T-S diagrams from TAO & Argo at (0, 145W→135W)#
In this notebook 140W really means (145W → 135W)
looking at TS diagrams across latitudes.
During TIW activity do we see more stirring near the equator or off the equator?
If salinity is a minor contributor to density, do we really expect to see T-S scatter as a signal of eddy stirring?
(Surprising conclusions)
T-S spread is similar in all ENSO phases. There is some separation in temperature (as expected)
Latitudinal variation in T-S spread is big; but again ENSO variation is minimal at all latitudes I’ve checked.
%load_ext watermark
import cf_xarray
import dcpy
import distributed
import matplotlib as mpl
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import pump
import xfilter
import xgcm
import xhistogram
import eddydiff as ed
import xarray as xr
xr.set_options(keep_attrs=True)
%watermark -iv
plt.rcParams["figure.dpi"] = 180
plt.rcParams["savefig.dpi"] = 200
plt.style.use("bmh")
numpy : 1.20.2
xarray : 0.17.1.dev3+g48378c4b1
xfilter : 0.1.dev41+geb0277f
dcpy : 0.1
cf_xarray : 0.4.1.dev21+gab9dc66
matplotlib : 3.4.1
xgcm : 0.5.1
eddydiff : 0.1
pump : 0.1
xhistogram : 0.1.3+37.g7b706e8
pandas : 1.2.3
distributed: 2021.4.0
if "client" in locals():
client.close()
client = distributed.Client(n_workers=3, processes=False, memory_limit="8GB")
client
TIWKE from TAO#
tao = xr.open_zarr(
"/home/deepak/work/pump/notebooks/tao_eq_hr_gridded.zarr/", consolidated=True
).sel(longitude=-140, time=slice("1996", None))
vmean = (
tao.v.sel(depth=slice(-80, 0))
.mean("depth")
.chunk({"time": -1})
.interpolate_na("time", max_gap="10D")
.compute()
)
# Moum et al. 2009 metric
v = xfilter.bandpass(
vmean,
"time",
freq=[1 / 12, 1 / 33],
cycles_per="D",
debug=False,
num_discard=120,
)
tiwke = xfilter.lowpass(
v**2 / 2,
"time",
freq=1 / 20,
cycles_per="D",
)
tiwke.attrs = {"long_name": "TIW KE", "units": "m²/s²"}
tiwke = tiwke
/home/deepak/miniconda3/envs/dcpy/lib/python3.8/site-packages/dask/array/numpy_compat.py:39: RuntimeWarning: invalid value encountered in true_divide
x = np.divide(x1, x2, out)
Argo#
Read data#
ds = xr.load_dataset("../datasets/argo/tao.nc")
ds.coords["year"] = ds.TIME.dt.year
ds
<xarray.Dataset>
Dimensions: (N_POINTS: 1327818)
Coordinates:
LATITUDE (N_POINTS) float64 6.221 6.221 ... -1.509 -1.509
LONGITUDE (N_POINTS) float64 -140.0 -140.0 ... -138.7 -138.7
TIME (N_POINTS) datetime64[ns] 2005-03-08T17:40:05 ... ...
year (N_POINTS) int64 2005 2005 2005 ... 2020 2020 2020
Dimensions without coordinates: N_POINTS
Data variables: (12/13)
CONFIG_MISSION_NUMBER (N_POINTS) int64 1 1 1 1 1 1 1 ... 84 84 84 84 84 84
CYCLE_NUMBER (N_POINTS) int64 1 1 1 1 1 1 1 ... 84 84 84 84 84 84
DATA_MODE (N_POINTS) object 'D' 'D' 'D' 'D' ... 'A' 'A' 'A' 'A'
DIRECTION (N_POINTS) object 'A' 'A' 'A' 'A' ... 'A' 'A' 'A' 'A'
PLATFORM_NUMBER (N_POINTS) int64 5900884 5900884 ... 1902197 1902197
POSITION_QC (N_POINTS) int64 1 1 1 1 1 1 1 1 ... 1 1 1 1 1 1 1 1
... ...
PRES_QC (N_POINTS) int64 1 1 1 1 1 1 1 1 ... 1 1 1 1 1 1 1 1
PSAL (N_POINTS) float64 34.85 34.85 34.85 ... 34.62 34.62
PSAL_QC (N_POINTS) int64 1 1 1 1 1 1 1 1 ... 1 1 1 1 1 1 1 1
TEMP (N_POINTS) float64 27.97 27.97 27.97 ... 7.57 7.555
TEMP_QC (N_POINTS) int64 1 1 1 1 1 1 1 1 ... 1 1 1 1 1 1 1 1
TIME_QC (N_POINTS) int64 1 1 1 1 1 1 1 1 ... 1 1 1 1 1 1 1 1
Attributes:
DATA_ID: ARGO
DOI: http://doi.org/10.17882/42182
Fetched_from: https://www.ifremer.fr/erddap
Fetched_by: deepak
Fetched_date: 2021/04/08
Fetched_constraints: [x=-145.00/-135.00; y=-2.00/8.00; z=0.0/500.0; t=20...
Fetched_uri: https://www.ifremer.fr/erddap/tabledap/ArgoFloats.n...
history: Variables filtered according to DATA_MODE; Variable...- N_POINTS: 1327818
- LATITUDE(N_POINTS)float646.221 6.221 6.221 ... -1.509 -1.509
- _CoordinateAxisType :
- Lat
- actual_range :
- [7.03662268e-315 4.07900597e-320]
- axis :
- Y
- colorBarMaximum :
- 90.0
- colorBarMinimum :
- -90.0
- ioos_category :
- Location
- long_name :
- Latitude of the station, best estimate
- standard_name :
- latitude
- units :
- degrees_north
- valid_max :
- 90.0
- valid_min :
- -90.0
array([ 6.22100019, 6.22100019, 6.22100019, ..., -1.509 , -1.509 , -1.509 ]) - LONGITUDE(N_POINTS)float64-140.0 -140.0 ... -138.7 -138.7
- _CoordinateAxisType :
- Lon
- actual_range :
- [1.04862074e-317 2.03851486e-312]
- axis :
- X
- colorBarMaximum :
- 180.0
- colorBarMinimum :
- -180.0
- ioos_category :
- Location
- long_name :
- Longitude of the station, best estimate
- standard_name :
- longitude
- units :
- degrees_east
- valid_max :
- 180.0
- valid_min :
- -180.0
array([-140.03300476, -140.03300476, -140.03300476, ..., -138.663 , -138.663 , -138.663 ]) - TIME(N_POINTS)datetime64[ns]2005-03-08T17:40:05 ... 2020-12-...
- _CoordinateAxisType :
- Time
- actual_range :
- [6.97812303e-310 1.67192352e-312]
- axis :
- T
- ioos_category :
- Time
- long_name :
- Julian day (UTC) of the station relative to REFERENCE_DATE_TIME
- standard_name :
- time
- time_origin :
- 01-JAN-1970 00:00:00
array(['2005-03-08T17:40:05.000000000', '2005-03-08T17:40:05.000000000', '2005-03-08T17:40:05.000000000', ..., '2020-12-29T17:08:18.000000000', '2020-12-29T17:08:18.000000000', '2020-12-29T17:08:18.000000000'], dtype='datetime64[ns]') - year(N_POINTS)int642005 2005 2005 ... 2020 2020 2020
array([2005, 2005, 2005, ..., 2020, 2020, 2020])
- CONFIG_MISSION_NUMBER(N_POINTS)int641 1 1 1 1 1 1 ... 84 84 84 84 84 84
- actual_range :
- [ 16777216 452984832]
- colorBarMaximum :
- 100.0
- colorBarMinimum :
- 0.0
- conventions :
- 1...N, 1 : first complete mission
- ioos_category :
- Statistics
- long_name :
