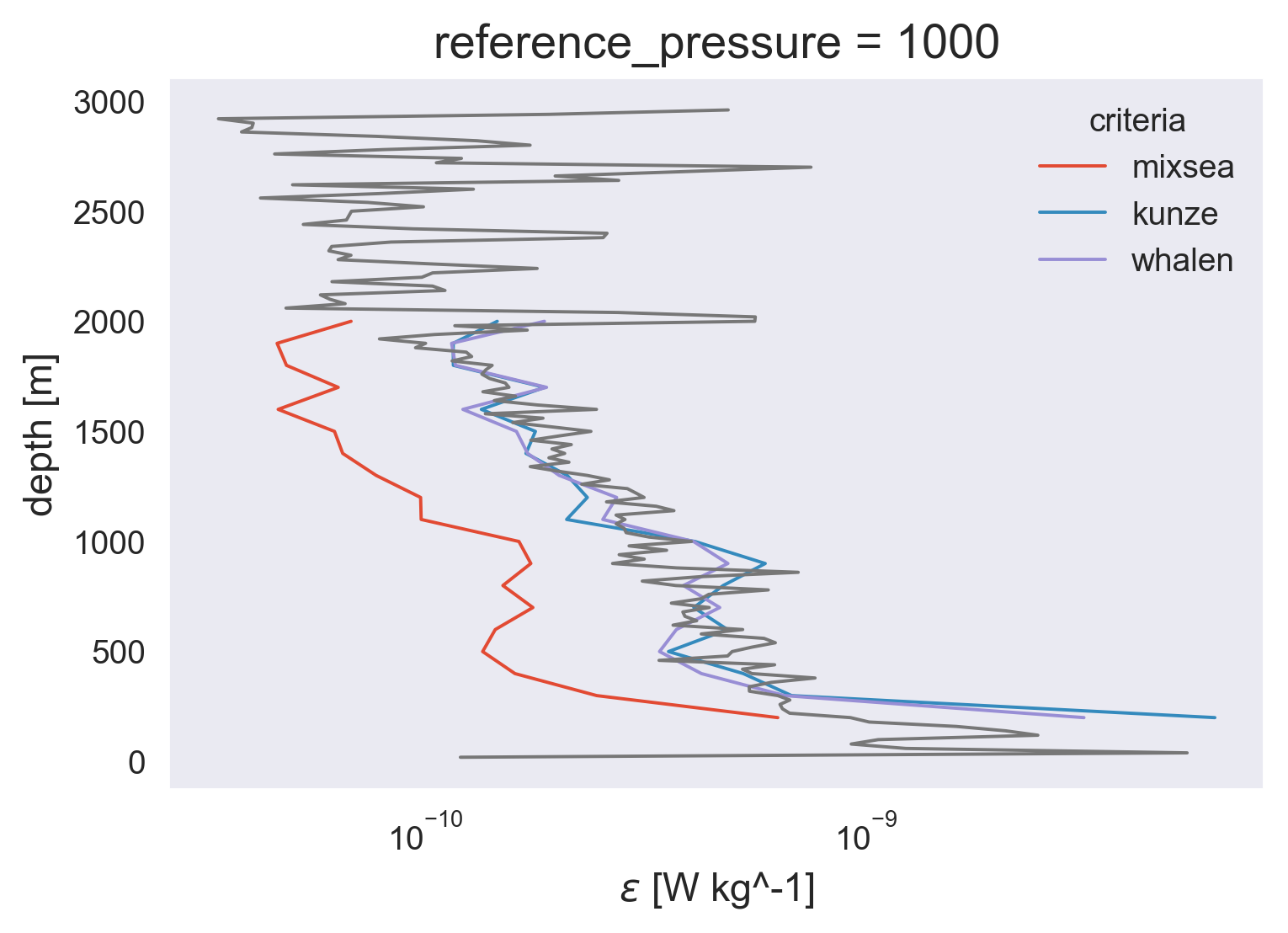
(avg.Kρ).plot(y="pressure_bins", hue="criteria", xscale="log")
natre.Krho.coarsen(depth=200, boundary="trim").mean().mean(
["latitude", "longitude"]
).plot.step(y="depth")
micro.Krho.plot.step(y="pres")