- Unique number denoting the missions performed by the float
- casted :
- 1
array([ 1, 1, 1, ..., 84, 84, 84])
- CYCLE_NUMBER(N_POINTS)int641 1 1 1 1 1 1 ... 84 84 84 84 84 84
- long_name :
- Float cycle number
- convention :
- 0..N, 0 : launch cycle (if exists), 1 : first complete cycle
- casted :
- 1
array([ 1, 1, 1, ..., 84, 84, 84])
- DATA_MODE(N_POINTS)object'D' 'D' 'D' 'D' ... 'A' 'A' 'A' 'A'
- long_name :
- Delayed mode or real time data
- convention :
- R : real time; D : delayed mode; A : real time with adjustment
- casted :
- 1
array(['D', 'D', 'D', ..., 'A', 'A', 'A'], dtype=object)
- DIRECTION(N_POINTS)object'A' 'A' 'A' 'A' ... 'A' 'A' 'A' 'A'
- long_name :
- Direction of the station profiles
- convention :
- A: ascending profiles, D: descending profiles
- casted :
- 1
array(['A', 'A', 'A', ..., 'A', 'A', 'A'], dtype=object)
- PLATFORM_NUMBER(N_POINTS)int645900884 5900884 ... 1902197 1902197
- long_name :
- Float unique identifier
- convention :
- WMO float identifier : A9IIIII
- casted :
- 1
array([5900884, 5900884, 5900884, ..., 1902197, 1902197, 1902197])
- POSITION_QC(N_POINTS)int641 1 1 1 1 1 1 1 ... 1 1 1 1 1 1 1 1
- long_name :
- Global quality flag of POSITION_QC profile
- convention :
- Argo reference table 2a
- casted :
- 1
array([1, 1, 1, ..., 1, 1, 1])
- PRES(N_POINTS)float644.3 6.9 11.0 ... 496.1 498.1 500.1
- long_name :
- Sea Pressure
- standard_name :
- sea_water_pressure
- units :
- decibar
- valid_min :
- 0.0
- valid_max :
- 12000.0
- resolution :
- 0.1
- axis :
- Z
- casted :
- 1
array([ 4.30000019, 6.9000001 , 11. , ..., 496.1000061 , 498.1000061 , 500.1000061 ]) - PRES_QC(N_POINTS)int641 1 1 1 1 1 1 1 ... 1 1 1 1 1 1 1 1
- long_name :
- Global quality flag of PRES_QC profile
- convention :
- Argo reference table 2a
- casted :
- 1
array([1, 1, 1, ..., 1, 1, 1])
- PSAL(N_POINTS)float6434.85 34.85 34.85 ... 34.62 34.62
- long_name :
- PRACTICAL SALINITY
- standard_name :
- sea_water_salinity
- units :
- psu
- valid_min :
- 0.0
- valid_max :
- 43.0
- resolution :
- 0.001
- casted :
- 1
array([34.84500122, 34.84500122, 34.84700012, ..., 34.61700058, 34.61600113, 34.61500168]) - PSAL_QC(N_POINTS)int641 1 1 1 1 1 1 1 ... 1 1 1 1 1 1 1 1
- long_name :
- Global quality flag of PSAL_QC profile
- convention :
- Argo reference table 2a
- casted :
- 1
array([1, 1, 1, ..., 1, 1, 1])
- TEMP(N_POINTS)float6427.97 27.97 27.97 ... 7.57 7.555
- long_name :
- SEA TEMPERATURE IN SITU ITS-90 SCALE
- standard_name :
- sea_water_temperature
- units :
- degree_Celsius
- valid_min :
- -2.0
- valid_max :
- 40.0
- resolution :
- 0.001
- casted :
- 1
array([27.96800041, 27.96699905, 27.96899986, ..., 7.59299994, 7.57000017, 7.55499983]) - TEMP_QC(N_POINTS)int641 1 1 1 1 1 1 1 ... 1 1 1 1 1 1 1 1
- long_name :
- Global quality flag of TEMP_QC profile
- convention :
- Argo reference table 2a
- casted :
- 1
array([1, 1, 1, ..., 1, 1, 1])
- TIME_QC(N_POINTS)int641 1 1 1 1 1 1 1 ... 1 1 1 1 1 1 1 1
- long_name :
- Global quality flag of TIME_QC profile
- convention :
- Argo reference table 2a
- casted :
- 1
array([1, 1, 1, ..., 1, 1, 1])
- DATA_ID :
- ARGO
- DOI :
- http://doi.org/10.17882/42182
- Fetched_from :
- https://www.ifremer.fr/erddap
- Fetched_by :
- deepak
- Fetched_date :
- 2021/04/08
- Fetched_constraints :
- [x=-145.00/-135.00; y=-2.00/8.00; z=0.0/500.0; t=2005-01-01/2010-12-31]
- Fetched_uri :
- https://www.ifremer.fr/erddap/tabledap/ArgoFloats.nc?data_mode,latitude,longitude,position_qc,time,time_qc,direction,platform_number,cycle_number,config_mission_number,vertical_sampling_scheme,pres,temp,psal,pres_qc,temp_qc,psal_qc,pres_adjusted,temp_adjusted,psal_adjusted,pres_adjusted_qc,temp_adjusted_qc,psal_adjusted_qc,pres_adjusted_error,temp_adjusted_error,psal_adjusted_error&longitude>=-145&longitude<=-135&latitude>=-2&latitude<=8&pres>=0&pres<=500&time>=1104537600.0&time<=1293753600.0&distinct()&orderBy("time,pres")
- history :
- Variables filtered according to DATA_MODE; Variables selected according to QC
Generate ENSO labels#
From Anna; Some Trenberth metric
Use NINO34 SST annomaly (w.r.t mean from 18xx)
At least 6 month long periods of SSTA > 0.4 (El-nino) & SSTA < -0.4 (La-Nina). Neutral everywhere else
from dcpy.util import block_lengths
from xarray.core.missing import _get_nan_block_lengths, get_clean_interp_index
nino34 = pump.obs.process_nino34()
ssta = nino34 - nino34.mean() # .rolling(time=6, center=True).mean()
enso = xr.full_like(ssta, fill_value="Neutral", dtype="U8")
index = ssta.indexes["time"] - ssta.indexes["time"][0]
en_mask = _get_nan_block_lengths(
xr.where(ssta > 0.4, np.nan, 0), dim="time", index=index
) >= pd.Timedelta("169d")
ln_mask = _get_nan_block_lengths(
xr.where(ssta < -0.4, np.nan, 0), dim="time", index=index
) >= pd.Timedelta("169d")
# neut_mask = _get_nan_block_lengths(xr.where((ssta < 0.5) & (ssta > -0.5), np.nan, 0), dim="time", index=index) >= pd.Timedelta("120d")
enso.loc[en_mask] = "El-Nino"
enso.loc[ln_mask] = "La-Nina"
# enso.loc[neut_mask] = "Neutral"
enso.name = "enso_phase"
enso.attrs[
"description"
] = "ENSO phase; El-Nino = NINO34 SSTA > 0.4 for at least 6 months; La-Nina = NINO34 SSTA < -0.4 for at least 6 months"
enso
<xarray.DataArray 'enso_phase' (time: 1812)>
array(['Neutral', 'Neutral', 'Neutral', ..., 'Neutral', 'Neutral',
'Neutral'], dtype='<U8')
Coordinates:
* time (time) datetime64[ns] 1870-01-31 1870-02-28 ... 2020-12-31
Attributes:
description: ENSO phase; El-Nino = NINO34 SSTA > 0.4 for at least 6 mont...- time: 1812
- 'Neutral' 'Neutral' 'Neutral' ... 'Neutral' 'Neutral' 'Neutral'
array(['Neutral', 'Neutral', 'Neutral', ..., 'Neutral', 'Neutral', 'Neutral'], dtype='<U8') - time(time)datetime64[ns]1870-01-31 ... 2020-12-31
array(['1870-01-31T00:00:00.000000000', '1870-02-28T00:00:00.000000000', '1870-03-31T00:00:00.000000000', ..., '2020-10-31T00:00:00.000000000', '2020-11-30T00:00:00.000000000', '2020-12-31T00:00:00.000000000'], dtype='datetime64[ns]')
- description :
- ENSO phase; El-Nino = NINO34 SSTA > 0.4 for at least 6 months; La-Nina = NINO34 SSTA < -0.4 for at least 6 months
Test against TIWKE
counts = (
ds.query({"N_POINTS": "PRES > 35 and PRES < 120"})
.TEMP.swap_dims({"N_POINTS": "TIME"})
.resample(TIME="M")
.count()
)
counts.attrs = {"long_name": "number of Argo samples", "units": "per month"}
from pump.plots import highlight_enso
ssta_ = dcpy.util.slice_like(ssta, tiwke)
enso_ = dcpy.util.slice_like(enso, tiwke)
f, axx = plt.subplots(3, 1, sharex=True, constrained_layout=True)
ssta_.plot(x="time", ax=axx[0])
highlight_enso(axx[0], enso_)
dcpy.plots.liney([-0.4, 0, 0.4], ax=axx[0], lw=0.3)
tiwke.plot(ax=axx[1], lw=1)
highlight_enso(axx[1], enso_)
counts.plot(ax=axx[2])
highlight_enso(axx[2], enso_.sel(time=slice("2005", None)))
dcpy.plots.clean_axes(np.atleast_2d(axx).T)
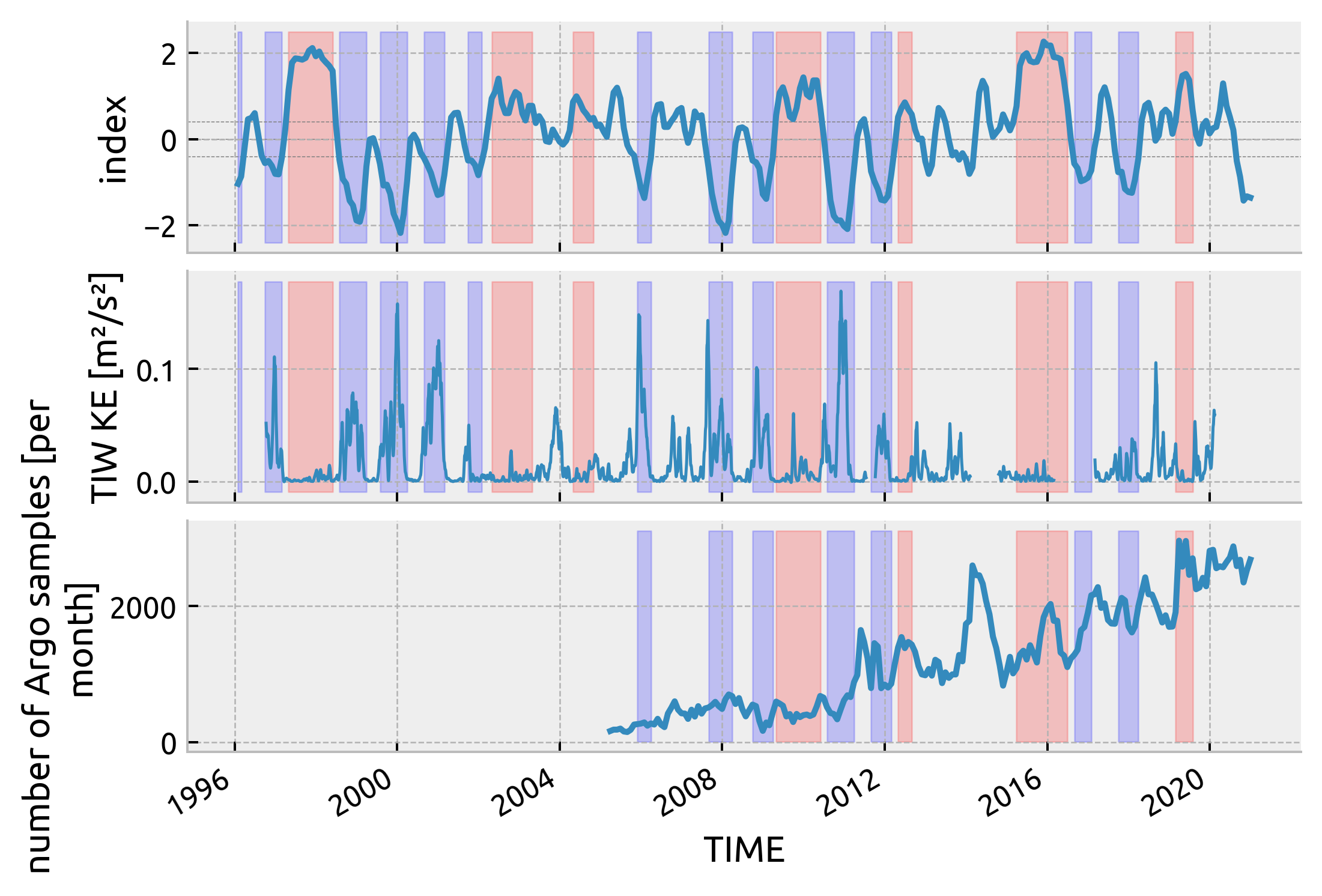
check ENSO phase calculation
f, ax = plt.subplots(2, 1, sharex=True)
(nino34 - nino34.mean()).sel(time=slice("2008", None)).plot(ax=ax[0])
ax[0].set_ylabel("SSTA")
ax[0].set_yticks([-0.4, 0, 0.4])
plt.plot(enso.time.sel(time=slice("2008", None)), enso.sel(time=slice("2008", None)))
[<matplotlib.lines.Line2D at 0x7fd20a9eac70>]
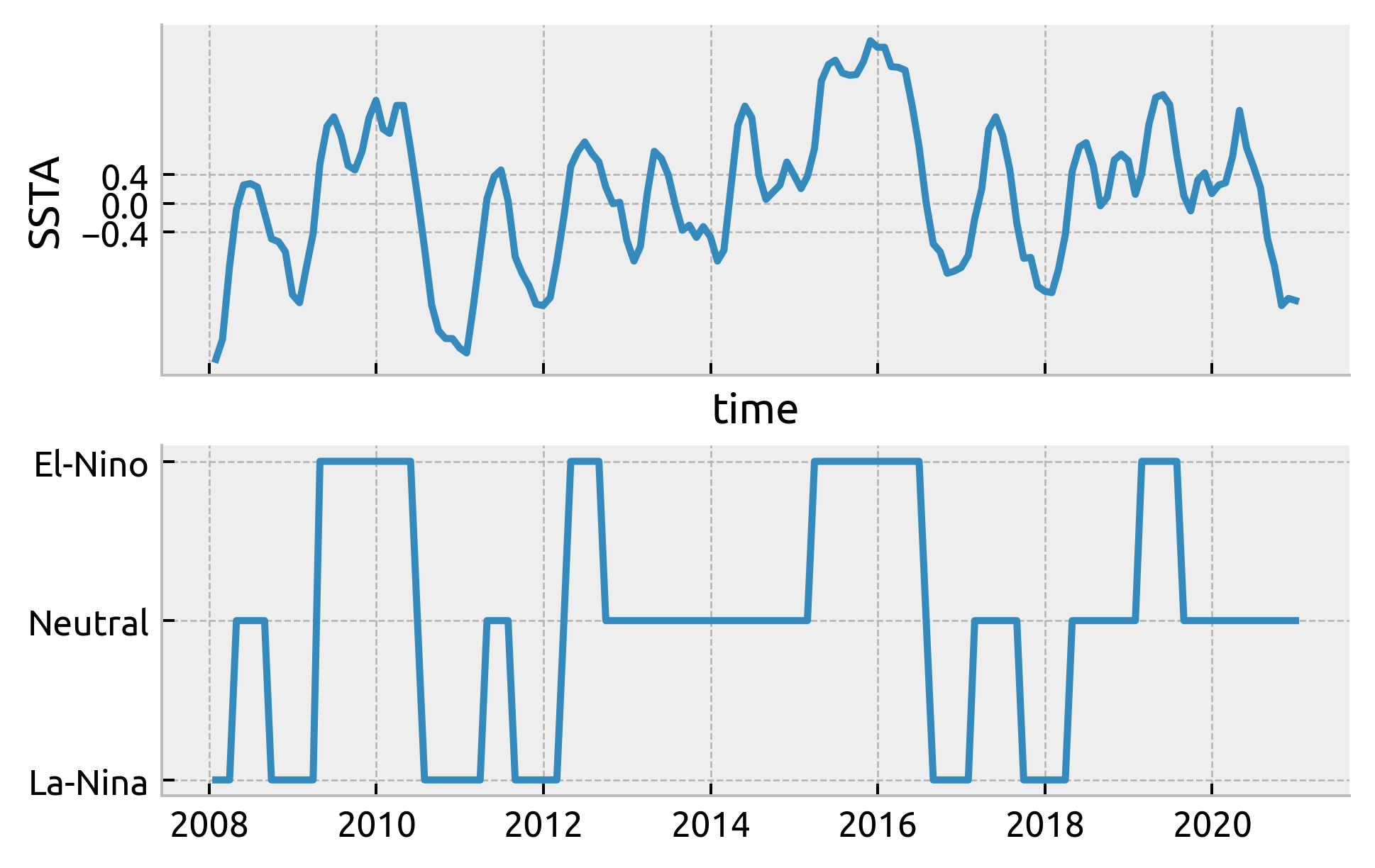
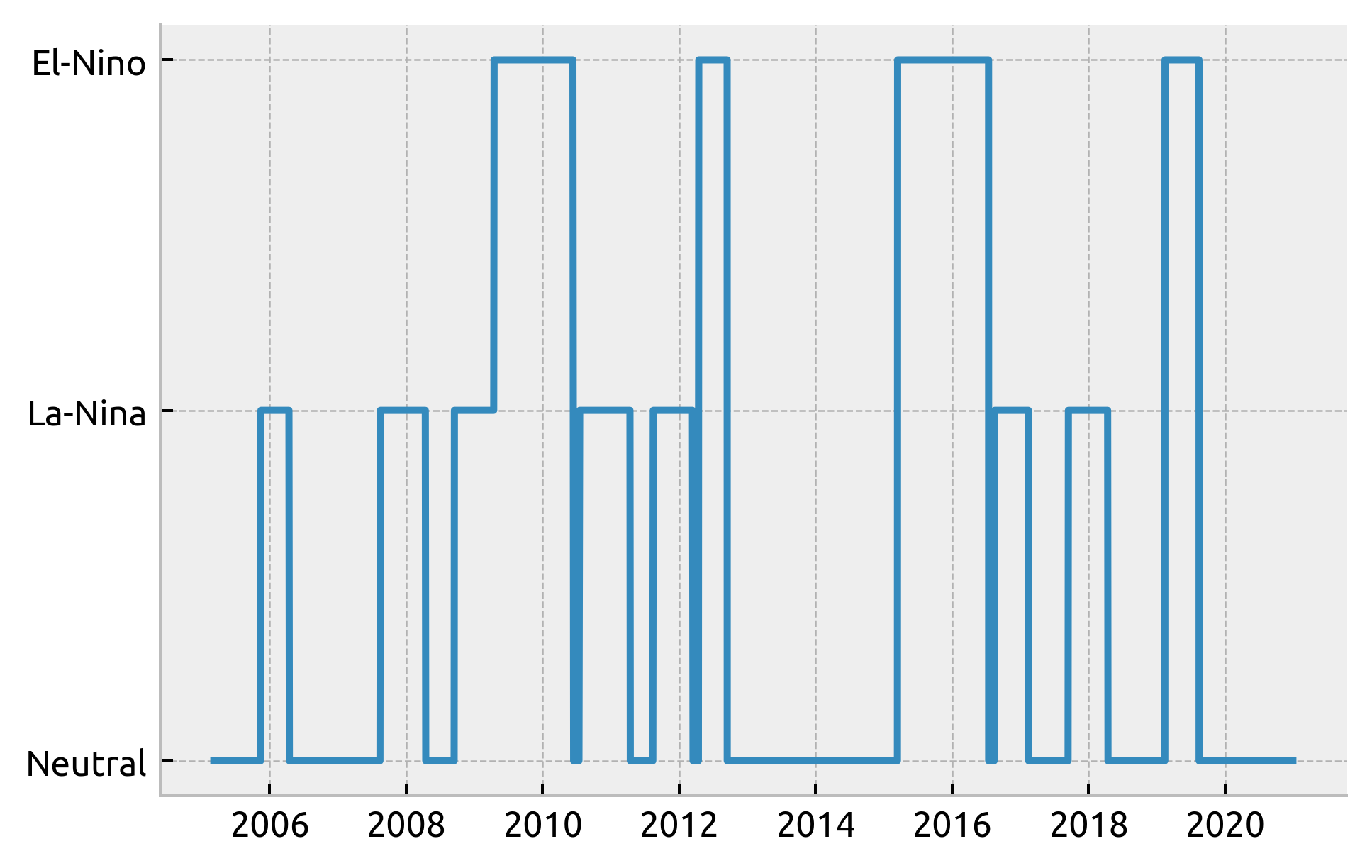
ds.coords["enso_phase"] = enso.sel(time=ds.TIME, method="nearest")
plt.plot(ds.TIME, ds.enso_phase)
[<matplotlib.lines.Line2D at 0x7fd4c8612d60>]
Latitudinal variation at 140W#
subset = ds.query({"N_POINTS": "LATITUDE > -2 & LATITUDE < 2"})
_, ax = dcpy.oceans.TSplot(subset.PSAL, subset.TEMP, hexbin=False)
subset = ds.query({"N_POINTS": "LATITUDE > 2 & LATITUDE < 4"})
dcpy.oceans.TSplot(subset.PSAL, subset.TEMP, hexbin=False, ax=ax)
subset = ds.query({"N_POINTS": "LATITUDE > 4 & LATITUDE < 6"})
dcpy.oceans.TSplot(subset.PSAL, subset.TEMP, hexbin=False, ax=ax)
ax["s"].legend(["-2->2", "2->4", "4->6"])
ax["s"].set_title("2005-2020")
Text(0.5, 1.0, '2005-2020')
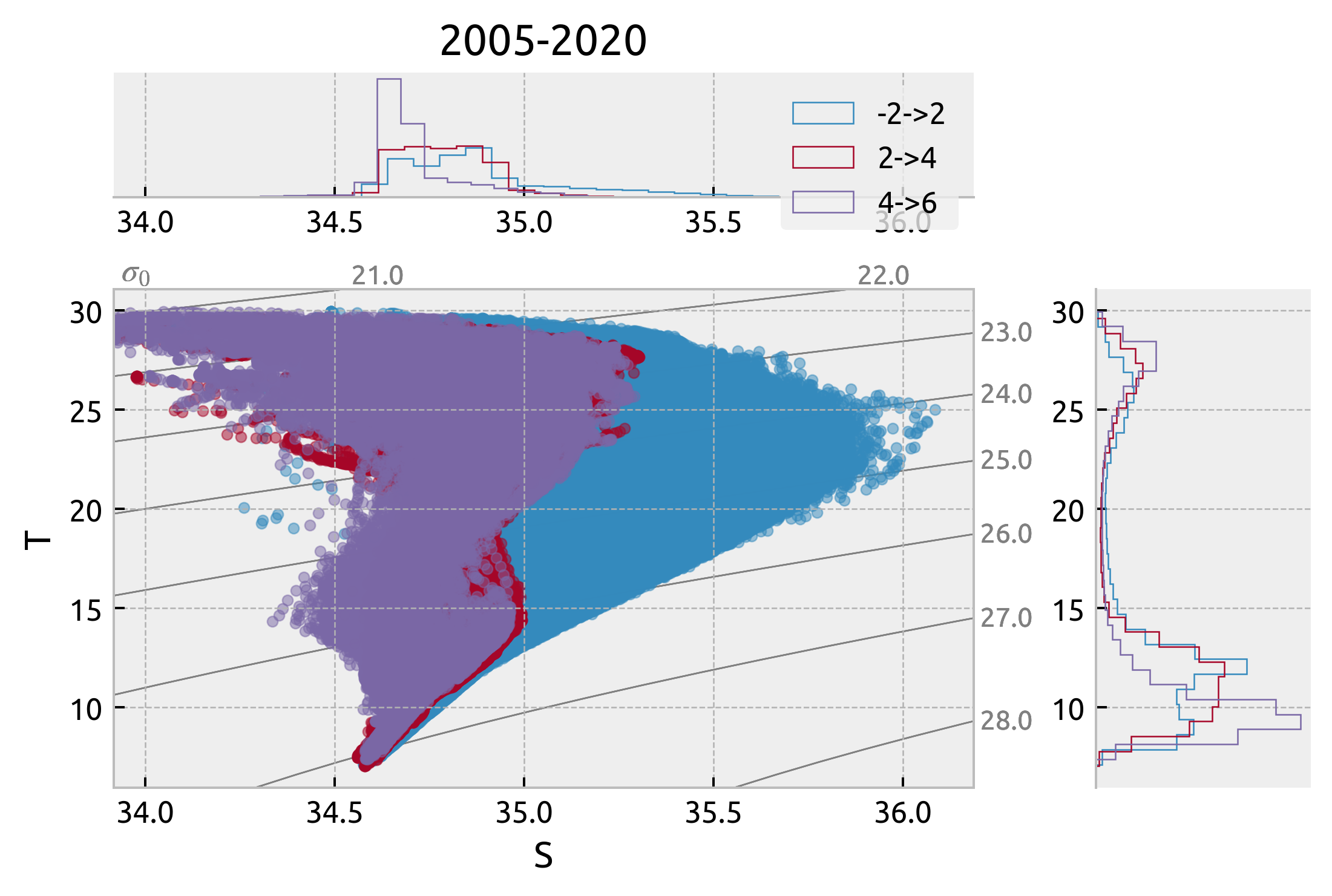
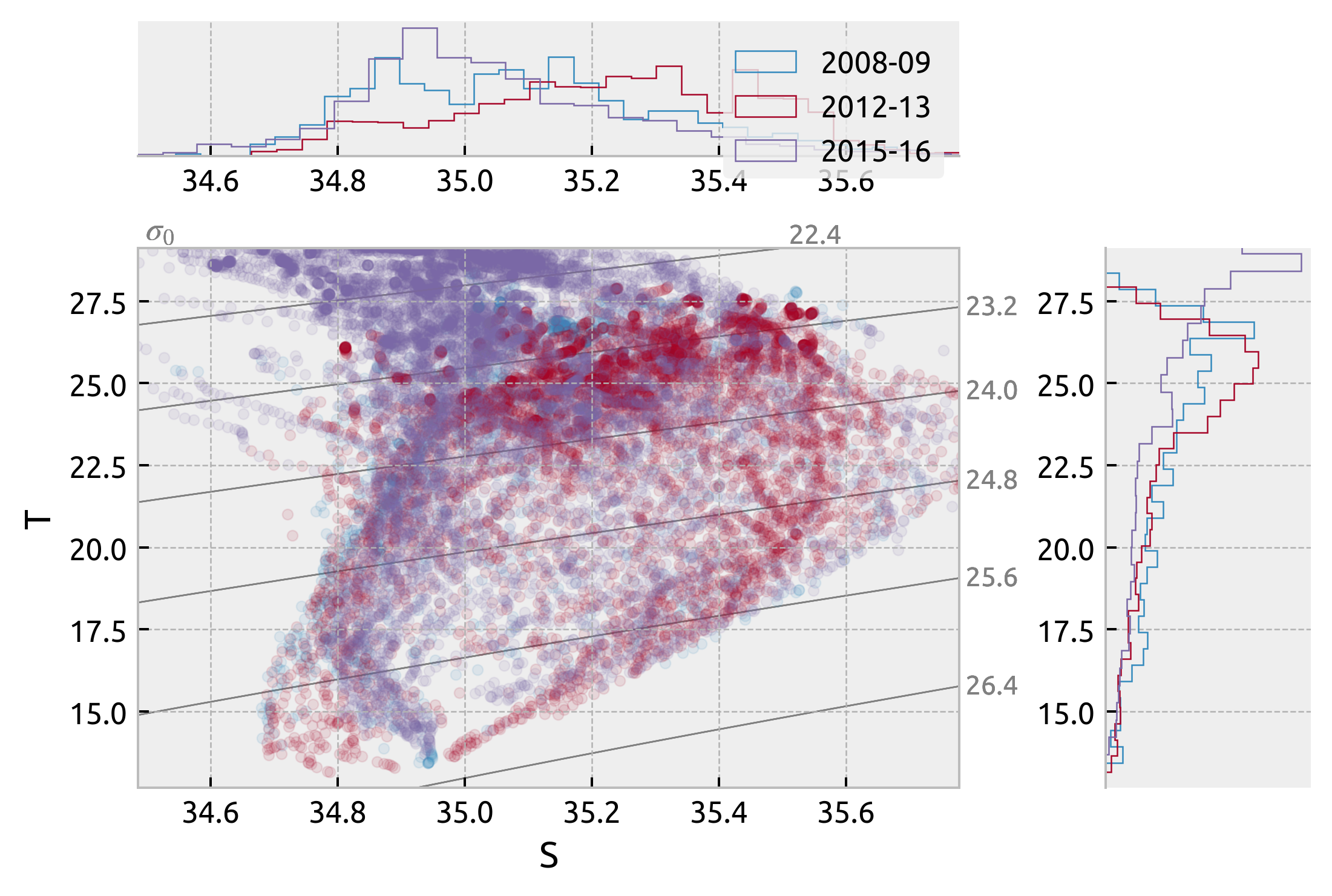
Interannual variation at (2S→2N, -140)#
Picked out two La-Nina(2008, 09; 2012, 13) and one El-Nino period (2015, 2016). They look very similar
kwargs = dict(kind="scatter", plot_kwargs=dict(alpha=0.1))
common = "LONGITUDE > -142 & LONGITUDE < -138 & LATITUDE > -2 & LATITUDE < 2 & PRES > 35 & PRES < 120"
subset = ds.query({"N_POINTS": f"{common} & (year == 2008 | year == 2009)"})
_, ax = dcpy.oceans.TSplot(subset.PSAL, subset.TEMP, **kwargs)
subset = ds.query({"N_POINTS": f"{common} & (year == 2012 | year == 2013)"})
dcpy.oceans.TSplot(subset.PSAL, subset.TEMP, **kwargs, ax=ax)
subset = ds.query({"N_POINTS": f"{common} & (year == 2015 | year == 2016)"})
dcpy.oceans.TSplot(subset.PSAL, subset.TEMP, **kwargs, ax=ax)
ax["s"].legend(["2008-09", "2012-13", "2015-16"])
<matplotlib.legend.Legend at 0x7fd1eab81970>
distributed.utils_perf - WARNING - full garbage collections took 10% CPU time recently (threshold: 10%)
distributed.utils_perf - WARNING - full garbage collections took 10% CPU time recently (threshold: 10%)
T-S diagram variation with ENSO phase#
This is a TS diagram of Argo profiles between 135W→145W, 2S→2N; grouped by ENSO phase.
El-Nino:
isopycnal scatter at warmer isotherms during El-Ninos (top 100m). Where is that coming from.
Below ≈ 100m the TS relationship is tight with two end-members? Some of this is a function of spatial extent.
La-Nina:
Similar scatter to El-Nino which is surprising
Neutral:
Lots of scatter! But also lots more points.
higher salinity water appears
could add depth contours.
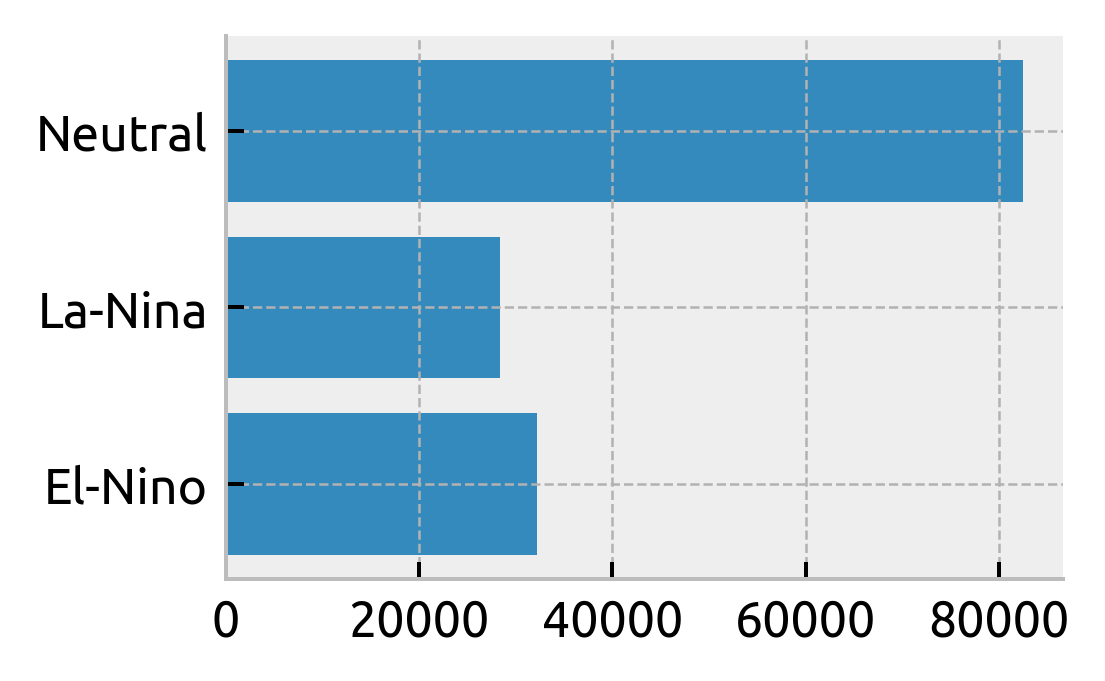
counts = (
ds.query({"N_POINTS": f"LATITUDE > -2 & LATITUDE < 2 & PRES > 35 & PRES < 120"})
.groupby("enso_phase")
.count()
)
plt.barh(counts.enso_phase, counts.TEMP)
plt.gcf().set_size_inches((3, 2))
def plot_TS(latitude_string):
"""Plot a TS diagram per ENSO phase"""
from matplotlib.lines import Line2D
kwargs = dict(
hexbin=True,
plot_kwargs={"alpha": 0.3, "mincnt": 10, "norm": mpl.colors.LogNorm(1, 100)},
)
# kwargs = dict(kind="scatter", plot_kwargs={"alpha": 0.1})
kwargs = dict(kind="hist", equalize=False, plot_kwargs={"alpha": 0.6})
ax = None
labels = []
lines = []
for phase, cmap, color in [
["El-Nino", mpl.cm.Reds, "red"],
# ["Neutral", mpl.cm.Greys, "black"],
["La-Nina", mpl.cm.Blues, "blue"],
]:
subset = ds.query(
{
"N_POINTS": f"{latitude_string} & enso_phase == {phase!r} & PRES > 35 & PRES < 100"
}
)
kwargs["plot_kwargs"]["cmap"] = cmap
if ax is not None:
kwargs.update({"label_spines": False, "rho_levels": None})
hdl, ax = dcpy.oceans.TSplot(
subset.PSAL, subset.TEMP, color=color, **kwargs, ax=ax
)
lines.append(Line2D([0], [0], color=color, lw=1))
labels.append(phase)
ax["ts"].set_ylim([None, 31])
ax["t"].get_figure().legend(handles=lines, labels=labels, loc="upper right")
ax["s"].set_title(latitude_string)
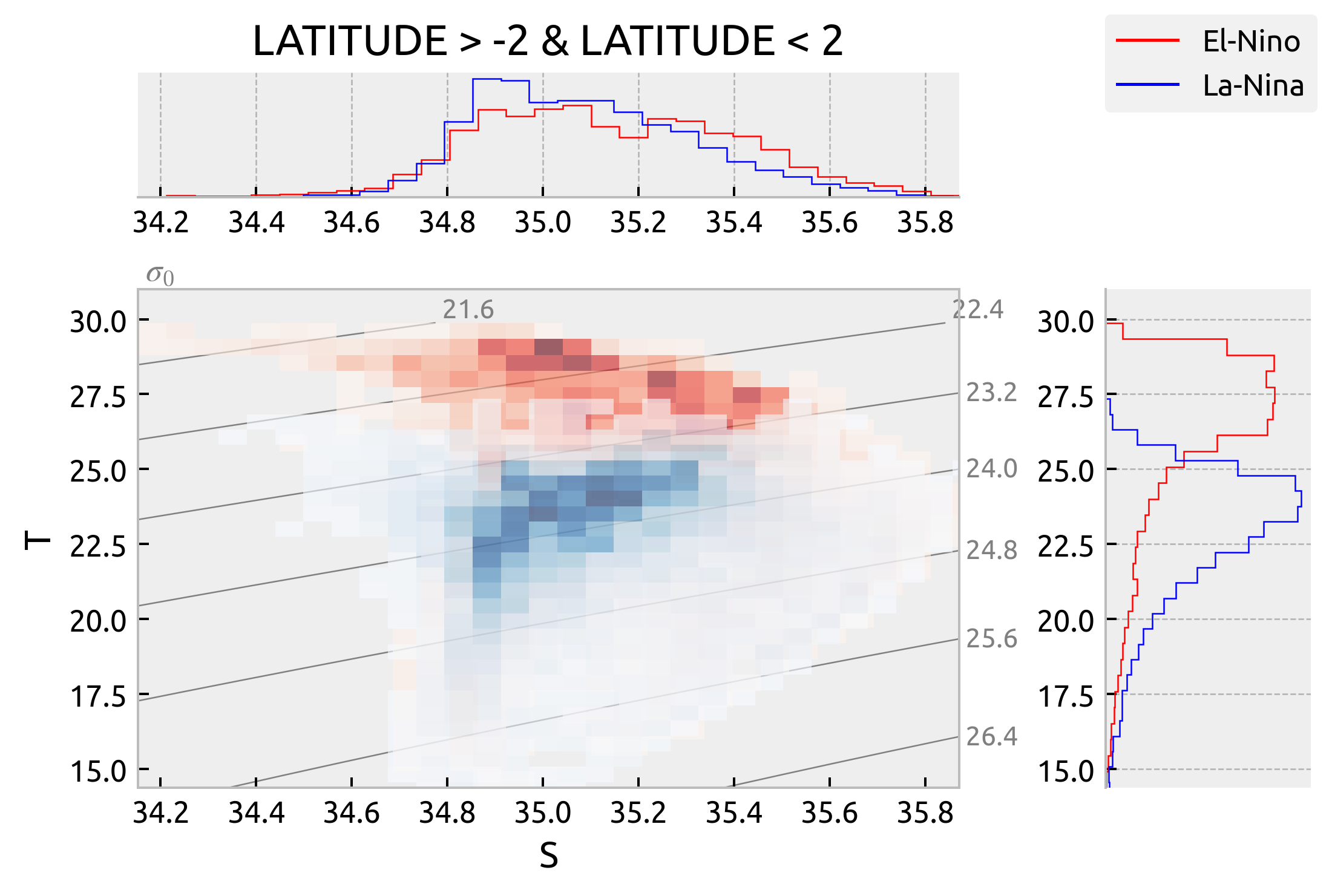
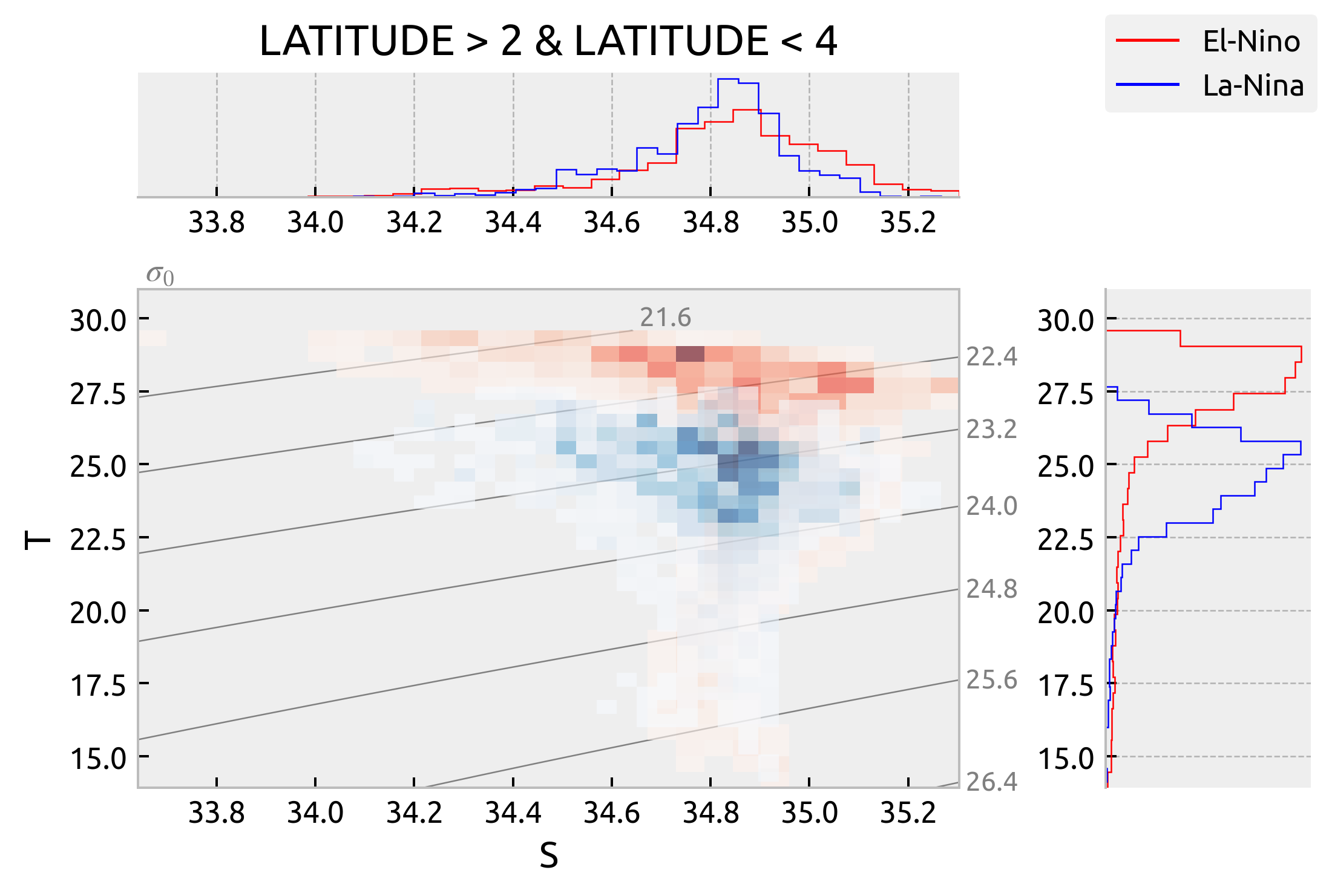
It looks like there is latitudinal variation in T-S spread along isopycnals but not really an ENSO phase variation. The La-Nina and Neutral TS diagrams basically overlap but we can see that the La-Nina and El-Nino diagrams are really only different in that the El-Nino one is warmer. The spread looks the same.