micro.Krho_m.plot.step(y="pres", yincrease=False)
plt.gca().grid(True)
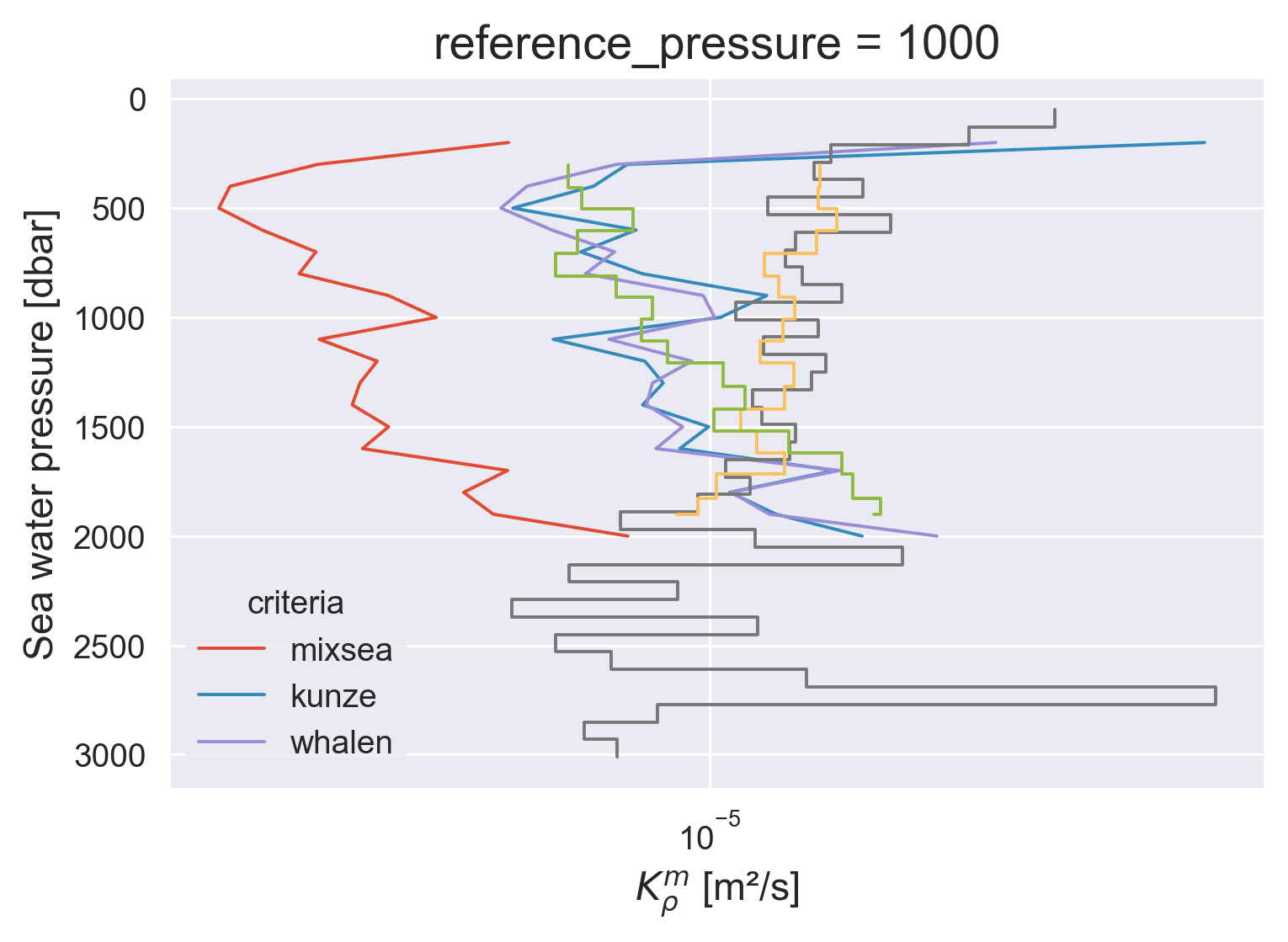
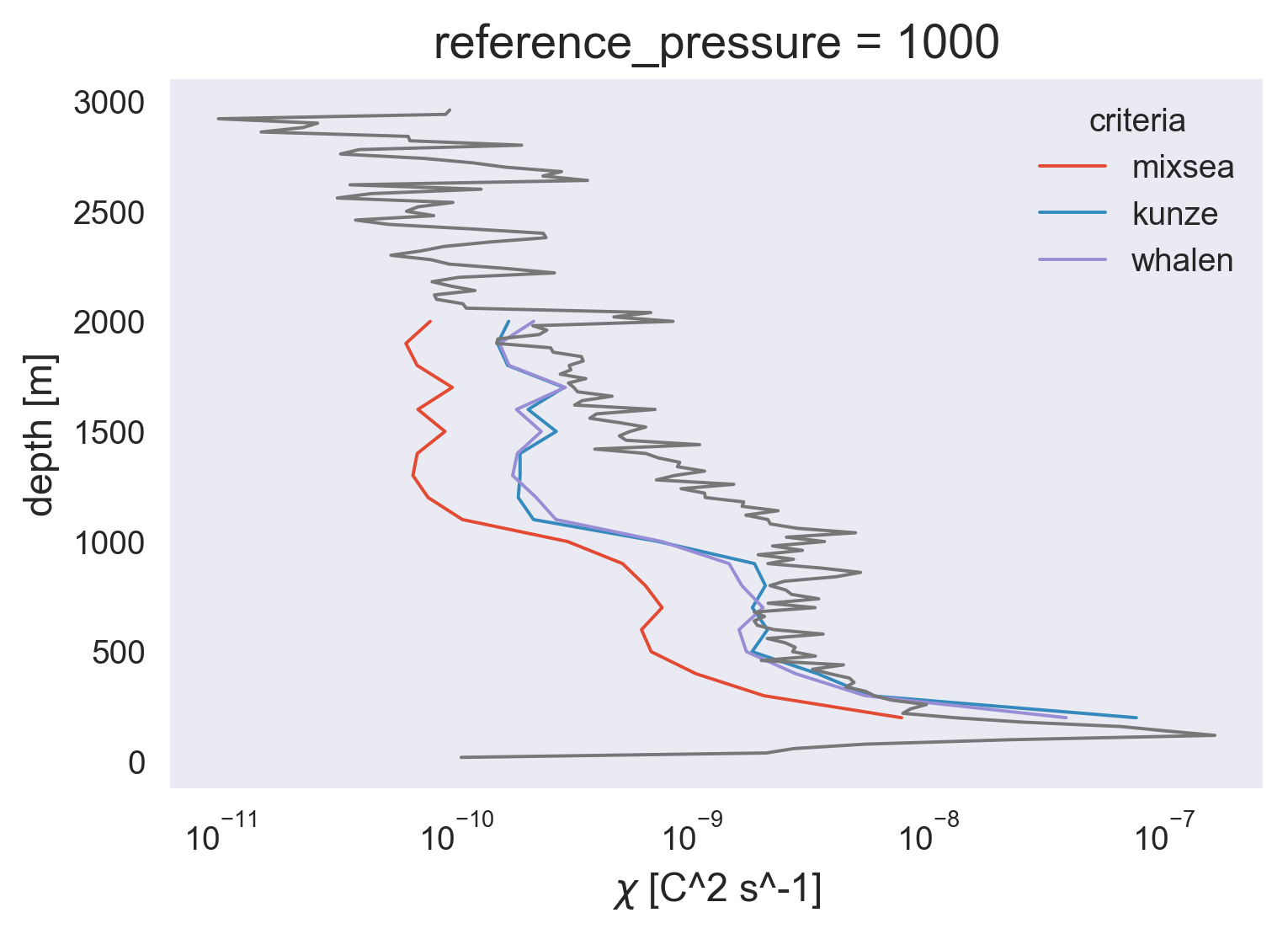
(avg.χ).plot(y="pressure_bins", hue="criteria", xscale="log")