plot distributions along 24.5 isopycnal
plot_TS(latitude_string="LATITUDE > -2 & LATITUDE < 2")
plot_TS(latitude_string="LATITUDE > 2 & LATITUDE < 4")
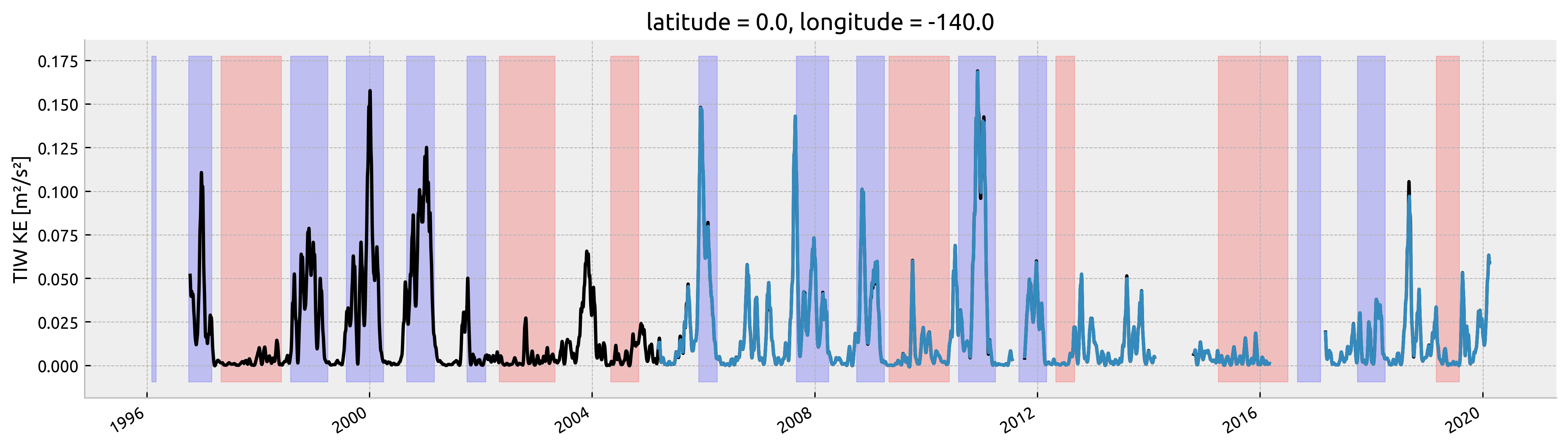
Variation of T-S with TIWKE at (0, 140W)#
Looks the same even for weak TIWs.
This won’t really recover the La-Nina/El-Nino plot because you can have periods of weak TIW KE during La-Nina conditions
TODO: I think I might need to set TIWKE for a profile as (mean TIWKE within a 5-10 day window)?
ds["tiwke"] = tiwke.sel(time=ds.TIME, method="pad")
da = tiwke.copy(deep=True)
tiwke.plot(x="time", color="k", size=4, aspect=4)
ds.tiwke.plot(x="TIME")
highlight_enso(ax=plt.gca(), enso=enso.sel(time=slice("1996", None)))
plt.gca().set_xlabel("")
Text(0.5, 0, '')
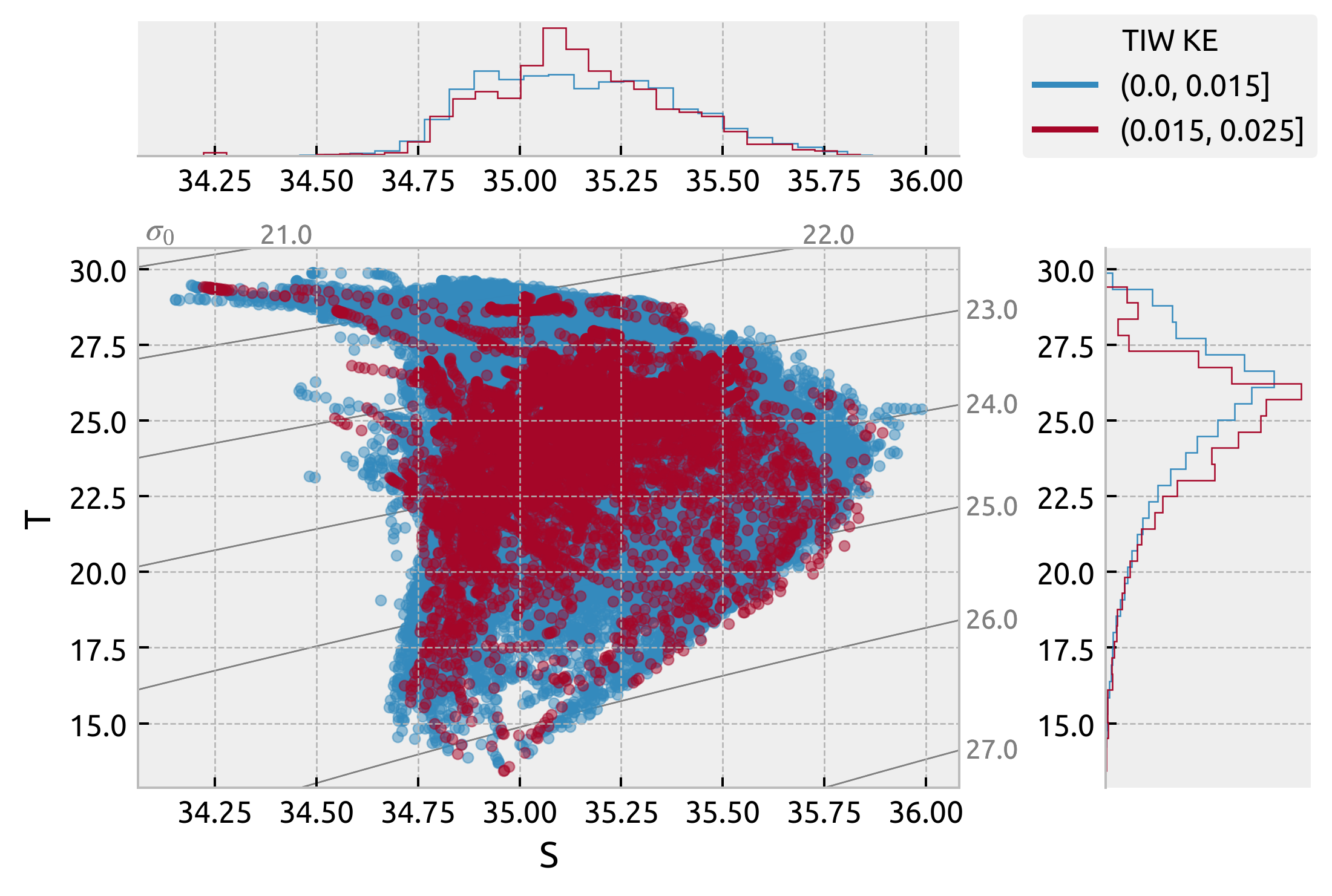
tiwkegrouped = ds.groupby_bins("tiwke", bins=[0, 0.015, 0.025])
from matplotlib.lines import Line2D
ax = None
labels = []
lines = []
for label, group in tiwkegrouped:
group = group.query(
{"N_POINTS": "LATITUDE > -2 & LATITUDE < 2 & PRES > 35 & PRES < 100"}
)
hdl, axxx = dcpy.oceans.TSplot(group.PSAL, group.TEMP, kind="scatter", ax=ax)
if ax is None:
ax = axxx
lines.append(Line2D([0], [0], color=hdl["Thist"][-1][0].get_edgecolor()))
labels.append(label)
ax["s"].get_figure().legend(lines, labels, title="TIW KE", loc="upper right")
<matplotlib.legend.Legend at 0x7fd46c679490>
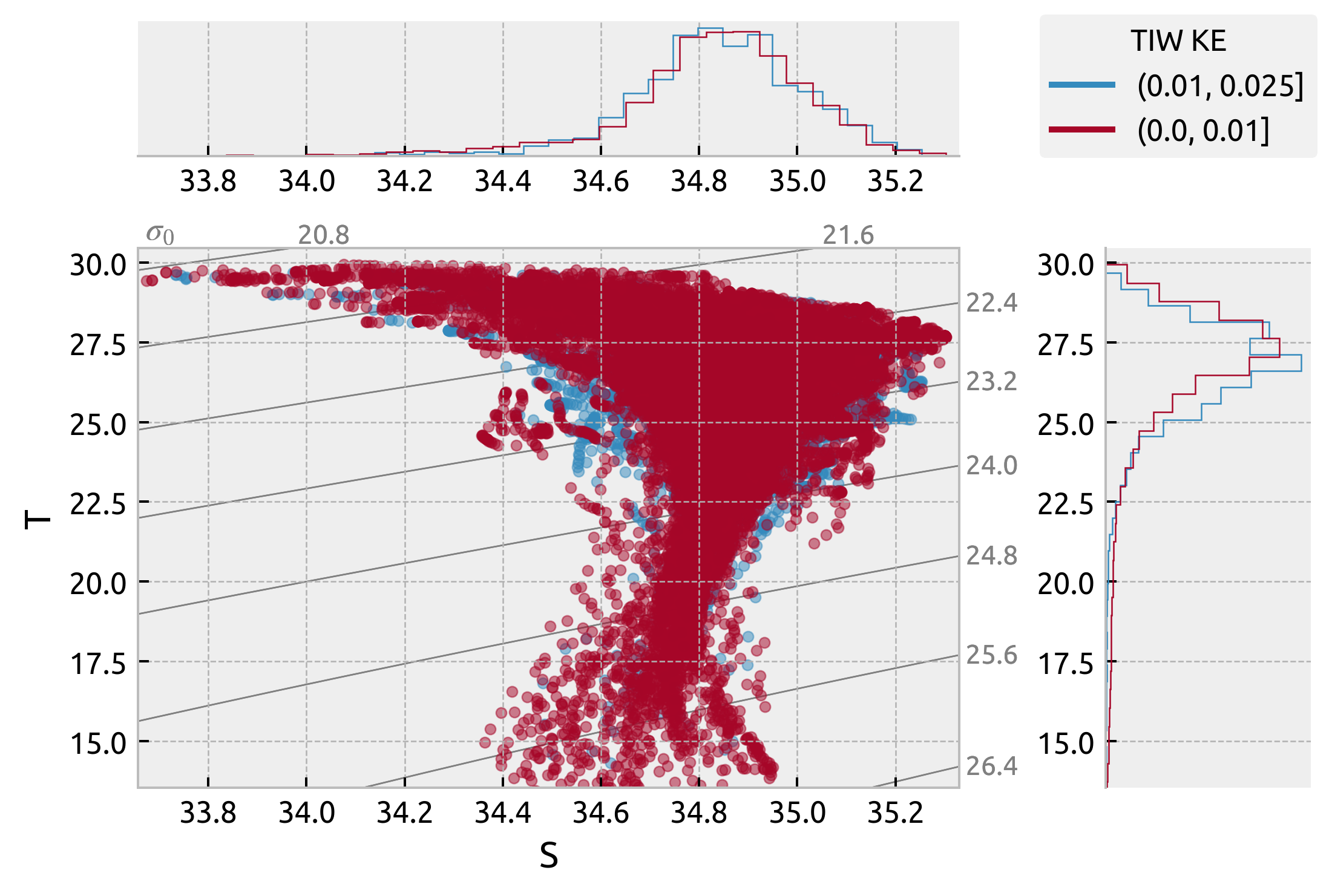
from matplotlib.lines import Line2D
ax = None
labels = []
lines = []
for label, group in tiwkegrouped:
group = group.query(
{"N_POINTS": "LATITUDE > 2 & LATITUDE < 6 & PRES > 35 & PRES < 100"}
)
hdl, axxx = dcpy.oceans.TSplot(group.PSAL, group.TEMP, kind="scatter", ax=ax)
if ax is None:
ax = axxx
lines.append(Line2D([0], [0], color=hdl["Thist"][-1][0].get_edgecolor()))
labels.append(label)
ax["s"].get_figure().legend(lines, labels, title="TIW KE", loc="upper right")
<matplotlib.legend.Legend at 0x7fd46c7bd6d0>
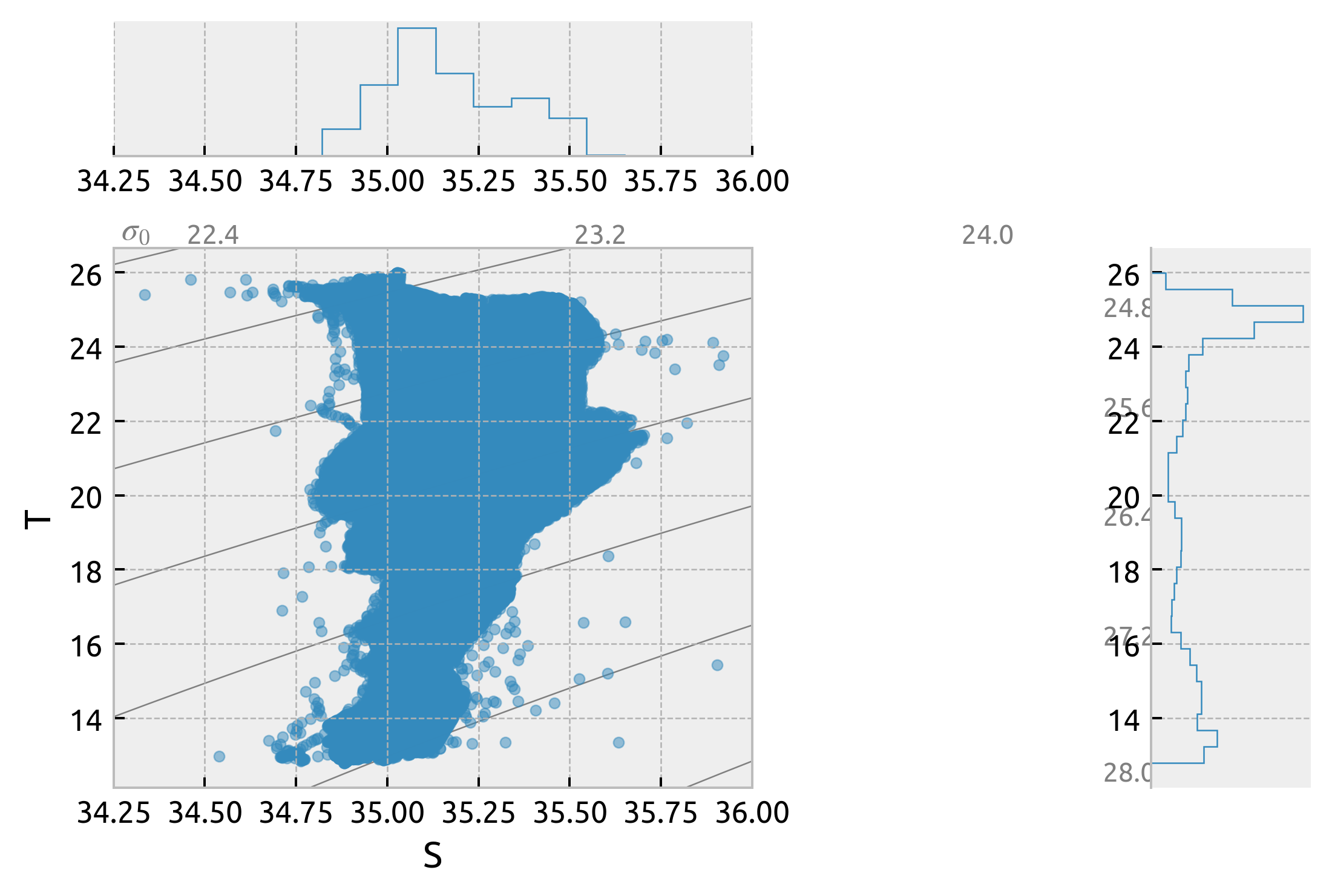
equix = xr.open_dataset("/home/deepak/datasets/microstructure/osu/equix.nc")
_, ax = dcpy.oceans.TSplot(equix.salt, equix.theta, hexbin=False)
ax["ts"].set_xlim([34.25, 36])
(34.25, 36.0)
argoclim = dcpy.oceans.read_argo_clim()
argoclim
<xarray.Dataset>
Dimensions: (lat: 145, lon: 360, pres: 58, time: 180)
Coordinates:
* lon (lon) float32 20.5 21.5 22.5 23.5 ... 377.5 378.5 379.5
* lat (lat) float32 -64.5 -63.5 -62.5 -61.5 ... 77.5 78.5 79.5
* pres (pres) float32 2.5 10.0 20.0 ... 1.8e+03 1.9e+03 1.975e+03
* time (time) datetime64[ns] 2004-01-16 2004-02-15 ... 2018-09-29
Data variables:
Tmean (pres, lat, lon) float32 dask.array<chunksize=(58, 20, 60), meta=np.ndarray>
Tanom (time, pres, lat, lon) float32 dask.array<chunksize=(180, 58, 20, 60), meta=np.ndarray>
BATHYMETRY_MASK (pres, lat, lon) float32 dask.array<chunksize=(58, 20, 60), meta=np.ndarray>
MAPPING_MASK (pres, lat, lon) float32 dask.array<chunksize=(58, 20, 60), meta=np.ndarray>
T (time, pres, lat, lon) float32 dask.array<chunksize=(180, 58, 20, 60), meta=np.ndarray>
Smean (pres, lat, lon) float32 dask.array<chunksize=(58, 20, 60), meta=np.ndarray>
Sanom (time, pres, lat, lon) float32 dask.array<chunksize=(180, 58, 20, 60), meta=np.ndarray>
S (time, pres, lat, lon) float32 dask.array<chunksize=(180, 58, 20, 60), meta=np.ndarray>- lat: 145
- lon: 360
- pres: 58
- time: 180
- lon(lon)float3220.5 21.5 22.5 ... 378.5 379.5
- units :
- degrees_east
- modulo :
- 360.0
- point_spacing :
- even
- axis :
- X
array([ 20.5, 21.5, 22.5, ..., 377.5, 378.5, 379.5], dtype=float32)
- lat(lat)float32-64.5 -63.5 -62.5 ... 78.5 79.5
- units :
- degrees_north
- point_spacing :
- even
- axis :
- Y
array([-64.5, -63.5, -62.5, -61.5, -60.5, -59.5, -58.5, -57.5, -56.5, -55.5, -54.5, -53.5, -52.5, -51.5, -50.5, -49.5, -48.5, -47.5, -46.5, -45.5, -44.5, -43.5, -42.5, -41.5, -40.5, -39.5, -38.5, -37.5, -36.5, -35.5, -34.5, -33.5, -32.5, -31.5, -30.5, -29.5, -28.5, -27.5, -26.5, -25.5, -24.5, -23.5, -22.5, -21.5, -20.5, -19.5, -18.5, -17.5, -16.5, -15.5, -14.5, -13.5, -12.5, -11.5, -10.5, -9.5, -8.5, -7.5, -6.5, -5.5, -4.5, -3.5, -2.5, -1.5, -0.5, 0.5, 1.5, 2.5, 3.5, 4.5, 5.5, 6.5, 7.5, 8.5, 9.5, 10.5, 11.5, 12.5, 13.5, 14.5, 15.5, 16.5, 17.5, 18.5, 19.5, 20.5, 21.5, 22.5, 23.5, 24.5, 25.5, 26.5, 27.5, 28.5, 29.5, 30.5, 31.5, 32.5, 33.5, 34.5, 35.5, 36.5, 37.5, 38.5, 39.5, 40.5, 41.5, 42.5, 43.5, 44.5, 45.5, 46.5, 47.5, 48.5, 49.5, 50.5, 51.5, 52.5, 53.5, 54.5, 55.5, 56.5, 57.5, 58.5, 59.5, 60.5, 61.5, 62.5, 63.5, 64.5, 65.5, 66.5, 67.5, 68.5, 69.5, 70.5, 71.5, 72.5, 73.5, 74.5, 75.5, 76.5, 77.5, 78.5, 79.5], dtype=float32) - pres(pres)float322.5 10.0 20.0 ... 1.9e+03 1.975e+03
- units :
- dbar
- positive :
- down
- point_spacing :
- uneven
- axis :
- Z
array([ 2.5, 10. , 20. , 30. , 40. , 50. , 60. , 70. , 80. , 90. , 100. , 110. , 120. , 130. , 140. , 150. , 160. , 170. , 182.5, 200. , 220. , 240. , 260. , 280. , 300. , 320. , 340. , 360. , 380. , 400. , 420. , 440. , 462.5, 500. , 550. , 600. , 650. , 700. , 750. , 800. , 850. , 900. , 950. , 1000. , 1050. , 1100. , 1150. , 1200. , 1250. , 1300. , 1350. , 1412.5, 1500. , 1600. , 1700. , 1800. , 1900. , 1975. ], dtype=float32) - time(time)datetime64[ns]2004-01-16 ... 2018-09-29
- axis :
- T
array(['2004-01-16T00:00:00.000000000', '2004-02-15T00:00:00.000000000', '2004-03-16T00:00:00.000000000', '2004-04-15T00:00:00.000000000', '2004-05-15T00:00:00.000000000', '2004-06-14T00:00:00.000000000', '2004-07-14T00:00:00.000000000', '2004-08-13T00:00:00.000000000', '2004-09-12T00:00:00.000000000', '2004-10-12T00:00:00.000000000', '2004-11-11T00:00:00.000000000', '2004-12-11T00:00:00.000000000', '2005-01-10T00:00:00.000000000', '2005-02-09T00:00:00.000000000', '2005-03-11T00:00:00.000000000', '2005-04-10T00:00:00.000000000', '2005-05-10T00:00:00.000000000', '2005-06-09T00:00:00.000000000', '2005-07-09T00:00:00.000000000', '2005-08-08T00:00:00.000000000', '2005-09-07T00:00:00.000000000', '2005-10-07T00:00:00.000000000', '2005-11-06T00:00:00.000000000', '2005-12-06T00:00:00.000000000', '2006-01-05T00:00:00.000000000', '2006-02-04T00:00:00.000000000', '2006-03-06T00:00:00.000000000', '2006-04-05T00:00:00.000000000', '2006-05-05T00:00:00.000000000', '2006-06-04T00:00:00.000000000', '2006-07-04T00:00:00.000000000', '2006-08-03T00:00:00.000000000', '2006-09-02T00:00:00.000000000', '2006-10-02T00:00:00.000000000', '2006-11-01T00:00:00.000000000', '2006-12-01T00:00:00.000000000', '2006-12-31T00:00:00.000000000', '2007-01-30T00:00:00.000000000', '2007-03-01T00:00:00.000000000', '2007-03-31T00:00:00.000000000', '2007-04-30T00:00:00.000000000', '2007-05-30T00:00:00.000000000', '2007-06-29T00:00:00.000000000', '2007-07-29T00:00:00.000000000', '2007-08-28T00:00:00.000000000', '2007-09-27T00:00:00.000000000', '2007-10-27T00:00:00.000000000', '2007-11-26T00:00:00.000000000', '2007-12-26T00:00:00.000000000', '2008-01-25T00:00:00.000000000', '2008-02-24T00:00:00.000000000', '2008-03-25T00:00:00.000000000', '2008-04-24T00:00:00.000000000', '2008-05-24T00:00:00.000000000', '2008-06-23T00:00:00.000000000', '2008-07-23T00:00:00.000000000', '2008-08-22T00:00:00.000000000', '2008-09-21T00:00:00.000000000', '2008-10-21T00:00:00.000000000', '2008-11-20T00:00:00.000000000', '2008-12-20T00:00:00.000000000', '2009-01-19T00:00:00.000000000', '2009-02-18T00:00:00.000000000', '2009-03-20T00:00:00.000000000', '2009-04-19T00:00:00.000000000', '2009-05-19T00:00:00.000000000', '2009-06-18T00:00:00.000000000', '2009-07-18T00:00:00.000000000', '2009-08-17T00:00:00.000000000', '2009-09-16T00:00:00.000000000', '2009-10-16T00:00:00.000000000', '2009-11-15T00:00:00.000000000', '2009-12-15T00:00:00.000000000', '2010-01-14T00:00:00.000000000', '2010-02-13T00:00:00.000000000', '2010-03-15T00:00:00.000000000', '2010-04-14T00:00:00.000000000', '2010-05-14T00:00:00.000000000', '2010-06-13T00:00:00.000000000', '2010-07-13T00:00:00.000000000', '2010-08-12T00:00:00.000000000', '2010-09-11T00:00:00.000000000', '2010-10-11T00:00:00.000000000', '2010-11-10T00:00:00.000000000', '2010-12-10T00:00:00.000000000', '2011-01-09T00:00:00.000000000', '2011-02-08T00:00:00.000000000', '2011-03-10T00:00:00.000000000', '2011-04-09T00:00:00.000000000', '2011-05-09T00:00:00.000000000', '2011-06-08T00:00:00.000000000', '2011-07-08T00:00:00.000000000', '2011-08-07T00:00:00.000000000', '2011-09-06T00:00:00.000000000', '2011-10-06T00:00:00.000000000', '2011-11-05T00:00:00.000000000', '2011-12-05T00:00:00.000000000', '2012-01-04T00:00:00.000000000', '2012-02-03T00:00:00.000000000', '2012-03-04T00:00:00.000000000', '2012-04-03T00:00:00.000000000', '2012-05-03T00:00:00.000000000', '2012-06-02T00:00:00.000000000', '2012-07-02T00:00:00.000000000', '2012-08-01T00:00:00.000000000', '2012-08-31T00:00:00.000000000', '2012-09-30T00:00:00.000000000', '2012-10-30T00:00:00.000000000', '2012-11-29T00:00:00.000000000', '2012-12-29T00:00:00.000000000', '2013-01-28T00:00:00.000000000', '2013-02-27T00:00:00.000000000', '2013-03-29T00:00:00.000000000', '2013-04-28T00:00:00.000000000', '2013-05-28T00:00:00.000000000', '2013-06-27T00:00:00.000000000', '2013-07-27T00:00:00.000000000', '2013-08-26T00:00:00.000000000', '2013-09-25T00:00:00.000000000', '2013-10-25T00:00:00.000000000', '2013-11-24T00:00:00.000000000', '2013-12-24T00:00:00.000000000', '2014-01-23T00:00:00.000000000', '2014-02-22T00:00:00.000000000', '2014-03-24T00:00:00.000000000', '2014-04-23T00:00:00.000000000', '2014-05-23T00:00:00.000000000', '2014-06-22T00:00:00.000000000', '2014-07-22T00:00:00.000000000', '2014-08-21T00:00:00.000000000', '2014-09-20T00:00:00.000000000', '2014-10-20T00:00:00.000000000', '2014-11-19T00:00:00.000000000', '2014-12-19T00:00:00.000000000', '2015-01-18T00:00:00.000000000', '2015-02-17T00:00:00.000000000', '2015-03-19T00:00:00.000000000', '2015-04-18T00:00:00.000000000', '2015-05-18T00:00:00.000000000', '2015-06-17T00:00:00.000000000', '2015-07-17T00:00:00.000000000', '2015-08-16T00:00:00.000000000', '2015-09-15T00:00:00.000000000', '2015-10-15T00:00:00.000000000', '2015-11-14T00:00:00.000000000', '2015-12-14T00:00:00.000000000', '2016-01-13T00:00:00.000000000', '2016-02-12T00:00:00.000000000', '2016-03-13T00:00:00.000000000', '2016-04-12T00:00:00.000000000', '2016-05-12T00:00:00.000000000', '2016-06-11T00:00:00.000000000', '2016-07-11T00:00:00.000000000', '2016-08-10T00:00:00.000000000