natre.chi.mean(["latitude", "longitude"]).coarsen(
depth=50, boundary="trim"
).mean().plot(y="depth")
[<matplotlib.lines.Line2D at 0x7fefe462fc10>]
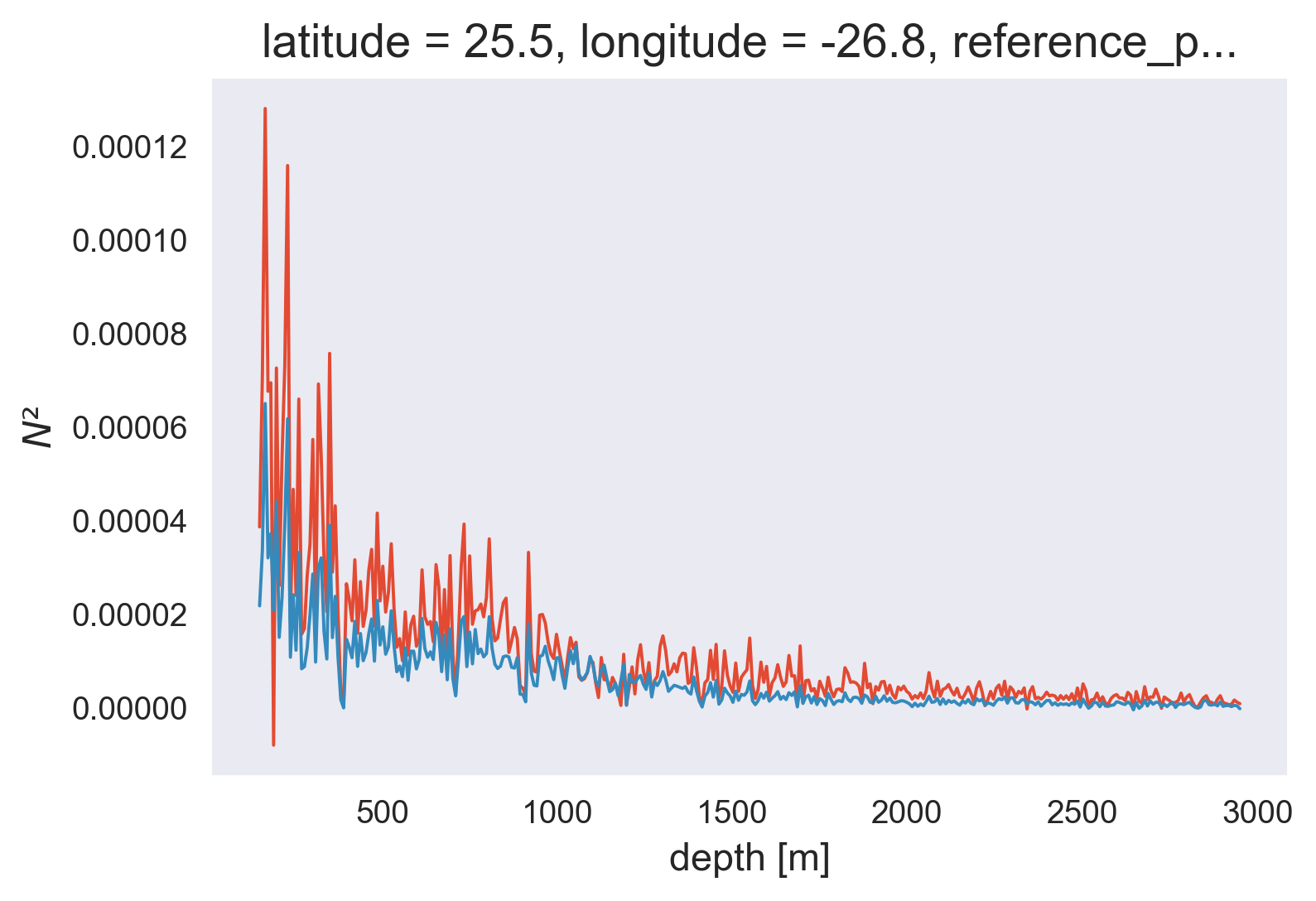
(9.81 * 1.7e-4 * profile.Tz).plot()
profile.N2.plot()
[<matplotlib.lines.Line2D at 0x7fefdc273d30>]
profile = natre.isel(latitude=5, longitude=9).coarsen(depth=20, boundary="trim").mean()
profile.Kt.plot()
profile.Krho.plot(yscale="log")
[<matplotlib.lines.Line2D at 0x7fefddf66670>]