', '2016-09-09T00:00:00.000000000', '2016-10-09T00:00:00.000000000', '2016-11-08T00:00:00.000000000', '2016-12-08T00:00:00.000000000', '2017-01-07T00:00:00.000000000', '2017-02-06T00:00:00.000000000', '2017-03-08T00:00:00.000000000', '2017-04-07T00:00:00.000000000', '2017-05-07T00:00:00.000000000', '2017-06-06T00:00:00.000000000', '2017-07-06T00:00:00.000000000', '2017-08-05T00:00:00.000000000', '2017-09-04T00:00:00.000000000', '2017-10-04T00:00:00.000000000', '2017-11-03T00:00:00.000000000', '2017-12-03T00:00:00.000000000', '2018-01-02T00:00:00.000000000', '2018-02-01T00:00:00.000000000', '2018-03-03T00:00:00.000000000', '2018-04-02T00:00:00.000000000', '2018-05-02T00:00:00.000000000', '2018-06-01T00:00:00.000000000', '2018-07-01T00:00:00.000000000', '2018-07-31T00:00:00.000000000', '2018-08-30T00:00:00.000000000', '2018-09-29T00:00:00.000000000'], dtype='datetime64[ns]')
- Tmean(pres, lat, lon)float32dask.array<chunksize=(58, 20, 60), meta=np.ndarray>
- units :
- degree celcius (ITS-90)
- long_name :
- ARGO TEMPERATURE MEAN Jan 2004 - Dec 2018 (15.0 year) RG CLIMATOLOGY
Array Chunk Bytes 11.55 MiB 271.88 kiB Shape (58, 145, 360) (58, 20, 60) Count 49 Tasks 48 Chunks Type float32 numpy.ndarray - Tanom(time, pres, lat, lon)float32dask.array<chunksize=(180, 58, 20, 60), meta=np.ndarray>
- units :
- degree celcius (ITS-90)
- long_name :
- ARGO TEMPERATURE ANOMALY defined by Jan 2004 - Dec 2018 (15.0 year) RG CLIMATOLOGY
Array Chunk Bytes 2.03 GiB 47.79 MiB Shape (180, 58, 145, 360) (180, 58, 20, 60) Count 49 Tasks 48 Chunks Type float32 numpy.ndarray - BATHYMETRY_MASK(pres, lat, lon)float32dask.array<chunksize=(58, 20, 60), meta=np.ndarray>
- long_name :
- BATHYMETRY MASK
Array Chunk Bytes 11.55 MiB 271.88 kiB Shape (58, 145, 360) (58, 20, 60) Count 49 Tasks 48 Chunks Type float32 numpy.ndarray - MAPPING_MASK(pres, lat, lon)float32dask.array<chunksize=(58, 20, 60), meta=np.ndarray>
- long_name :
- MAPPING MASK: pressure limits of mapping can be shallower than 2000dbar in marginal seas
Array Chunk Bytes 11.55 MiB 271.88 kiB Shape (58, 145, 360) (58, 20, 60) Count 49 Tasks 48 Chunks Type float32 numpy.ndarray - T(time, pres, lat, lon)float32dask.array<chunksize=(180, 58, 20, 60), meta=np.ndarray>
- units :
- degree celcius (ITS-90)
- long_name :
- ARGO TEMPERATURE ANOMALY defined by Jan 2004 - Dec 2018 (15.0 year) RG CLIMATOLOGY
- standard_name :
- sea_water_potential_temperature
Array Chunk Bytes 2.03 GiB 47.79 MiB Shape (180, 58, 145, 360) (180, 58, 20, 60) Count 194 Tasks 48 Chunks Type float32 numpy.ndarray - Smean(pres, lat, lon)float32dask.array<chunksize=(58, 20, 60), meta=np.ndarray>
- units :
- Practical Salinity Scale 78
- long_name :
- ARGO SALINITY MEAN Jan 2004 - Dec 2018 (15.0 year) RG CLIMATOLOGY
Array Chunk Bytes 11.55 MiB 271.88 kiB Shape (58, 145, 360) (58, 20, 60) Count 49 Tasks 48 Chunks Type float32 numpy.ndarray - Sanom(time, pres, lat, lon)float32dask.array<chunksize=(180, 58, 20, 60), meta=np.ndarray>
- units :
- Practical Salinity Scale 78
- long_name :
- ARGO SALINITY ANOMALY defined by Jan 2004 - Dec 2018 (15.0 year) RG CLIMATOLOGY
Array Chunk Bytes 2.03 GiB 47.79 MiB Shape (180, 58, 145, 360) (180, 58, 20, 60) Count 49 Tasks 48 Chunks Type float32 numpy.ndarray - S(time, pres, lat, lon)float32dask.array<chunksize=(180, 58, 20, 60), meta=np.ndarray>
- units :
- Practical Salinity Scale 78
- long_name :
- ARGO SALINITY ANOMALY defined by Jan 2004 - Dec 2018 (15.0 year) RG CLIMATOLOGY
- standard_name :
- sea_water_salinity
Array Chunk Bytes 2.03 GiB 47.79 MiB Shape (180, 58, 145, 360) (180, 58, 20, 60) Count 194 Tasks 48 Chunks Type float32 numpy.ndarray
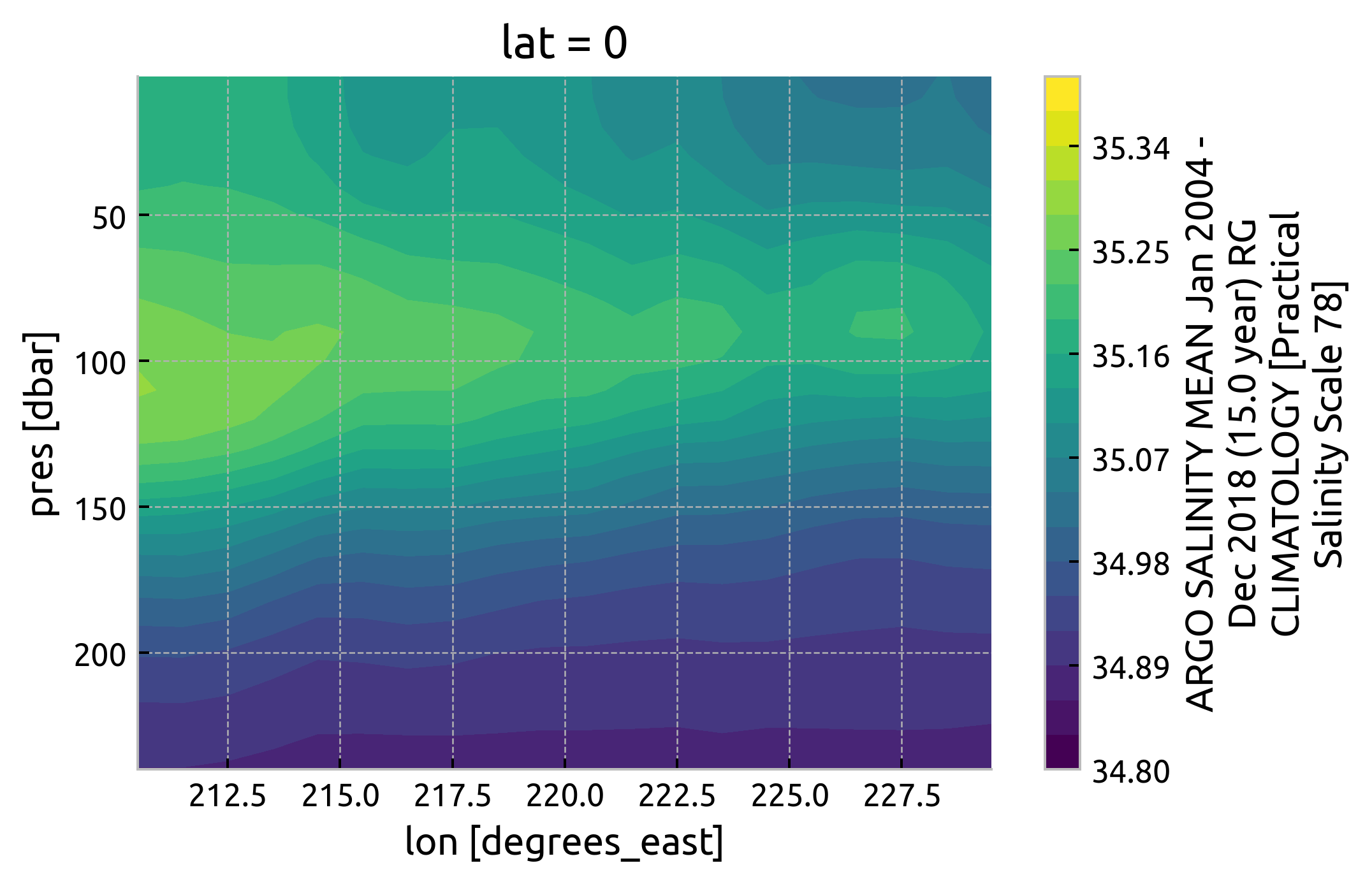
argoclim.Smean.interp(lat=0).sel(
# lon=slice(165, 360 - 80),
lon=slice(210, 230),
pres=slice(250),
).cf.plot.contourf(levels=21, vmin=34.8, vmax=35.4)
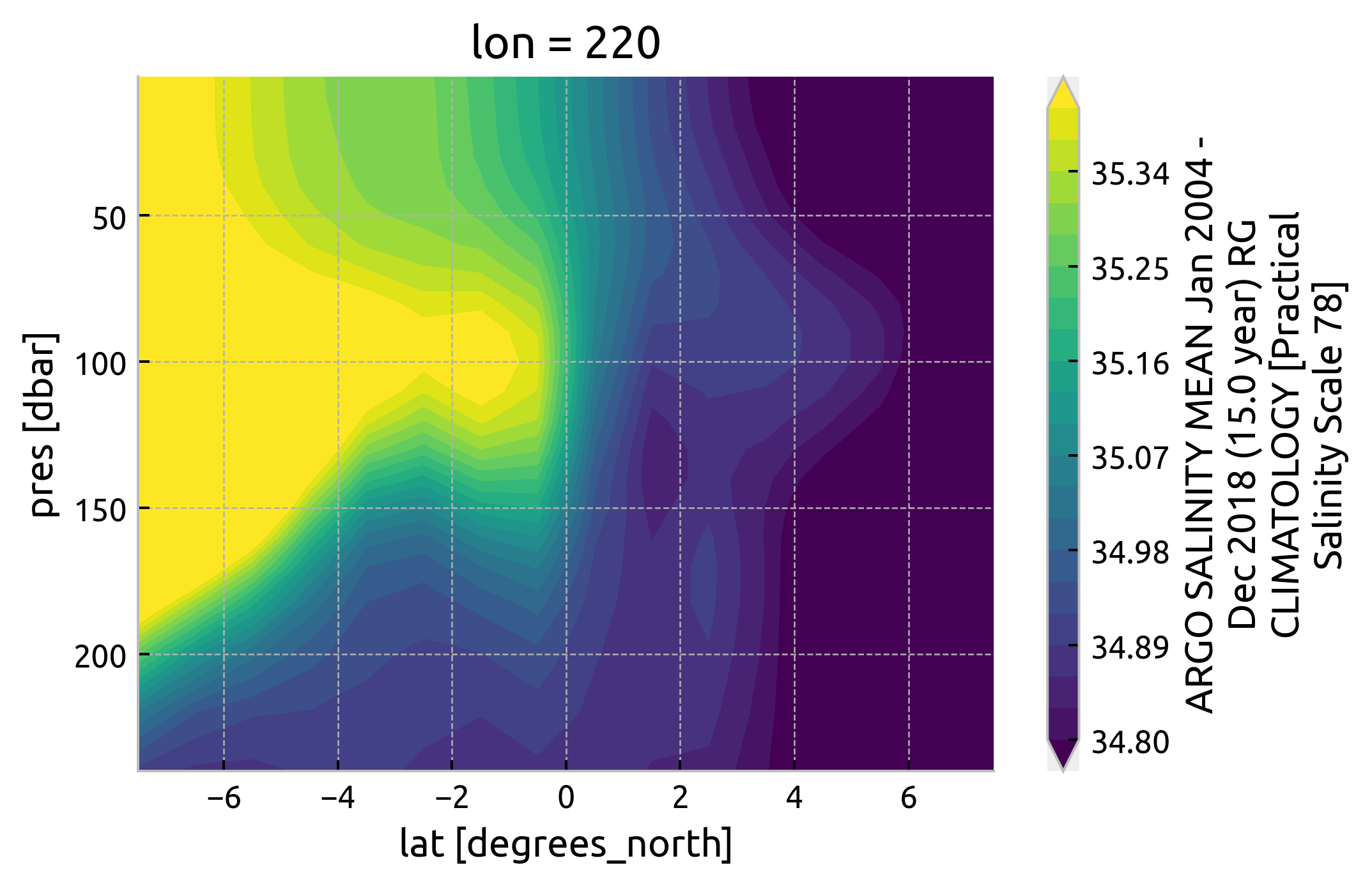
plt.figure()
argoclim.Smean.interp(lon=220).sel(lat=slice(-8, 8), pres=slice(250)).cf.plot.contourf(
levels=21, vmin=34.8, vmax=35.4
)
<matplotlib.contour.QuadContourSet at 0x7fd4a331e640>
older T-S attempt#
I first picked out three years 2008, 2010, 2015-16
ds = xr.open_mfdataset(
["argo_140_2008.nc", "argo_140_2010.nc", "argo_140_2016.nc"],
preprocess=lambda ds: ds.drop_vars("N_POINTS"),
concat_dim="N_POINTS",
combine="nested",
)
ds
<xarray.Dataset>
Dimensions: (N_POINTS: 62174)
Coordinates:
LATITUDE (N_POINTS) float64 dask.array<chunksize=(8464,), meta=np.ndarray>
LONGITUDE (N_POINTS) float64 dask.array<chunksize=(8464,), meta=np.ndarray>
TIME (N_POINTS) datetime64[ns] dask.array<chunksize=(8464,), meta=np.ndarray>
Dimensions without coordinates: N_POINTS
Data variables: (12/13)
CONFIG_MISSION_NUMBER (N_POINTS) int64 dask.array<chunksize=(8464,), meta=np.ndarray>
CYCLE_NUMBER (N_POINTS) int64 dask.array<chunksize=(8464,), meta=np.ndarray>
DATA_MODE (N_POINTS) object dask.array<chunksize=(8464,), meta=np.ndarray>
DIRECTION (N_POINTS) object dask.array<chunksize=(8464,), meta=np.ndarray>
PLATFORM_NUMBER (N_POINTS) int64 dask.array<chunksize=(8464,), meta=np.ndarray>
POSITION_QC (N_POINTS) int64 dask.array<chunksize=(8464,), meta=np.ndarray>
... ...
PRES_QC (N_POINTS) int64 dask.array<chunksize=(8464,), meta=np.ndarray>
PSAL (N_POINTS) float64 dask.array<chunksize=(8464,), meta=np.ndarray>
PSAL_QC (N_POINTS) int64 dask.array<chunksize=(8464,), meta=np.ndarray>
TEMP (N_POINTS) float64 dask.array<chunksize=(8464,), meta=np.ndarray>
TEMP_QC (N_POINTS) int64 dask.array<chunksize=(8464,), meta=np.ndarray>
TIME_QC (N_POINTS) int64 dask.array<chunksize=(8464,), meta=np.ndarray>
Attributes:
DATA_ID: ARGO
DOI: http://doi.org/10.17882/42182
Fetched_from: https://www.ifremer.fr/erddap
Fetched_by: deepak
Fetched_date: 2021/04/02
Fetched_constraints: [x=-145.00/-135.00; y=-1.00/1.00; z=0.0/500.0; t=20...
Fetched_uri: https://www.ifremer.fr/erddap/tabledap/ArgoFloats.n...
history: Variables filtered according to DATA_MODE; Variable...- N_POINTS: 62174
- LATITUDE(N_POINTS)float64dask.array<chunksize=(8464,), meta=np.ndarray>
- _CoordinateAxisType :
- Lat
- actual_range :
- [ 1.36981406e-312 -1.99480466e-201]
- axis :
- Y
- colorBarMaximum :
- 90.0
- colorBarMinimum :
- -90.0
- ioos_category :
- Location
- long_name :
- Latitude of the station, best estimate
- standard_name :
- latitude
- units :
- degrees_north
- valid_max :
- 90.0
- valid_min :
- -90.0
Array Chunk Bytes 497.39 kB 324.62 kB Shape (62174,) (40578,) Count 9 Tasks 3 Chunks Type float64 numpy.ndarray - LONGITUDE(N_POINTS)float64dask.array<chunksize=(8464,), meta=np.ndarray>
- _CoordinateAxisType :
- Lon
- actual_range :
- [-1.91533933e+093 2.05277202e-312]
- axis :
- X
- colorBarMaximum :
- 180.0
- colorBarMinimum :
- -180.0
- ioos_category :
- Location
- long_name :
- Longitude of the station, best estimate
- standard_name :
- longitude
- units :
- degrees_east
- valid_max :
- 180.0
- valid_min :
- -180.0
Array Chunk Bytes 497.39 kB 324.62 kB Shape (62174,) (40578,) Count 9 Tasks 3 Chunks Type float64 numpy.ndarray - TIME(N_POINTS)datetime64[ns]dask.array<chunksize=(8464,), meta=np.ndarray>
- _CoordinateAxisType :
- Time
- actual_range :
- [6.99589853e-310 6.97419820e-310]
- axis :
- T
- ioos_category :
- Time
- long_name :
- Julian day (UTC) of the station relative to REFERENCE_DATE_TIME
- standard_name :
- time
- time_origin :
- 01-JAN-1970 00:00:00
Array Chunk Bytes 497.39 kB 324.62 kB Shape (62174,) (40578,) Count 9 Tasks 3 Chunks Type datetime64[ns] numpy.ndarray
- CONFIG_MISSION_NUMBER(N_POINTS)int64dask.array<chunksize=(8464,), meta=np.ndarray>
- actual_range :
- [16777216 16777216]
- colorBarMaximum :
- 100.0
- colorBarMinimum :
- 0.0
- conventions :
- 1...N, 1 : first complete mission
- ioos_category :
- Statistics
- long_name :
- Unique number denoting the missions performed by the float
- casted :
- 1
Array Chunk Bytes 497.39 kB 324.62 kB Shape (62174,) (40578,) Count 9 Tasks 3 Chunks Type int64 numpy.ndarray - CYCLE_NUMBER(N_POINTS)int64dask.array<chunksize=(8464,), meta=np.ndarray>
- long_name :
- Float cycle number
- convention :
- 0..N, 0 : launch cycle (if exists), 1 : first complete cycle
- casted :
- 1
Array Chunk Bytes 497.39 kB 324.62 kB Shape (62174,) (40578,) Count 9 Tasks 3 Chunks Type int64 numpy.ndarray - DATA_MODE(N_POINTS)objectdask.array<chunksize=(8464,), meta=np.ndarray>
- long_name :
- Delayed mode or real time data
- convention :
- R : real time; D : delayed mode; A : real time with adjustment
- casted :
- 1
Array Chunk Bytes 497.39 kB 324.62 kB Shape (62174,) (40578,) Count 9 Tasks 3 Chunks Type object numpy.ndarray - DIRECTION(N_POINTS)objectdask.array<chunksize=(8464,), meta=np.ndarray>
- long_name :
- Direction of the station profiles
- convention :
- A: ascending profiles, D: descending profiles
- casted :
- 1
Array Chunk Bytes 497.39 kB 324.62 kB Shape (62174,) (40578,) Count 9 Tasks 3 Chunks Type object numpy.ndarray - PLATFORM_NUMBER(N_POINTS)int64dask.array<chunksize=(8464,), meta=np.ndarray>
- long_name :
- Float unique identifier
- convention :
- WMO float identifier : A9IIIII
- casted :
- 1
Array Chunk Bytes 497.39 kB 324.62 kB Shape (62174,) (40578,) Count 9 Tasks 3 Chunks Type int64 numpy.ndarray - POSITION_QC(N_POINTS)int64dask.array<chunksize=(8464,), meta=np.ndarray>
- long_name :
- Global quality flag of POSITION_QC profile
- convention :
- Argo reference table 2a
- casted :
- 1
Array Chunk Bytes 497.39 kB 324.62 kB Shape (62174,) (40578,) Count 9 Tasks 3 Chunks Type int64 numpy.ndarray - PRES(N_POINTS)float64dask.array<chunksize=(8464,), meta=np.ndarray>
- long_name :
- Sea Pressure
- standard_name :
- sea_water_pressure
- units :
- decibar
- valid_min :
- 0.0
- valid_max :
- 12000.0
- resolution :
- 0.1
- axis :
- Z
- casted :
- 1
Array Chunk Bytes 497.39 kB 324.62 kB Shape (62174,) (40578,) Count 9 Tasks 3 Chunks Type float64 numpy.ndarray - PRES_QC(N_POINTS)int64dask.array<chunksize=(8464,), meta=np.ndarray>
- long_name :
- Global quality flag of PRES_QC profile
- convention :
- Argo reference table 2a
- casted :
- 1
Array Chunk Bytes 497.39 kB 324.62 kB Shape (62174,) (40578,) Count 9 Tasks 3 Chunks Type int64 numpy.ndarray - PSAL(N_POINTS)float64dask.array<chunksize=(8464,), meta=np.ndarray>
- long_name :
- PRACTICAL SALINITY
- standard_name :
- sea_water_salinity
- units :
- psu
- valid_min :
- 0.0
- valid_max :
- 43.0
- resolution :
- 0.001
- casted :
- 1
Array Chunk Bytes 497.39 kB 324.62 kB Shape (62174,) (40578,) Count 9 Tasks 3 Chunks Type float64 numpy.ndarray - PSAL_QC(N_POINTS)int64dask.array<chunksize=(8464,), meta=np.ndarray>
- long_name :
- Global quality flag of PSAL_QC profile
- convention :
- Argo reference table 2a
- casted :
- 1
Array Chunk Bytes 497.39 kB 324.62 kB Shape (62174,) (40578,) Count 9 Tasks 3 Chunks Type int64 numpy.ndarray - TEMP(N_POINTS)float64dask.array<chunksize=(8464,), meta=np.ndarray>
- long_name :
- SEA TEMPERATURE IN SITU ITS-90 SCALE
- standard_name :
- sea_water_temperature
- units :
- degree_Celsius
- valid_min :
- -2.0
- valid_max :
- 40.0
- resolution :
- 0.001
- casted :
- 1
Array Chunk Bytes 497.39 kB 324.62 kB Shape (62174,) (40578,) Count 9 Tasks 3 Chunks Type float64 numpy.ndarray - TEMP_QC(N_POINTS)int64dask.array<chunksize=(8464,), meta=np.ndarray>
- long_name :
- Global quality flag of TEMP_QC profile
- convention :
- Argo reference table 2a
- casted :
- 1
Array Chunk Bytes 497.39 kB 324.62 kB Shape (62174,) (40578,) Count 9 Tasks 3 Chunks Type int64 numpy.ndarray - TIME_QC(N_POINTS)int64dask.array<chunksize=(8464,), meta=np.ndarray>
- long_name :
- Global quality flag of TIME_QC profile
- convention :
- Argo reference table 2a
- casted :
- 1
Array Chunk Bytes 497.39 kB 324.62 kB Shape (62174,) (40578,) Count 9 Tasks 3 Chunks Type int64 numpy.ndarray
- DATA_ID :
- ARGO
- DOI :
- http://doi.org/10.17882/42182
- Fetched_from :
- https://www.ifremer.fr/erddap
- Fetched_by :
- deepak
- Fetched_date :
- 2021/04/02
- Fetched_constraints :
- [x=-145.00/-135.00; y=-1.00/1.00; z=0.0/500.0; t=2008-05-01/2009-05-01]
- Fetched_uri :
- https://www.ifremer.fr/erddap/tabledap/ArgoFloats.nc?data_mode,latitude,longitude,position_qc,time,time_qc,direction,platform_number,cycle_number,config_mission_number,vertical_sampling_scheme,pres,temp,psal,pres_qc,temp_qc,psal_qc,pres_adjusted,temp_adjusted,psal_adjusted,pres_adjusted_qc,temp_adjusted_qc,psal_adjusted_qc,pres_adjusted_error,temp_adjusted_error,psal_adjusted_error&longitude>=-145&longitude<=-135&latitude>=-1&latitude<=1&pres>=0&pres<=500&time>=1209600000.0&time<=1241136000.0&distinct()&orderBy("time,pres")
- history :
- Variables filtered according to DATA_MODE; Variables selected according to QC
ds.TIME.dt.year.compute()
<xarray.DataArray 'year' (N_POINTS: 62174)>
array([2008, 2008, 2008, ..., 2016, 2016, 2016])
Coordinates:
LATITUDE (N_POINTS) float64 0.707 0.707 0.707 ... -0.202 -0.202 -0.202
LONGITUDE (N_POINTS) float64 -136.5 -136.5 -136.5 ... -135.6 -135.6 -135.6
TIME (N_POINTS) datetime64[ns] 2008-05-01T16:43:14 ... 2016-04-29T0...
Dimensions without coordinates: N_POINTS- N_POINTS: 62174
- 2008 2008 2008 2008 2008 2008 2008 ... 2016 2016 2016 2016 2016 2016
array([2008, 2008, 2008, ..., 2016, 2016, 2016])
- LATITUDE(N_POINTS)float640.707 0.707 0.707 ... -0.202 -0.202
- _CoordinateAxisType :
- Lat
- actual_range :
- [ 1.36981406e-312 -1.99480466e-201]
- axis :
- Y
- colorBarMaximum :
- 90.0
- colorBarMinimum :
- -90.0
- ioos_category :
- Location
- long_name :
- Latitude of the station, best estimate
- standard_name :
- latitude
- units :
- degrees_north
- valid_max :
- 90.0
- valid_min :
- -90.0
array([ 0.707, 0.707, 0.707, ..., -0.202, -0.202, -0.202])
- LONGITUDE(N_POINTS)float64-136.5 -136.5 ... -135.6 -135.6
- _CoordinateAxisType :
- Lon
- actual_range :
- [-1.91533933e+093 2.05277202e-312]
- axis :
- X
- colorBarMaximum :
- 180.0
- colorBarMinimum :
- -180.0
- ioos_category :
- Location
- long_name :
- Longitude of the station, best estimate
- standard_name :
- longitude
- units :
- degrees_east
- valid_max :
- 180.0
- valid_min :
- -180.0
array([-136.527, -136.527, -136.527, ..., -135.644, -135.644, -135.644])
- TIME(N_POINTS)datetime64[ns]2008-05-01T16:43:14 ... 2016-04-...
- _CoordinateAxisType :
- Time
- actual_range :
- [6.99589853e-310 6.97419820e-310]
- axis :
- T
- ioos_category :
- Time
- long_name :
- Julian day (UTC) of the station relative to REFERENCE_DATE_TIME
- standard_name :
- time
- time_origin :
- 01-JAN-1970 00:00:00
array(['2008-05-01T16:43:14.000000000', '2008-05-01T16:43:14.000000000', '2008-05-01T16:43:14.000000000', ..., '2016-04-29T08:37:40.000000000', '2016-04-29T08:37:40.000000000', '2016-04-29T08:37:40.000000000'], dtype='datetime64[ns]')
ds.coords["year"] = ds.TIME.dt.year
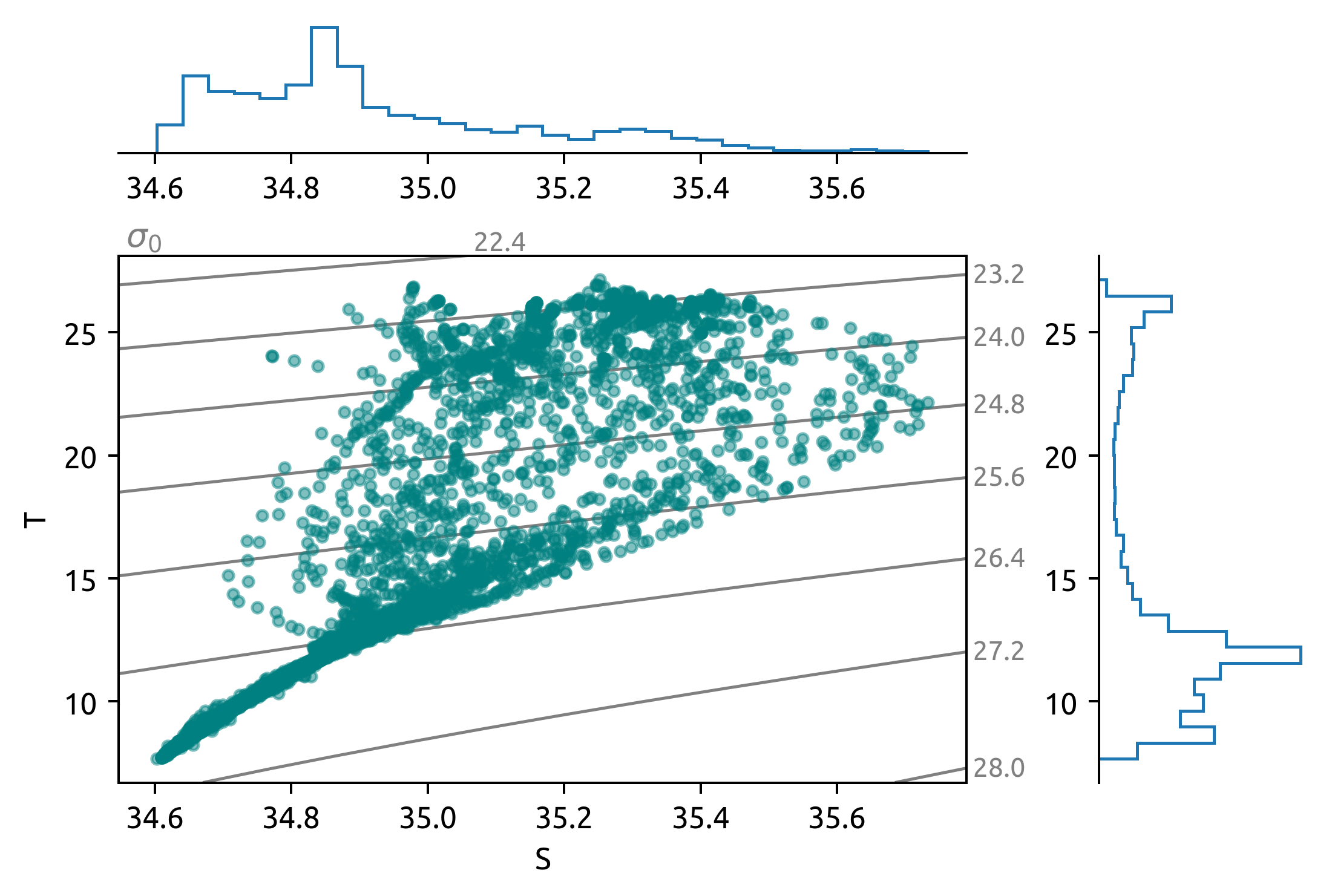
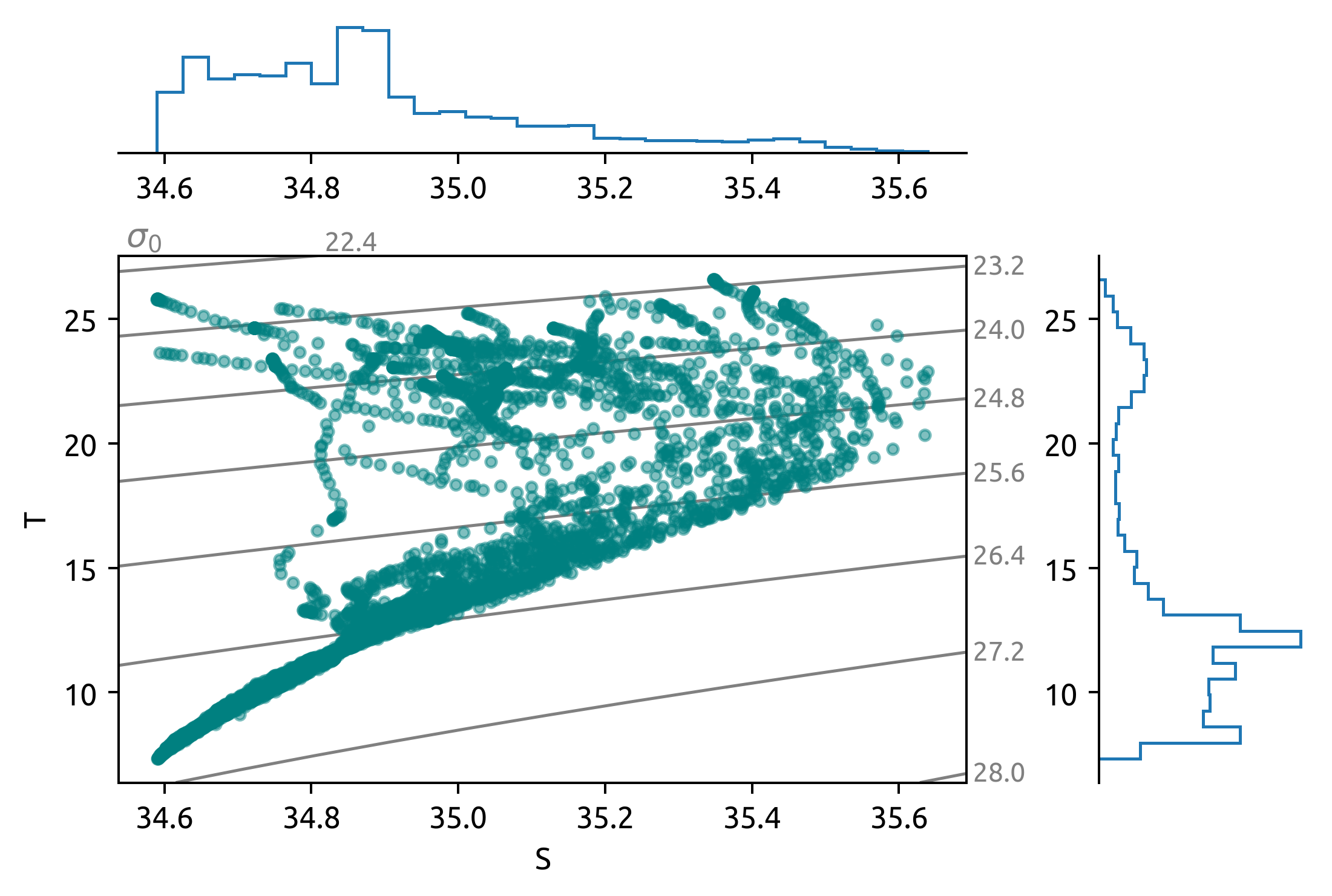
sub = ds.query({"N_POINTS": "year == 2008"})
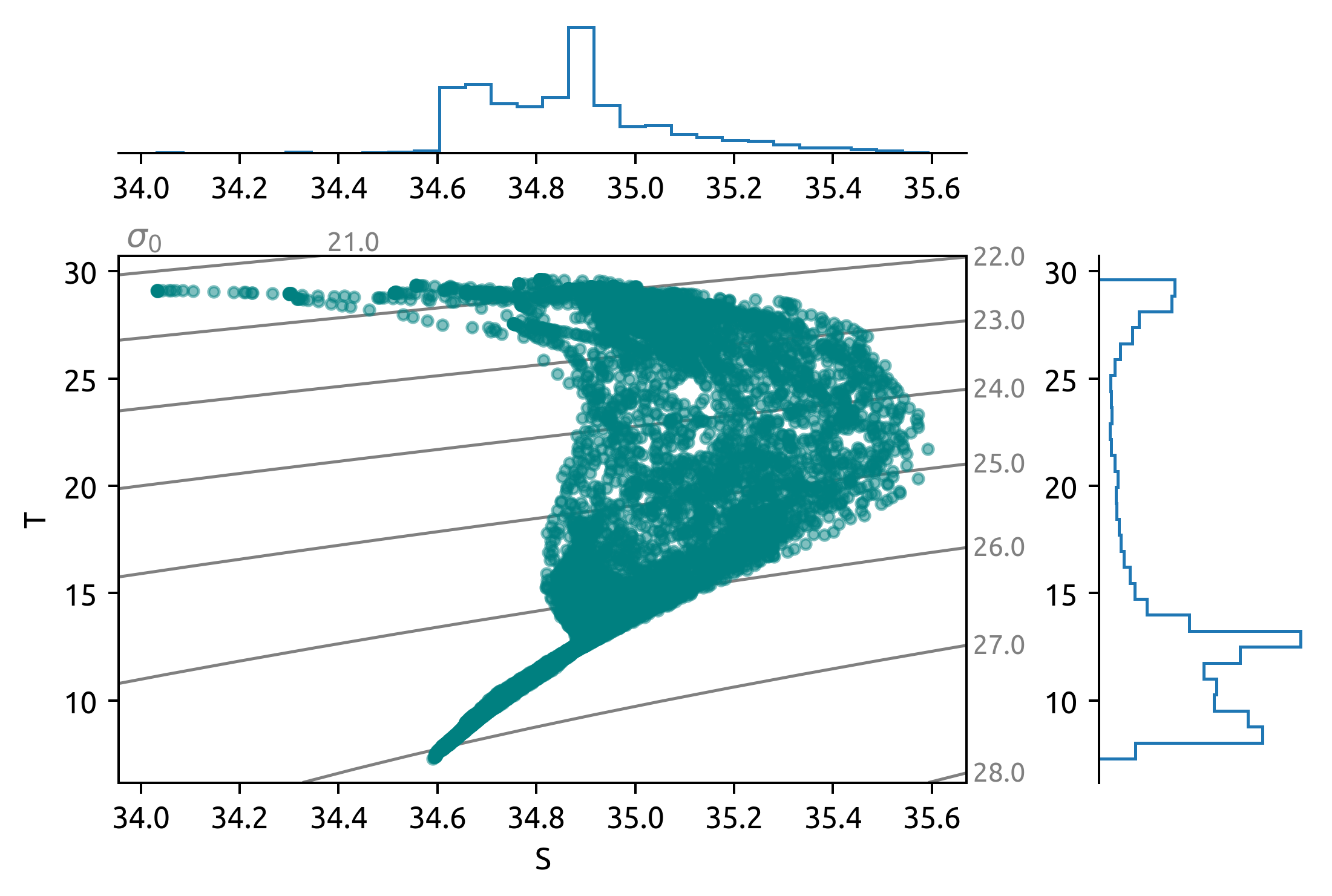
dcpy.oceans.TSplot(sub.PSAL, sub.TEMP, hexbin=False)
sub = ds.query({"N_POINTS": "year == 2010"})
dcpy.oceans.TSplot(sub.PSAL, sub.TEMP, hexbin=False)
sub = ds.query({"N_POINTS": "year == 2016"})
dcpy.oceans.TSplot(sub.PSAL, sub.TEMP, hexbin=False);
Download Argo data#
from argopy import DataFetcher as ArgoDataFetcher
argo_loader = ArgoDataFetcher()
NATRE#
ds = xr.open_mfdataset(
"argo_natre_*.nc",
preprocess=lambda ds: ds.drop_vars("N_POINTS"),
concat_dim="N_POINTS",
combine="nested",
).load()
ds.to_netcdf("../datasets/argo/natre.nc")
ds = argo_loader.region(
[-35, -25, 23, 28, 0, 2000, "2005-01", "2010-12-31"]
).to_xarray()
ds.to_netcdf("argo_natre_2005_2010.nc")
ds = argo_loader.region(
[-35, -25, 23, 28, 0, 2000, "2011-01", "2015-12-31"]
).to_xarray()
ds.to_netcdf("argo_natre_20010_2015.nc")
ds = argo_loader.region(
[-35, -25, 23, 28, 0, 2000, "2016-01", "2020-12-31"]
).to_xarray()
ds.to_netcdf("argo_natre_2016_2020.nc")
TAO#
ds = xr.open_mfdataset(
"argo_140_20*_*.nc",
preprocess=lambda ds: ds.drop_vars("N_POINTS"),
concat_dim="N_POINTS",
combine="nested",
).load()
ds.to_netcdf("../datasets/argo/tao.nc")
ds = argo_loader.region(
[-145, -135, -2, 8, 0, 500, "2005-01", "2010-12-31"]
).to_xarray()
ds.to_netcdf("argo_140_2005_2010.nc")
ds = argo_loader.region(
[-145, -135, -2, 8, 0, 500, "2011-01", "2015-12-31"]
).to_xarray()
ds.to_netcdf("argo_140_2011_2015.nc")
# Too much data end up getting HTTP 413: Payload too large errors.
for year in range(2016, 2021):
ds = argo_loader.region(
[-145, -135, -2, 8, 0, 500, f"{year}-01-01", f"{year}-12-31"]
).to_xarray()
ds.to_netcdf(f"argo_140_{year}_{year}.nc")
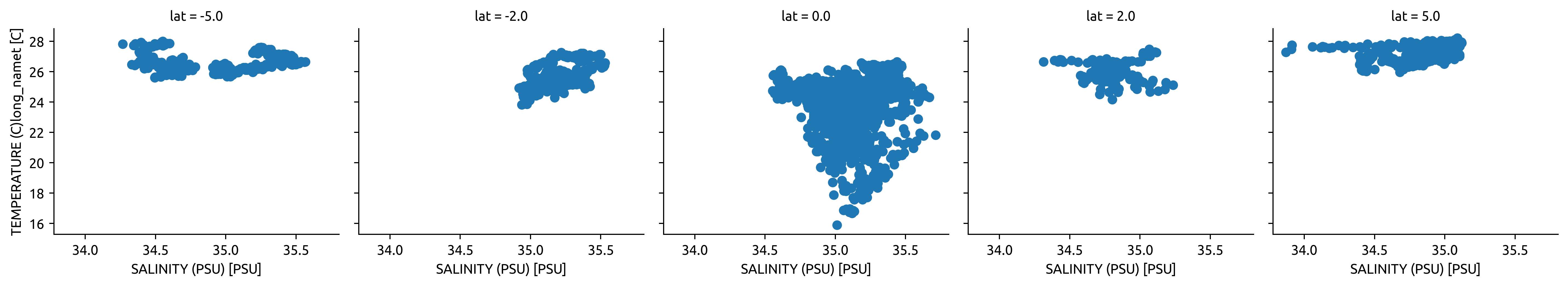
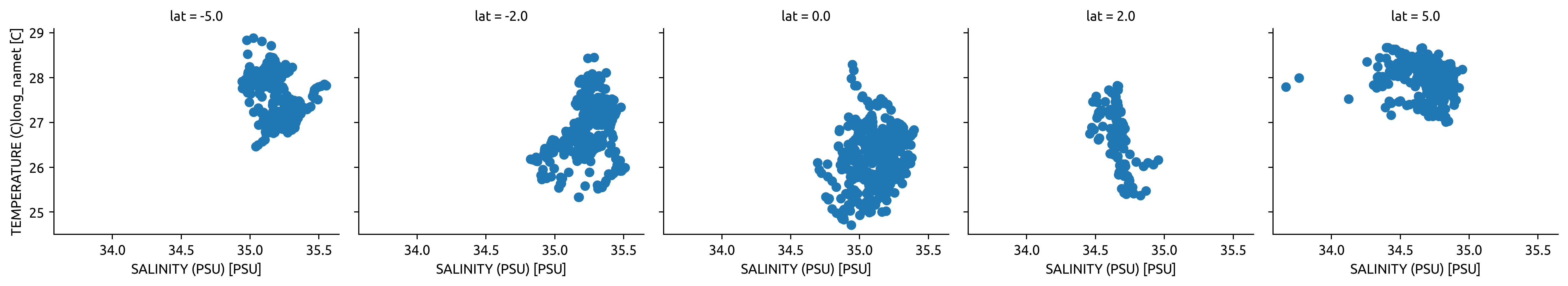
TAO mooring T-S#
Conclusion There is not enough data. Basically no salinity observations off the equator.
import pump
temp = (
xr.open_dataset("/home/deepak/TaoTritonPirataRama/TAO_TRITON/t_xyzt_dy.cdf")
.rename({"T_20": "temp"})
.cf.guess_coord_axis()
.sel(lon=220)
)
salt = (
xr.open_dataset("/home/deepak/TaoTritonPirataRama/TAO_TRITON/s_xyzt_dy.cdf")
.rename({"S_41": "salt"})
.cf.guess_coord_axis()
.sel(lon=220)
)
tao = xr.merge([temp, salt], join="inner").sel(lat=[-5, -2, 0, 2, 5])
tao = tao.where(tao < 1000)
tao.sel(time="2003").plot.scatter("salt", "temp", col="lat")
tao.sel(time=slice("2008-Jun", "2009-Mar")).plot.scatter("salt", "temp", col="lat")
<xarray.plot.facetgrid.FacetGrid at 0x7f78fe81bee0>


